

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC141111.4 - phase: 0 /pseudo
(1069 letters)
Database: MTGI
36,976 sequences; 27,044,181 total letters
Searching..................................................done
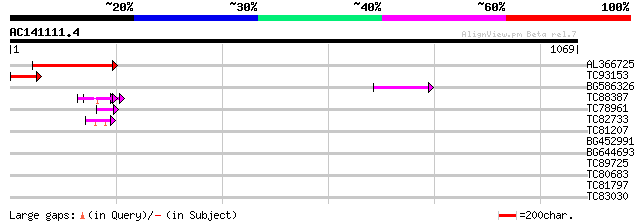
Score E
Sequences producing significant alignments: (bits) Value
AL366725 259 5e-69
TC93153 similar to GP|14715220|emb|CAC44106. gag polyprotein {Ci... 116 5e-26
BG586326 similar to PIR|G84493|G8 probable retroelement pol poly... 47 3e-05
TC88387 similar to GP|13357253|gb|AAK20050.1 putative zinc finge... 46 7e-05
TC78961 similar to GP|18252179|gb|AAL61922.1 unknown protein {Ar... 44 4e-04
TC82733 similar to GP|10177404|dbj|BAB10535. gene_id:K24M7.12~pi... 43 7e-04
TC81207 similar to GP|21322752|dbj|BAB78536. cold shock protein-... 41 0.002
BG452991 PIR|A25875|A25 histone H4 - Tetrahymena thermophila, pa... 37 0.003
BG644693 weakly similar to GP|18767374|g Putative 22 kDa kafirin... 38 0.018
TC89725 similar to PIR|T05494|T05494 glycine-rich protein T19K4.... 32 1.3
TC80683 homologue to GP|8777424|dbj|BAA97014.1 gb|AAF56406.1~gen... 31 2.9
TC81797 similar to GP|17065224|gb|AAL32766.1 Unknown protein {Ar... 30 3.8
TC83030 weakly similar to GP|18855061|gb|AAL79753.1 putative RNA... 30 3.8
>AL366725
Length = 485
Score = 259 bits (661), Expect = 5e-69
Identities = 119/161 (73%), Positives = 136/161 (83%)
Frame = +2
Query: 43 KFYPHYTAETAEFSKCIKIENGLRADIKRAIGYQKIRNFSELVSSCRIYEEDTKAHYKVM 102
KFYPHY AETAEFSKCIK ENGLR DIKRAIGYQ++R F +LV++CRIYEEDTKAH KV+
Sbjct: 2 KFYPHYAAETAEFSKCIKFENGLRPDIKRAIGYQQLRVFPDLVNTCRIYEEDTKAHDKVV 181
Query: 103 SERRGKGQLSRPKPYSAPPDKGK*RLKDERRPKMRDAPTDIVCFKCGEKGHKSNVCDRDE 162
+ER+ KGQ SRPKPYSAP DKGK R+ D+RRPK +DAP +IVCF GEKGHKSNVC ++
Sbjct: 182 NERKTKGQ*SRPKPYSAPADKGKQRMVDDRRPKKKDAPAEIVCFNYGEKGHKSNVCPKEI 361
Query: 163 KKCFRCGKKGHTLADCKRGDIVCYNFNEEGHISLQCTQPKK 203
KKC RC KKGH +ADCKR DIVC+N NEEGHI QC QPK+
Sbjct: 362 KKCVRCDKKGHIVADCKRNDIVCFNCNEEGHIGSQCKQPKR 484
>TC93153 similar to GP|14715220|emb|CAC44106. gag polyprotein {Cicer
arietinum}, partial (8%)
Length = 516
Score = 116 bits (290), Expect = 5e-26
Identities = 55/59 (93%), Positives = 57/59 (96%)
Frame = +2
Query: 1 RREFLNRYFPEDVRGKKEIEFLELKQGDMSVTEYAAKFVELAKFYPHYTAETAEFSKCI 59
R+EFL RYFPEDVRGKKEIEFLELKQGDMSVTEYAAKFVELA FYPHY+AETAEFSKCI
Sbjct: 338 RKEFLGRYFPEDVRGKKEIEFLELKQGDMSVTEYAAKFVELATFYPHYSAETAEFSKCI 514
>BG586326 similar to PIR|G84493|G8 probable retroelement pol polyprotein
[imported] - Arabidopsis thaliana, partial (13%)
Length = 736
Score = 47.4 bits (111), Expect = 3e-05
Identities = 45/113 (39%), Positives = 56/113 (48%)
Frame = +3
Query: 686 FIVMRPSWA*VVF*CKMVKW*LMLLGS*RFMKRIILHMIWSWLRWFLF*KFGGIICTVPD 745
FI R S *V +* M + GS* M+ MI WLR + *+FG C VP
Sbjct: 54 FIQTRLSLD*VAY*PSMRRSSPTRQGS*ENMRETTPPMI*KWLR*YSP*RFGAHTCMVPR 233
Query: 746 LKCLVITKV*NIYLIRKS*I*GREDGLTC*RIMILV*ITIRVKLMCLQML*AG 798
+ + KV*+I+L S* *GR G * I + IR KL+ Q L*AG
Sbjct: 234 FRYIRTIKV*SIFLPSLS*T*GRGGGWNS*LTTI*TSLIIREKLIW*QTL*AG 392
