

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC141111.3 + phase: 0 /pseudo
(64 letters)
Database: MTGI
36,976 sequences; 27,044,181 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
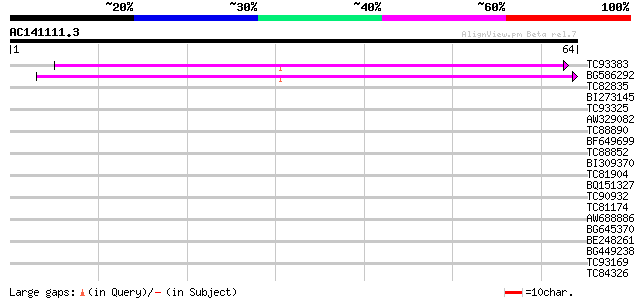
TC93383 similar to PIR|T47786|T47786 hypothetical protein F17J16... 43 3e-05
BG586292 similar to PIR|T46163|T46 nodulin / glutamate-ammonia l... 39 5e-04
TC82835 weakly similar to GP|21741712|emb|CAD41335. oj991113_30.... 36 0.003
BI273145 weakly similar to PIR|A84474|A84 hypothetical protein A... 35 0.004
TC93325 similar to GP|20268762|gb|AAM14084.1 putative salt-induc... 34 0.013
AW329082 weakly similar to PIR|F86151|F86 hypothetical protein A... 34 0.013
TC88890 similar to PIR|T47786|T47786 hypothetical protein F17J16... 33 0.017
BF649699 similar to GP|9757911|dbj gb|AAF19552.1~gene_id:MMN10.1... 33 0.022
TC88852 weakly similar to GP|22773243|gb|AAN06849.1 Hypothetical... 32 0.037
BI309370 weakly similar to PIR|T02656|T026 probable salt-inducib... 31 0.11
TC81904 weakly similar to GP|8778410|gb|AAF79418.1| F16A14.3 {Ar... 31 0.11
BQ151327 30 0.24
TC90932 similar to PIR|A86237|A86237 protein F14N23.15 [imported... 29 0.41
TC81174 29 0.41
AW688886 similar to GP|3258568|gb|A Unknown protein {Arabidopsis... 28 0.70
BG645370 weakly similar to PIR|T00405|T004 hypothetical protein ... 28 0.91
BE248261 weakly similar to GP|23616943|dbj contains EST AU069907... 27 1.2
BG449238 weakly similar to GP|19071826|dbj PnC401 homologue {Ara... 27 1.6
TC93169 weakly similar to GP|9294448|dbj|BAB02667.1 gb|AAF26996.... 27 1.6
TC84326 weakly similar to PIR|F85225|F85225 hypothetical protein... 27 1.6
>TC93383 similar to PIR|T47786|T47786 hypothetical protein F17J16.90 -
Arabidopsis thaliana, partial (43%)
Length = 1038
Score = 42.7 bits (99), Expect = 3e-05
Identities = 22/59 (37%), Positives = 35/59 (59%), Gaps = 1/59 (1%)
Frame = +1
Query: 6 YNTMITGYGKASNMDGAEGVFLTL-GGRIEPDETSYRSMIEGWGRAGNYEKARWYYEEL 63
Y TM++ Y A +M+GAE F L EP+ +Y ++I+G+ +A + EK YEE+
Sbjct: 319 YTTMLSAYVNAPDMEGAEKFFKRLIQDGFEPNVVTYGTLIKGYAKANDIEKVMEKYEEM 495
Score = 26.9 bits (58), Expect = 1.6
Identities = 20/64 (31%), Positives = 32/64 (49%), Gaps = 4/64 (6%)
Frame = +1
Query: 4 VVYNTMI---TGYGKASNM-DGAEGVFLTLGGRIEPDETSYRSMIEGWGRAGNYEKARWY 59
V YN+++ T Y + SN+ D + L PD SY +I +G+A E+A
Sbjct: 7 VTYNSLMSFETNYKEVSNIYDQMQRADL------RPDVVSYALLINAYGKARREEEALAV 168
