

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC140913.9 - phase: 2 /pseudo
(273 letters)
Database: MTGI
36,976 sequences; 27,044,181 total letters
Searching..................................................done
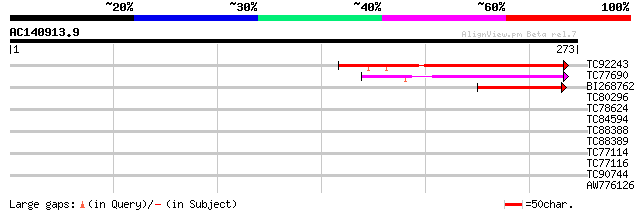
Score E
Sequences producing significant alignments: (bits) Value
TC92243 similar to PIR|T06661|T06661 hypothetical protein T6G15.... 87 5e-18
TC77690 similar to PIR|E86385|E86385 probable transmembrane prot... 63 1e-10
BI268762 similar to GP|15450794|gb| transmembrane protein FT27/P... 42 3e-04
TC80296 similar to PIR|H86265|H86265 protein F3F19.18 [imported]... 31 0.45
TC78624 similar to GP|4850385|gb|AAD31055.1| EST gb|T22808 comes... 29 1.7
TC84594 similar to GP|23497092|gb|AAN36641.1 hypothetical protei... 29 2.2
TC88388 similar to PIR|T04590|T04590 hypothetical protein F23E13... 28 2.9
TC88389 similar to PIR|T04590|T04590 hypothetical protein F23E13... 28 2.9
TC77114 similar to GP|6686786|emb|CAB64720.1 AKIN gamma {Arabido... 28 3.8
TC77116 similar to GP|9294182|dbj|BAB02084.1 protein kinases-lik... 28 3.8
TC90744 similar to GP|20147217|gb|AAM10324.1 AT5g19390/F7K24_140... 28 4.9
AW776126 similar to GP|12643044|gb| putative AT-Hook DNA-binding... 28 4.9
>TC92243 similar to PIR|T06661|T06661 hypothetical protein T6G15.140 -
Arabidopsis thaliana, partial (43%)
Length = 723
Score = 87.4 bits (215), Expect = 5e-18
Identities = 53/120 (44%), Positives = 74/120 (61%), Gaps = 9/120 (7%)
Frame = +3
Query: 159 LLVYFGVSTLLDA---SSSDSQKSD------DEQKEAELAVSDFSGDGAGILAAASTIVS 209
LL++FG+ ++ DA S D + D DE EAE V + + + I
Sbjct: 3 LLLFFGLKSIKDAWDLPSKDVKNGDNSSPELDELAEAEELVKEKASQR--LSNPLEIIWK 176
Query: 210 TFLLVFVAEWGDKSFFSTIALAAASSPLGVIAGSLAGHGVATLIAVLGGSLLGTFLSEKV 269
+F LVF AEWGD+S +TIALAAA SP GV +G++AGH +AT IA++GGSLL ++SEK+
Sbjct: 177 SFSLVFFAEWGDRSMLATIALAAAQSPWGVASGAIAGHLLATCIAIVGGSLLANYISEKL 356
>TC77690 similar to PIR|E86385|E86385 probable transmembrane protein
[imported] - Arabidopsis thaliana, partial (95%)
Length = 1215
Score = 63.2 bits (152), Expect = 1e-10
Identities = 36/101 (35%), Positives = 55/101 (53%), Gaps = 1/101 (0%)
Frame = +1
Query: 170 DASSSDSQKSDDEQKEAELA-VSDFSGDGAGILAAASTIVSTFLLVFVAEWGDKSFFSTI 228
+ ++ DS+K DD K+ + +S F + ++ F + F EWGDKS +TI
Sbjct: 475 NGATKDSKKVDDATKKHKRPFLSQFF---------SPILLQAFSITFFGEWGDKSQLATI 627
Query: 229 ALAAASSPLGVIAGSLAGHGVATLIAVLGGSLLGTFLSEKV 269
LAA +P GV+ G + G + T AV+GG L + +SEKV
Sbjct: 628 GLAADENPFGVVLGGILGQALCTTAAVIGGKSLASQISEKV 750
Score = 31.2 bits (69), Expect = 0.45
Identities = 21/77 (27%), Positives = 36/77 (46%)