>TC88387 similar to GP|13357253|gb|AAK20050.1 putative zinc finger protein
{Oryza sativa (japonica cultivar-group)}, partial (96%)
Length = 1286
Score = 46.2 bits (108), Expect = 7e-05
Identities = 24/77 (31%), Positives = 39/77 (50%), Gaps = 2/77 (2%)
Frame = +1
Query: 129 KDERRPK--MRDAPTDIVCFKCGEKGHKSNVCDRDEKKCFRCGKKGHTLADCKRGDIVCY 186
K+ +RP +R+ P VC C GH ++ C + C+ C + GH + C I C+
Sbjct: 427 KNCKRPGHYVRECPNVAVCHNCSLPGHIASECST-KSLCWNCKEPGHMASSCPNEGI-CH 600
Query: 187 NFNEEGHISLQCTQPKK 203
+ GH + +CT P+K
Sbjct: 601 TCGKAGHRARECTVPQK 651
Score = 44.3 bits (103), Expect = 3e-04
Identities = 26/77 (33%), Positives = 38/77 (48%), Gaps = 1/77 (1%)
Frame = +1
Query: 140 PTDI-VCFKCGEKGHKSNVCDRDEKKCFRCGKKGHTLADCKRGDIVCYNFNEEGHISLQC 198
P D+ +C C ++GH + C +EK C C K GH DC D +C N GH++ QC
Sbjct: 655 PGDLRLCNNCYKQGHIAVECT-NEKACNNCRKTGHLARDCP-NDPICNLCNISGHVARQC 828
Query: 199 TQPKKVRTGGKVFALTG 215
+ + G +L G
Sbjct: 829 PKSNVIGDRGGGGSLRG 879
Score = 43.1 bits (100), Expect = 6e-04
Identities = 19/65 (29%), Positives = 31/65 (47%), Gaps = 6/65 (9%)
Frame = +1
Query: 140 PTDIVCFKCGEKGHKSNVCDRDEKK------CFRCGKKGHTLADCKRGDIVCYNFNEEGH 193
P + +C CG+ GH++ C +K C C K+GH +C + C N + GH
Sbjct: 580 PNEGICHTCGKAGHRARECTVPQKPPGDLRLCNNCYKQGHIAVEC-TNEKACNNCRKTGH 756
Query: 194 ISLQC 198
++ C
Sbjct: 757 LARDC 771
Score = 41.6 bits (96), Expect = 0.002
Identities = 23/78 (29%), Positives = 31/78 (39%)
Frame = +1
Query: 132 RRPKMRDAPTDIVCFKCGEKGHKSNVCDRDEKKCFRCGKKGHTLADCKRGDIVCYNFNEE 191
RR R D +C C GH C + C C GH ++C + C+N E
Sbjct: 385 RRDSRRGFSQDNLCKNCKRPGHYVRECP-NVAVCHNCSLPGHIASECSTKSL-CWNCKEP 558
Query: 192 GHISLQCTQPKKVRTGGK 209
GH++ C T GK
Sbjct: 559 GHMASSCPNEGICHTCGK 612
Score = 36.2 bits (82), Expect = 0.069
Identities = 21/85 (24%), Positives = 30/85 (34%), Gaps = 23/85 (27%)
Frame = +1
Query: 137 RDAPTDIVCFKCGEKGHKSNVCDRD----------------------EKKCFRCGKKGHT 174
RD P D +C C GH + C + + C C + GH
Sbjct: 763 RDCPNDPICNLCNISGHVARQCPKSNVIGDRGGGGSLRGGYRDGGFRDVVCRSCQQFGHM 942
Query: 175 LADCKRGDI-VCYNFNEEGHISLQC 198
DC G + +C N GH + +C
Sbjct: 943 SRDCMGGPLMICQNCGGRGHQAYEC 1017
Score = 33.1 bits (74), Expect = 0.59
Identities = 16/44 (36%), Positives = 20/44 (45%), Gaps = 1/44 (2%)
Frame = +1
Query: 142 DIVCFKCGEKGHKSNVCDRDEKK-CFRCGKKGHTLADCKRGDIV 184
D+VC C + GH S C C CG +GH +C G V
Sbjct: 904 DVVCRSCQQFGHMSRDCMGGPLMICQNCGGRGHQAYECPSGRFV 1035
>TC78961 similar to GP|18252179|gb|AAL61922.1 unknown protein {Arabidopsis
thaliana}, partial (71%)
Length = 974
Score = 43.5 bits (101), Expect = 4e-04
Identities = 21/44 (47%), Positives = 23/44 (51%), Gaps = 3/44 (6%)
Frame = +2
Query: 164 KCFRCGKKGHTLADCKRGD--IVCYNFNEEGHISLQC-TQPKKV 204
+CF CG GH DCK GD CY E GHI C PKK+
Sbjct: 356 RCFNCGIDGHWARDCKAGDWKNKCYRCGERGHIEKNCKNSPKKL 487
Score = 40.8 bits (94), Expect = 0.003
Identities = 16/37 (43%), Positives = 22/37 (59%), Gaps = 2/37 (5%)
Frame = +2
Query: 145 CFKCGEKGHKSNVCDRDE--KKCFRCGKKGHTLADCK 179
CF CG GH + C + KC+RCG++GH +CK
Sbjct: 359 CFNCGIDGHWARDCKAGDWKNKCYRCGERGHIEKNCK 469
>TC82733 similar to GP|10177404|dbj|BAB10535.