Query: 60 YEEL 63
+EE+
Sbjct: 169FEEM 180
>BG586292 similar to PIR|T46163|T46 nodulin / glutamate-ammonia ligase-like
protein - Arabidopsis thaliana, partial (50%)
Length = 719
Score = 38.5 bits (88), Expect = 5e-04
Identities = 21/63 (33%), Positives = 30/63 (47%), Gaps = 2/63 (3%)
Frame = -3
Query: 4 VVYNTMITGYGKASNMDGAEGVFLTL--GGRIEPDETSYRSMIEGWGRAGNYEKARWYYE 61
V YNT++ GYGKA + E V + G PD + S+I +G GN K +Y
Sbjct: 405 VTYNTILDGYGKAGLFEEMENVLADMIEDGDSLPDVFTLNSIIGSYGNGGNIRKMESWYN 226
Query: 62 ELK 64
+
Sbjct: 225 RFQ 217
Score = 32.0 bits (71), Expect = 0.048
Identities = 18/56 (32%), Positives = 30/56 (53%), Gaps = 3/56 (5%)
Frame = -3
Query: 3 IVVYNTMITGYGKASN---MDGAEGVFLTLGGRIEPDETSYRSMIEGWGRAGNYEK 55
+ N++I YG N M+ F +G +EPD T++ +I +G+AG Y+K
Sbjct: 300 VFTLNSIIGSYGNGGNIRKMESWYNRFQLMG--VEPDITTFNVLILSFGKAGMYKK 139
>TC82835 weakly similar to GP|21741712|emb|CAD41335. oj991113_30.17 {Oryza
sativa}, partial (5%)
Length = 915
Score = 36.2 bits (82), Expect = 0.003
Identities = 21/62 (33%), Positives = 33/62 (52%), Gaps = 1/62 (1%)
Frame = +3
Query: 4 VVYNTMITGYGKASNMDGAEGVFLTLGGR-IEPDETSYRSMIEGWGRAGNYEKARWYYEE 62
+ YNT+I G K D F + EPD +Y ++I+ GRAGN E++ ++ E
Sbjct: 222 ISYNTLINGLRKVGRSDMCFEYFKEMKENGNEPDLLTYTALIDISGRAGNIEESLKFFME 401
Query: 63 LK 64
+K
Sbjct: 402 MK 407
>BI273145 weakly similar to PIR|A84474|A84 hypothetical protein At2g06000
[imported] - Arabidopsis thaliana, partial (28%)
Length = 466
Score = 35.4 bits (80), Expect = 0.004
Identities = 18/60 (30%), Positives = 32/60 (53%), Gaps = 1/60 (1%)
Frame = +2
Query: 5 VYNTMITGYGKASNMDGAEGVFLTLGGRIEPDETSYRSMIEGWGRAGN-YEKARWYYEEL 63
+YN +I GY K+ N+D A + + + + +PD+ ++ +I G G YE +Y L
Sbjct: 203 IYNPVIDGYCKSGNVDEANAIVVXMEKKCKPDKLTFTILIIGHCMKGRAYEAIGIFYRML 382
>TC93325 similar to GP|20268762|gb|AAM14084.1 putative salt-inducible
protein {Arabidopsis thaliana}, partial (31%)
Length = 929
Score = 33.9 bits (76), Expect = 0.013
Identities = 20/55 (36%), Positives = 29/55 (52%), Gaps = 1/55 (1%)
Frame = +3
Query: 5 VYNTMITGYGKASNMDGAEG-VFLTLGGRIEPDETSYRSMIEGWGRAGNYEKARW 58
+YN MI + + + D A G VF R +PD +Y ++I GRAG + RW
Sbjct: 450 IYNMMIRLHARHNTTDQARGLVFEMQKCRCKPDAETYNALINAHGRAGQW---RW 605
>AW329082 weakly similar to PIR|F86151|F86 hypothetical protein AAF76475.1
[imported] - Arabidopsis thaliana, partial (32%)
Length = 481
Score = 33.9 bits (76), Expect = 0.013