Frame = +1
Query: 182 EQKEAELAVSDFSGDGAGILAAASTIVSTFLLVFVAEWGDKSFFSTIALAAASSPLGVIA 241
E+ L++S A + + + + ++E GDK+FF+ LA V+
Sbjct: 79 EETTTTLSLSLSLSLSANMSSIVQGFTKSLAMTVLSEIGDKTFFAAAILAMRHPRRLVLT 258
Query: 242 GSLAGHGVATLIAVLGG 258
G LA V T+++VL G
Sbjct: 259 GCLAALIVMTILSVLVG 309
>BI268762 similar to GP|15450794|gb| transmembrane protein FT27/PFT27-like
{Arabidopsis thaliana}, partial (21%)
Length = 210
Score = 41.6 bits (96), Expect = 3e-04
Identities = 19/43 (44%), Positives = 30/43 (69%)
Frame = +2
Query: 226 STIALAAASSPLGVIAGSLAGHGVATLIAVLGGSLLGTFLSEK 268
+TIALA + +GV AG+ GH + T AV+GGS+L + +S++
Sbjct: 8 ATIALATHKNAIGVAAGATIGHTICTSXAVVGGSMLASRISQR 136
>TC80296 similar to PIR|H86265|H86265 protein F3F19.18 [imported] -
Arabidopsis thaliana, partial (34%)
Length = 1734
Score = 31.2 bits (69), Expect = 0.45
Identities = 15/30 (50%), Positives = 17/30 (56%)
Frame = +3
Query: 8 GFVRASEDDSVAENSDSQCQLATQNVSSDD 37
G A EDD V+ NSD QL + N SDD
Sbjct: 798 GSDNAQEDDQVSLNSDDDNQLGSDNTGSDD 887
>TC78624 similar to GP|4850385|gb|AAD31055.1| EST gb|T22808 comes from this
gene. {Arabidopsis thaliana}, partial (51%)
Length = 2052
Score = 29.3 bits (64), Expect = 1.7
Identities = 21/59 (35%), Positives = 29/59 (48%), Gaps = 2/59 (3%)
Frame = -2
Query: 212 LLVFVAEWGDKSFFSTIALAAASSPLGVIAGSLAGHGVATLIAVLGGSL--LGTFLSEK 268
LLVF A + SFF + +A +S LG+ A +G+ + G SL L FL K
Sbjct: 1091 LLVFFASISESSFFQSYPFSANTSSLGL----AAPNGLGLFVFTTGDSLTELSPFLGSK 927
>TC84594 similar to GP|23497092|gb|AAN36641.1 hypothetical protein
{Plasmodium falciparum 3D7}, partial (0%)
Length = 791
Score = 28.9 bits (63), Expect = 2.2
Identities = 12/37 (32%), Positives = 20/37 (53%), Gaps = 3/37 (8%)
Frame = -1
Query: 77 GDVGDLSTGFA---SYHRHFC*YFSLNWETRLFSLQH 110
G++ + GF+ YH ++C*+ LNW + QH
Sbjct: 611 GNISGMKLGFSIL*HYHHYYC*HQ*LNWNCACYRHQH 501
>TC88388 similar to PIR|T04590|T04590 hypothetical protein F23E13.100 -
Arabidopsis thaliana, partial (20%)
Length = 930
Score = 28.5 bits (62), Expect = 2.9
Identities = 17/41 (41%), Positives = 23/41 (55%)
Frame = +3
Query: 231 AAASSPLGVIAGSLAGHGVATLIAVLGGSLLGTFLSEKVTS 271
AAA+S G +AGS+A VA G L GT ++ +V S
Sbjct: 63 AAAASAAGTVAGSVA---VAASFGAAGAGLTGTKMARRVGS 176
>TC88389 similar to PIR|T04590|T04590 hypothetical protein F23E13.100 -
Arabidopsis thaliana, partial (37%)
Length = 630
Score = 28.5 bits (62), Expect = 2.9
Identities = 17/41 (41%), Positives = 23/41 (55%)
Frame = +2
Query: 231 AAASSPLGVIAGSLAGHGVATLIAVLGGSLLGTFLSEKVTS 271
AAA+S G +AGS+A VA G L GT ++ +V S
Sbjct: 200 AAAASAAGTVAGSVA---VAASFGAAGAGLTGTKMARRVGS 313