gene_id:K24M7.12~pir||S42136~similar to unknown protein
{Arabidopsis thaliana}, partial (57%)
Length = 710
Score = 42.7 bits (99), Expect = 7e-04
Identities = 21/67 (31%), Positives = 34/67 (50%), Gaps = 12/67 (17%)
Frame = +3
Query: 144 VCFKCGEKGHKSNVCD-----RDEKKCFRCGKKGHTLADC----KRGDIV---CYNFNEE 191
+C +C +GH++ C D K + CG GH+LA+C + G + C+ E+
Sbjct: 354 ICLRCRRRGHRAQNCPDGGSKEDFKY*YNCGDNGHSLANCPHPLQEGGTMFAQCFVCKEQ 533
Query: 192 GHISLQC 198
GH+S C
Sbjct: 534 GHLSKNC 554
Score = 41.6 bits (96), Expect = 0.002
Identities = 35/119 (29%), Positives = 50/119 (41%), Gaps = 11/119 (9%)
Frame = +3
Query: 105 RRGKGQLSRPKPYS-APPDKGK*RLKDERRPKMRDAPTDIVCFKCGEKGHKSNVCDRD-- 161
++ K + R KP S + P GK L R P M+ + CF C H + C +
Sbjct: 177 KKKKNKFKRKKPDSNSKPRTGKRPL---RVPGMKPGDS---CFICKGLDHIAKFCTQKAE 338
Query: 162 ---EKKCFRCGKKGHTLADCKRGDI-----VCYNFNEEGHISLQCTQPKKVRTGGKVFA 212
K C RC ++GH +C G YN + GH C P ++ GG +FA
Sbjct: 339 WEKNKICLRCRRRGHRAQNCPDGGSKEDFKY*YNCGDNGHSLANCPHP--LQEGGTMFA 509
>TC81207 similar to GP|21322752|dbj|BAB78536. cold shock protein-1 {Triticum
aestivum}, partial (39%)
Length = 630
Score = 41.2 bits (95), Expect = 0.002
Identities = 23/88 (26%), Positives = 30/88 (33%), Gaps = 31/88 (35%)
Frame = +3
Query: 145 CFKCGEKGHKSNVCDRDEKK-----------------CFRCGKKGHTLADCKR------- 180
C+ CG+ GH + CDR ++ C+ CG H DC R
Sbjct: 351 CYTCGDTGHIARDCDRSDRNDRNDRSGGGGGGDRDRACYTCGSFEHFARDCMRGGGNNNN 530
Query: 181 -------GDIVCYNFNEEGHISLQCTQP 201
G CY GHI+ C P
Sbjct: 531 GGGGYGGGGTSCYRCGGVGHIARDCATP 614
>BG452991 PIR|A25875|A25 histone H4 - Tetrahymena thermophila, partial (33%)
Length = 560
Score = 37.4 bits (85), Expect(2) = 0.003
Identities = 26/61 (42%), Positives = 36/61 (58%)
Frame = +1
Query: 818 ET*VLSVSCHLRVYSWVC*RLIVIS*TVSEKHIKLMSSLLI*WLLVMKLKIMTSRLMIKV 877
ET*V V C RV +W C*R VS++ + M S I* L +++LK++ +LMIK
Sbjct: 82 ET*VWFVKCRHRV*NWGC*RSTTNFWIVSKRLRRWM*SW*I*CLGIIRLKMVILKLMIKE 261
Query: 878 C 878
C
Sbjct: 262 C 264
Score = 22.3 bits (46), Expect(2) = 0.003
Identities = 9/12 (75%), Positives = 10/12 (83%)
Frame = +3
Query: 802 TCLL*W*KSLSC 813
TCLL*W +S SC
Sbjct: 33 TCLL*WLESWSC 68
>BG644693 weakly similar to GP|18767374|g Putative 22 kDa kafirin cluster;
Ty3-Gypsy type {Oryza sativa}, partial (15%)
Length = 716
Score = 38.1 bits (87), Expect = 0.018
Identities = 16/27 (59%), Positives = 20/27 (73%)
Frame = +2
Query: 557 LDKFVVVFIDDILIYSETEEEHAEHLK 583
LD V+VF +DILIYS+ E EH HL+
Sbjct: 506 LDSLVIVFSNDILIYSKNENEHENHLR 586
>TC89725 similar to PIR|T05494|T05494 glycine-rich protein T19K4.150 -
Arabidopsis thaliana, partial (17%)