Identities = 20/62 (32%), Positives = 32/62 (51%), Gaps = 2/62 (3%)
Frame = +2
Query: 5 VYNTMITGYGKASNMDGAEGVF--LTLGGRIEPDETSYRSMIEGWGRAGNYEKARWYYEE 62
VY ++ Y N +GA+ VF + L G I PD+ +I + AG +KAR ++
Sbjct: 8 VYKALLRAYSVIGNAEGAQRVFDAIQLAGII-PDDKMCSLLIYAYSMAGQSQKARIAFDN 184
Query: 63 LK 64
+K
Sbjct: 185MK 190
>TC88890 similar to PIR|T47786|T47786 hypothetical protein F17J16.90 -
Arabidopsis thaliana, partial (23%)
Length = 749
Score = 33.5 bits (75), Expect = 0.017
Identities = 17/57 (29%), Positives = 29/57 (50%), Gaps = 1/57 (1%)
Frame = +1
Query: 9 MITGYGKASNMDGAEGVF-LTLGGRIEPDETSYRSMIEGWGRAGNYEKARWYYEELK 64
+IT YGK + +GAE V L P+ S +++E +G+ G Y A + ++
Sbjct: 496 LITAYGKLGDFNGAEKVLGLMNKNGYAPNVVSQTALMEAYGKGGRYNNAEAIFRRMQ 666
>BF649699 similar to GP|9757911|dbj gb|AAF19552.1~gene_id:MMN10.14~similar to
unknown protein {Arabidopsis thaliana}, partial (11%)
Length = 617
Score = 33.1 bits (74), Expect = 0.022
Identities = 18/62 (29%), Positives = 34/62 (54%), Gaps = 1/62 (1%)
Frame = +1
Query: 3 IVVYNTMITGYGKASNMDGAEGVFLTLGGR-IEPDETSYRSMIEGWGRAGNYEKARWYYE 61
I YN++I G+ K ++ AE ++ + +EP T++ ++I G+ + EKA Y
Sbjct: 289 IYPYNSLINGHCKFGDLSAAEFLYTKMINEGLEPTATTFTTLISGYCKDLQVEKAFKLYR 468
Query: 62 EL 63
E+
Sbjct: 469 EM 474
>TC88852 weakly similar to GP|22773243|gb|AAN06849.1 Hypothetical protein
{Oryza sativa (japonica cultivar-group)}, partial (36%)
Length = 1351
Score = 32.3 bits (72), Expect = 0.037
Identities = 18/60 (30%), Positives = 30/60 (50%), Gaps = 1/60 (1%)
Frame = +3
Query: 6 YNTMITGYGKASNMDGAEGVFLTLGGR-IEPDETSYRSMIEGWGRAGNYEKARWYYEELK 64
YN I+ + K+ N+D + + PD ++ S+I G + NYE A +EE+K
Sbjct: 519 YNNFISAFSKSGNVDAMLAWYSAKKATGLGPDLQTFESVISGCVNSKNYEIADRVFEEMK 698
>BI309370 weakly similar to PIR|T02656|T026 probable salt-inducible protein
[imported] - Arabidopsis thaliana, partial (8%)
Length = 637
Score = 30.8 bits (68), Expect = 0.11
Identities = 16/63 (25%), Positives = 34/63 (53%), Gaps = 1/63 (1%)
Frame = +1
Query: 3 IVVYNTMITGYGK-ASNMDGAEGVFLTLGGRIEPDETSYRSMIEGWGRAGNYEKARWYYE 61
+V YN + G+ + ++ + + + +EP+ T+++ +IEG AG E+A ++
Sbjct: 67 VVAYNVLAAGFFRNRTDFEAMDLLNYMESQGVEPNSTTHKIIIEGLCSAGKVEEAEEFFN 246
Query: 62 ELK 64
LK
Sbjct: 247 WLK 255
>TC81904 weakly similar to GP|8778410|gb|AAF79418.1| F16A14.3 {Arabidopsis
thaliana}, partial (7%)
Length = 978
Score = 30.8 bits (68), Expect = 0.11
Identities = 16/63 (25%), Positives = 34/63 (53%), Gaps = 1/63 (1%)