>TC77114 similar to GP|6686786|emb|CAB64720.1 AKIN gamma {Arabidopsis
thaliana}, partial (85%)
Length = 1417
Score = 28.1 bits (61), Expect = 3.8
Identities = 23/70 (32%), Positives = 34/70 (47%), Gaps = 2/70 (2%)
Frame = -2
Query: 198 AGILAAASTIVSTFLLV--FVAEWGDKSFFSTIALAAASSPLGVIAGSLAGHGVATLIAV 255
A I A++ + S F+ + F+A G K+ ++ A S LG+ S H +TLI
Sbjct: 249 ASIFTASAKVASDFISISWFLAGKG-KAETGMLSKQALSLSLGLNCAS*TSHRSSTLIPS 73
Query: 256 LGGSLLGTFL 265
L LLG L
Sbjct: 72 LASGLLGISL 43
>TC77116 similar to GP|9294182|dbj|BAB02084.1 protein kinases-like protein
{Arabidopsis thaliana}, partial (28%)
Length = 724
Score = 28.1 bits (61), Expect = 3.8
Identities = 23/70 (32%), Positives = 34/70 (47%), Gaps = 2/70 (2%)
Frame = -2
Query: 198 AGILAAASTIVSTFLLV--FVAEWGDKSFFSTIALAAASSPLGVIAGSLAGHGVATLIAV 255
A I A++ + S F+ + F+A G K+ ++ A S LG+ S H +TLI
Sbjct: 231 ASIFTASAKVASDFISISWFLAGKG-KAETGMLSKQALSLSLGLNCAS*TSHRSSTLIPS 55
Query: 256 LGGSLLGTFL 265
L LLG L
Sbjct: 54 LASGLLGISL 25
>TC90744 similar to GP|20147217|gb|AAM10324.1 AT5g19390/F7K24_140
{Arabidopsis thaliana}, partial (22%)
Length = 968
Score = 27.7 bits (60), Expect = 4.9
Identities = 13/33 (39%), Positives = 20/33 (60%)
Frame = -1
Query: 149 LPIDDIAAVCLLVYFGVSTLLDASSSDSQKSDD 181
LP+ A +C +F +S+L ASSS ++S D
Sbjct: 308 LPLASFAILCCKSFFAISSL*IASSSSPEESTD 210
>AW776126 similar to GP|12643044|gb| putative AT-Hook DNA-binding protein
{Oryza sativa}, partial (19%)
Length = 742
Score = 27.7 bits (60), Expect = 4.9
Identities = 17/60 (28%), Positives = 27/60 (44%)
Frame = +2
Query: 28 LATQNVSSDDASMTVLKFMLFSAFFALQDAFPAVAASDFATGLNSIPIFGDVGDLSTGFA 87
L+T +S F+ + F + + FP+ AA DFA + ++ DVG FA
Sbjct: 332 LSTNEFASKKGRGKSTGFVNYQTFSSFGEVFPSTAAVDFAPHVVTVYAGEDVGGKILSFA 511
Database: MTGI
Posted date: Oct 22, 2004 3:39 PM
Number of letters in database: 27,044,181
Number of sequences in database: 36,976
Lambda K H
0.325 0.138 0.401
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 7,125,439
Number of Sequences: 36976
Number of extensions: 91999
Number of successful extensions: 537
Number of sequences better than 10.0: 24
Number of HSP's better than 10.0 without gapping: 531
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 537
length of query: 273
length of database: 9,014,727
effective HSP length: 95
effective length of query: 178
effective length of database: 5,502,007
effective search space: 979357246
effective search space used: 979357246
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 15 ( 7.0 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 40 (21.6 bits)
S2: 57 (26.6 bits)
Medicago: description of AC140913.9