Length = 378
Score = 32.0 bits (71), Expect = 1.3
Identities = 15/57 (26%), Positives = 19/57 (33%), Gaps = 20/57 (35%)
Frame = +1
Query: 145 CFKCGEKGHKSNVCDRDEK--------------------KCFRCGKKGHTLADCKRG 181
C+ CGE GH + C C+ CG+ GH DC G
Sbjct: 16 CYNCGESGHMARECTSGGGGGGGRYGGGGGGGGGGGGGGSCYSCGESGHFARDCPTG 186
>TC80683 homologue to GP|8777424|dbj|BAA97014.1
gb|AAF56406.1~gene_id:K9P8.7~strong similarity to
unknown protein {Arabidopsis thaliana}, partial (16%)
Length = 1360
Score = 30.8 bits (68), Expect = 2.9
Identities = 22/81 (27%), Positives = 38/81 (46%), Gaps = 6/81 (7%)
Frame = +1
Query: 105 RRGKGQLSRPKPYSAPPDKGK*RLK-----DERRPKMRDAPTDIVCFKCGEKGHKSNVCD 159
R KG+L + K A D+ + ++ +P ++ ++ + +KG KS+
Sbjct: 886 RGQKGKLKKMKEKYADQDEEERSIRMSLLASSGKPIKKEETLPVI--ETSDKGKKSDSGP 1059
Query: 160 RDEKK-CFRCGKKGHTLADCK 179
D K C++C K GH DCK
Sbjct: 1060IDAPKICYKCKKVGHLSRDCK 1122
>TC81797 similar to GP|17065224|gb|AAL32766.1 Unknown protein {Arabidopsis
thaliana}, partial (17%)
Length = 758
Score = 30.4 bits (67), Expect = 3.8
Identities = 17/43 (39%), Positives = 29/43 (66%)
Frame = -2
Query: 233 IVLL*LLL*ILVLLIVSLLLIVHISWVWLYLI*KDKWLLKLQL 275
+VL+ +++ +L+LL++ LLL+ W+WL+L WLL L L
Sbjct: 493 LVLMAIMMFLLLLLLLVLLLL----WLWLWL-----WLLLLML 392
>TC83030 weakly similar to GP|18855061|gb|AAL79753.1 putative RNA helicase
{Oryza sativa}, partial (7%)
Length = 624
Score = 30.4 bits (67), Expect = 3.8
Identities = 12/20 (60%), Positives = 14/20 (70%), Gaps = 2/20 (10%)
Frame = +1
Query: 165 CFRCGKKGHTLADC--KRGD 182
CF CG+ GH +DC KRGD
Sbjct: 166 CFTCGESGHRASDCPNKRGD 225
Database: MTGI
Posted date: Oct 22, 2004 3:39 PM
Number of letters in database: 27,044,181
Number of sequences in database: 36,976
Lambda K H
0.369 0.167 0.628
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 36,590,598
Number of Sequences: 36976
Number of extensions: 599713
Number of successful extensions: 7899
Number of sequences better than 10.0: 27
Number of HSP's better than 10.0 without gapping: 1701
Number of HSP's successfully gapped in prelim test: 293
Number of HSP's that attempted gapping in prelim test: 5861
Number of HSP's gapped (non-prelim): 2411
length of query: 1069
length of database: 9,014,727
effective HSP length: 106
effective length of query: 963
effective length of database: 5,095,271
effective search space: 4906745973
effective search space used: 4906745973
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 14 ( 7.5 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 36 (21.7 bits)
S2: 63 (28.9 bits)
Medicago: description of AC141111.4