Frame = +1
Query: 3 IVVYNTMITGYGK-ASNMDGAEGVFLTLGGRIEPDETSYRSMIEGWGRAGNYEKARWYYE 61
+V YN + G+ + ++ + + + +EP+ T+++ +IEG AG E+A ++
Sbjct: 487 VVAYNVLAAGFFRNRTDFEAMDLLNYMESQGVEPNSTTHKIIIEGLCSAGKVEEAEEFFN 666
Query: 62 ELK 64
LK
Sbjct: 667 WLK 675
>BQ151327
Length = 774
Score = 29.6 bits (65), Expect = 0.24
Identities = 12/22 (54%), Positives = 14/22 (63%)
Frame = -3
Query: 43 MIEGWGRAGNYEKARWYYEELK 64
+I GWGRAG E WY EL+
Sbjct: 679 IIMGWGRAGIVEYVYWYIAELR 614
>TC90932 similar to PIR|A86237|A86237 protein F14N23.15 [imported] -
Arabidopsis thaliana, partial (36%)
Length = 1095
Score = 28.9 bits (63), Expect = 0.41
Identities = 12/30 (40%), Positives = 18/30 (60%)
Frame = +1
Query: 35 PDETSYRSMIEGWGRAGNYEKARWYYEELK 64
PD Y ++I G+ GN +KA ++ELK
Sbjct: 655 PDSMVYNNLISGFLELGNLDKANELFDELK 744
Score = 27.3 bits (59), Expect = 1.2
Identities = 12/29 (41%), Positives = 19/29 (65%)
Frame = +1
Query: 4 VVYNTMITGYGKASNMDGAEGVFLTLGGR 32
+VYN +I+G+ + N+D A +F L GR
Sbjct: 664 MVYNNLISGFLELGNLDKANELFDELKGR 750
>TC81174
Length = 1058
Score = 28.9 bits (63), Expect = 0.41
Identities = 18/65 (27%), Positives = 33/65 (50%), Gaps = 1/65 (1%)
Frame = +1
Query: 1 MCIVVYNTMITGYGKASNMDGAEGVFLTLGGR-IEPDETSYRSMIEGWGRAGNYEKARWY 59
+ + +Y+ I GK ++ AE +F + + ++PD + MI + + GN EKA
Sbjct: 373 LLLPLYSDTILLLGKNKMIEKAEEMFYEVVEKGLKPDTRLFNEMIGVYLQVGNTEKAMEV 552
Query: 60 YEELK 64
Y +K
Sbjct: 553 YRSMK 567
>AW688886 similar to GP|3258568|gb|A Unknown protein {Arabidopsis thaliana},
partial (11%)
Length = 660
Score = 28.1 bits (61), Expect = 0.70
Identities = 17/56 (30%), Positives = 27/56 (47%), Gaps = 1/56 (1%)
Frame = +2
Query: 6 YNTMITGYGKASNMDGAEGVFLTLGG-RIEPDETSYRSMIEGWGRAGNYEKARWYY 60
+NT+I + A N+D A VF + + D SY +I + G+Y KA +
Sbjct: 383 FNTLINSHCCAGNLDEAFKVFENMKKLEVSADSASYSVLIRTLCQKGDYGKAEMLF 550
>BG645370 weakly similar to PIR|T00405|T004 hypothetical protein At2g44880
[imported] - Arabidopsis thaliana, partial (11%)
Length = 784
Score = 27.7 bits (60), Expect = 0.91
Identities = 18/58 (31%), Positives = 24/58 (41%)
Frame = -2
Query: 6 YNTMITGYGKASNMDGAEGVFLTLGGRIEPDETSYRSMIEGWGRAGNYEKARWYYEEL 63
+N MIT YG + PD Y +I+ + R GN KAR EE+
Sbjct: 723 FNKMITNYG------------------LSPDIDIYACLIDLYARNGNLRKARDLMEEM 604
>BE248261 weakly similar to GP|23616943|dbj contains EST
AU069907(E11940)~similar to salt-inducible protein,
partial (11%)
Length = 282
