

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC140850.4 - phase: 0 /pseudo
(1010 letters)
Database: MTGI
36,976 sequences; 27,044,181 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
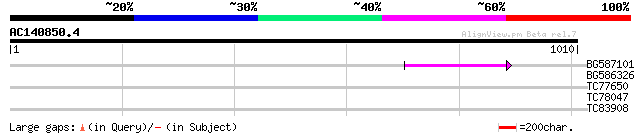
BG587101 similar to GP|6691191|gb F7F22.15 {Arabidopsis thaliana... 63 6e-10
BG586326 similar to PIR|G84493|G8 probable retroelement pol poly... 35 0.19
TC77650 similar to GP|9795587|gb|AAF98405.1| Unknown protein {Ar... 30 4.7
TC78047 similar to PIR|T02706|T02706 hypothetical protein At2g03... 30 6.1
TC83908 similar to PIR|T47792|T47792 hypothetical protein F17J16... 29 8.0
>BG587101 similar to GP|6691191|gb F7F22.15 {Arabidopsis thaliana}, partial
(10%)
Length = 624
Score = 62.8 bits (151), Expect = 6e-10
Identities = 57/191 (29%), Positives = 87/191 (44%)
Frame = +3
Query: 703 GMNHTCLKLERITSCVGVLIMRRQRKYCGNVTIPPMEVTSTGRELLQKYSNPVSIGQNFS 762
GM+H+C+ I GVL+ +R + Y +P + ++ + VS G S
Sbjct: 27 GMSHSCISSVPIIFT*GVLLKKRYQAYSSTAMVPTTRDILL*AKPFRRSNKLVSGGLRCS 206
Query: 763 KMLTLIASFVMNVKEPDQCQSGMKCLYKVFLKWKCLIVGVLILLAPSHHRTQMNTSW*P* 822
KM T + V++ ++ + GM+C K + L G I SH T ++TS
Sbjct: 207 KMPTPSSLSVIHARDREISARGMRCHRTSSWKLRYLTCGGSISWDHSHPLTTISTSLLLL 386
Query: 823 IMSQSGWKPSRRQKPTARQW*SSSKRISSPG*EHQEF**VMEAHVSATHSFLRH*IIMV* 882
SG +P + Q+ T + W* S S QE+* VME H+S+T * M *
Sbjct: 387 TTFPSG*RP*QVQQTTPQWW*RCSNLSSFQDLGCQEW*LVMEDHISSTKCSRSC*RRMA* 566
Query: 883 NTR*HHHTILK 893
+TR HT L+
Sbjct: 567 DTRWRQHTTLR 599
>BG586326 similar to PIR|G84493|G8 probable retroelement pol polyprotein
[imported] - Arabidopsis thaliana, partial (13%)
Length = 736
Score = 34.7 bits (78), Expect = 0.19
Identities = 20/49 (40%), Positives = 25/49 (50%)
Frame = -2
Query: 3 NNIHRMSLGFESDLVRRYLIAAWSV*TTILEPTRYERNFSNAYTIASNS 51
N+IH + L F S V+ Y WSV L P +YER A T A+ S
Sbjct: 318 NSIHLLCLKFNSGWVKIYFRLLWSVCI*TLAPYKYERQIFKANTTAAIS 172
>TC77650 similar to GP|9795587|gb|AAF98405.1| Unknown protein {Arabidopsis
thaliana}, partial (91%)
Length = 1530
Score = 30.0 bits (66), Expect = 4.7
Identities = 19/70 (27%), Positives = 33/70 (47%)
Frame = -2
Query: 687 IGNNARNFSMKLFTMYGMNHTCLKLERITSCVGVLIMRRQRKYCGNVTIPPMEVTSTGRE 746
IGNN N + + + G + TC+ L V RQR + +P ++ +S+ +
Sbjct: 1340 IGNNVGNNHLHMPSGVGRSFTCICLYIHCQFSDVTSPPRQRMTDPPLHVPTLQHSSSDKP 1161
Query: 747 LLQKYSNPVS 756
L Y NP++
Sbjct: 1160 LTTPYKNPLA 1131
>TC78047 similar to PIR|T02706|T02706 hypothetical protein At2g03200
[imported] - Arabidopsis thaliana, partial (67%)
Length = 1727
Score = 29.6 bits (65), Expect = 6.1
Identities = 13/43 (30%), Positives = 22/43 (50%)
Frame = -1
Query: 682 FLKIIIGNNARNFSMKLFTMYGMNHTCLKLERITSCVGVLIMR 724
+LK ++ N + + + NH C KLE++ C+G MR
Sbjct: 818 YLKYLVS*NNHHMNK-----HNCNHCCCKLEKVKHCIGCYYMR 705
>TC83908 similar to PIR|T47792|T47792 hypothetical protein F17J16.150 -
Arabidopsis thaliana, partial (12%)
Length = 841
Score = 29.3 bits (64), Expect = 8.0
Identities = 15/43 (34%), Positives = 22/43 (50%)
Frame = +1
Query: 569 NARTKFSMLYIMRVKF*MKNKLIMLPLKKSCSQ*YMHWKNFVH 611
N +TK LY++ K + I + K S+* M+WKN H
Sbjct: 535 NLKTKIMPLYLLAEKLCRTST*IRIITMKKLSK*EMYWKNSTH 663
Database: MTGI
Posted date: Oct 22, 2004 3:39 PM
Number of letters in database: 27,044,181
Number of sequences in database: 36,976
Lambda K H
0.358 0.155 0.550
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 33,802,731
Number of Sequences: 36976
Number of extensions: 499429
Number of successful extensions: 5104
Number of sequences better than 10.0: 10
Number of HSP's better than 10.0 without gapping: 1506
Number of HSP's successfully gapped in prelim test: 205
Number of HSP's that attempted gapping in prelim test: 3532
Number of HSP's gapped (non-prelim): 1878
length of query: 1010
length of database: 9,014,727
effective HSP length: 106
effective length of query: 904
effective length of database: 5,095,271
effective search space: 4606124984
effective search space used: 4606124984
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 14 ( 7.2 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 37 (21.8 bits)
S2: 63 (28.9 bits)
Medicago: description of AC140850.4