Score = 27.3 bits (59), Expect = 1.2
Identities = 11/30 (36%), Positives = 19/30 (62%)
Frame = +2
Query: 35 PDETSYRSMIEGWGRAGNYEKARWYYEELK 64
PD SY S++ +GR+ +KAR ++ +K
Sbjct: 89 PDVVSYTSLLNAYGRSRKPQKAREIFKMIK 178
>BG449238 weakly similar to GP|19071826|dbj PnC401 homologue {Arabidopsis
thaliana}, partial (11%)
Length = 617
Score = 26.9 bits (58), Expect = 1.6
Identities = 12/49 (24%), Positives = 25/49 (50%), Gaps = 1/49 (2%)
Frame = +3
Query: 4 VVYNTMITGYGKASNMDGAEGVFLTL-GGRIEPDETSYRSMIEGWGRAG 51
+ YN +I + ++ NM+ A+ + P +Y ++I+G+G G
Sbjct: 471 IFYNAVINAFAESGNMEDAKKTVXKMKESGFRPSTGTYSNLIKGYGIEG 617
>TC93169 weakly similar to GP|9294448|dbj|BAB02667.1
gb|AAF26996.1~gene_id:MSL1.5~similar to unknown protein
{Arabidopsis thaliana}, partial (18%)
Length = 632
Score = 26.9 bits (58), Expect = 1.6
Identities = 15/59 (25%), Positives = 29/59 (48%), Gaps = 1/59 (1%)
Frame = +3
Query: 6 YNTMITGYGKASNMDGAEGVFLTL-GGRIEPDETSYRSMIEGWGRAGNYEKARWYYEEL 63
+N ++ G + A +F + I+PD SY +++ RAG +E+A +E+
Sbjct: 105 HNIILNGLARTGGPKRAMEMFAKMKSSTIKPDAVSYNTVLGCLSRAGLFEEATKLMKEM 281
>TC84326 weakly similar to PIR|F85225|F85225 hypothetical protein AT4g19900
[imported] - Arabidopsis thaliana, partial (18%)
Length = 756
Score = 26.9 bits (58), Expect = 1.6
Identities = 17/60 (28%), Positives = 33/60 (54%), Gaps = 3/60 (5%)
Frame = +1
Query: 6 YNTMITGYGKASNMDGAEGVF---LTLGGRIEPDETSYRSMIEGWGRAGNYEKARWYYEE 62
YN +++ + K N+ A +F L +G I+PD SY ++I + R +++ ++EE
Sbjct: 316 YNILMSEHCKQENIRQALALFNKMLKIG--IQPDIHSYTTLIAVFCRENRMKESEMFFEE 489
Database: MTGI
Posted date: Oct 22, 2004 3:39 PM
Number of letters in database: 27,044,181
Number of sequences in database: 36,976
Lambda K H
0.317 0.137 0.428
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 1,832,919
Number of Sequences: 36976
Number of extensions: 15348
Number of successful extensions: 112
Number of sequences better than 10.0: 48
Number of HSP's better than 10.0 without gapping: 105
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 111
length of query: 64
length of database: 9,014,727
effective HSP length: 40
effective length of query: 24
effective length of database: 7,535,687
effective search space: 180856488
effective search space used: 180856488
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.3 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.6 bits)
S2: 51 (24.3 bits)
Medicago: description of AC141111.3


