

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC140850.3 + phase: 0 /pseudo
(650 letters)
Database: MTGI
36,976 sequences; 27,044,181 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
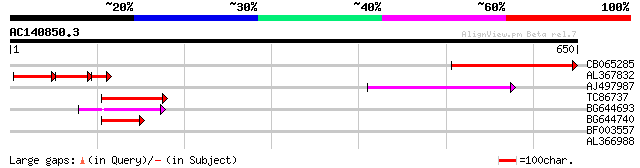
CB065285 weakly similar to PIR|A84500|A84 probable retroelement ... 218 4e-57
AL367832 weakly similar to GP|14091845|gb Putative retroelement ... 101 2e-50
AJ497987 weakly similar to GP|9927273|dbj Similar to Arabidopsis... 69 7e-12
TC86737 weakly similar to GP|6683624|dbj|BAA89272.1 Pol {Alterna... 56 5e-08
BG644693 weakly similar to GP|18767374|g Putative 22 kDa kafirin... 54 2e-07
BG644740 similar to PIR|A84460|A84 probable retroelement pol pol... 49 8e-06
BF003557 weakly similar to GP|21554257|gb| unknown {Arabidopsis ... 32 1.00
AL366988 30 2.9
>CB065285 weakly similar to PIR|A84500|A84 probable retroelement gag/pol
polyprotein [imported] - Arabidopsis thaliana, partial
(2%)
Length = 592
Score = 218 bits (556), Expect = 4e-57
Identities = 119/144 (82%), Positives = 123/144 (84%)
Frame = -3
Query: 507 NQKTAKNR*LVKVWIPIASGD*SSMVLSMLMVKESGQSLYPRRGITSLLPPGFCSNVQTI 566
NQKTAKNR*LVKV IPIA+G *S MVLSMLMVKE QSLYPRRGITSLLPPGFCSNVQTI
Sbjct: 590 NQKTAKNR*LVKVQIPIANGV*SLMVLSMLMVKELEQSLYPRRGITSLLPPGFCSNVQTI 411
Query: 567 WPNTKRVSLGSMKQLT*ESNTSTSMEIRHLSSTR*RVNGRPTMPS*FLTVIMRDVC*HIL 626
W + K VSLGS KQLT*ESNTSTSMEI HLSSTR RVNGRPTMP *FLT MR VC* IL
Sbjct: 410 WLSMKHVSLGSRKQLT*ESNTSTSMEILHLSSTRSRVNGRPTMPI*FLTATMRGVC*PIL 231
Query: 627 QRSNCTISLGMRTKWLMLLLLYPP 650
Q+ +CTI L RTKWLML LLYPP
Sbjct: 230 QKLSCTIFLVTRTKWLMLWLLYPP 159
>AL367832 weakly similar to GP|14091845|gb Putative retroelement {Oryza
sativa}, partial (2%)
Length = 384
Score = 101 bits (252), Expect(3) = 2e-50
Identities = 47/49 (95%), Positives = 48/49 (97%)
Frame = -3
Query: 5 P*RD*AYQPRYRGEQARDQGRCCSRGRG*KEDLPAPPRIPGYFCMVV*R 53
P RD*AYQPRYRGEQARDQGRCCSRGRG*KEDLPAPPRIPGYFCM+V*R
Sbjct: 358 PGRD*AYQPRYRGEQARDQGRCCSRGRG*KEDLPAPPRIPGYFCMLV*R 212
Score = 86.3 bits (212), Expect(3) = 2e-50
Identities = 42/43 (97%), Positives = 43/43 (99%)
Frame = -2
Query: 53 RLWNIGSQQSLNVLPSGRN*EGLIQIWLSRLRVRFKSRLMRVF 95
+LWNIGSQQSLNVLPSGRN*EGLIQIWLSRLRVRFKSRLMRVF
Sbjct: 191 KLWNIGSQQSLNVLPSGRN*EGLIQIWLSRLRVRFKSRLMRVF 63
Score = 51.2 bits (121), Expect(3) = 2e-50
Identities = 22/22 (100%), Positives = 22/22 (100%)
Frame = -1
Query: 95 FLMTVEYPEWVANIVPVPKKDG 116
FLMTVEYPEWVANIVPVPKKDG
Sbjct: 66 FLMTVEYPEWVANIVPVPKKDG 1
>AJ497987 weakly similar to GP|9927273|dbj Similar to Arabidopsis thaliana
chromosome II BAC F26H6; putative retroelement pol
polyprotein, partial (1%)
Length = 636
Score = 68.6 bits (166), Expect = 7e-12
Identities = 64/169 (37%), Positives = 78/169 (45%)
Frame = -3
Query: 411 RLVVLWPGPPNACVIIW*IIQLG*YPEWIR*SISSRNLLLLGRLHAGKCSCPNMTLCSRL 470
R VVL PG P+A VII I LG Y +WI+ S R+ L LH G+C NM +
Sbjct: 634 RHVVL*PGLPSASVII*LITLLGWYLKWIQSSTYLRSPL*QEGLHGGRCYYRNMISSTVP 455
Query: 471 KKQSKVAFLPIILPTNLLMITNQLSLISPMKRSCI*NQKTAKNR*LVKVWIPIASGD*SS 530
++Q K FL I TN L + SL MKR CI* K + KV I I G *
Sbjct: 454 RRQLKAVFLLITWLTNHLKTIDLSSLTFLMKRLCI*R*KIVTSHYSEKVLIQIQCGV*YL 275
Query: 531 MVLSMLMVKESGQSLYPRRGITSLLPPGFCSNVQTIWPNTKRVSLGSMK 579
M LSM GQ + +TSL +T NTK S S +
Sbjct: 274 MGLSMYTATA*GQYSLLLKVLTSLSQQDSGLTARTTSLNTKPASWASTR 128
>TC86737 weakly similar to GP|6683624|dbj|BAA89272.1 Pol {Alternaria
alternata}, partial (21%)
Length = 1540
Score = 55.8 bits (133), Expect = 5e-08
Identities = 27/75 (36%), Positives = 46/75 (61%)
Frame = +1
Query: 106 ANIVPVPKKDGKVRMCVDFRDLNKASPKDNFPLPHIDVLVDNTAQSKVFSFMDGFSGYNL 165
A ++ V K G +R CVD+R LN + KD +PLP I + A ++ F+ +D + ++
Sbjct: 577 APVLFVRKPGGGIRFCVDYRALNAITKKDRYPLPLISETLRRVAGARWFTKLDVVAAFHK 756
Query: 166 IKMSPEDREKTSFIT 180
+++ ED+EKT+F T
Sbjct: 757 MRIKDEDQEKTAFRT 801
>BG644693 weakly similar to GP|18767374|g Putative 22 kDa kafirin cluster;
Ty3-Gypsy type {Oryza sativa}, partial (15%)
Length = 716
Score = 53.5 bits (127), Expect = 2e-07
Identities = 36/99 (36%), Positives = 54/99 (54%)
Frame = +2
Query: 80 LSRLRVRFKSRLMRVFLMTVEYPEWVANIVPVPKKDGKVRMCVDFRDLNKASPKDNFPLP 139
L L+++ K L + F+ YP V ++ + KKDG +RM +D+ LN + K +PLP
Sbjct: 110 LKVLKLQLKDLLEKGFIQPSIYP*GVV-VLFLKKKDGFLRMSIDYPQLNNVNIKIKYPLP 286
Query: 140 HIDVLVDNTAQSKVFSFMDGFSGYNLIKMSPEDREKTSF 178
ID L DN SK F +D G + ++ ED KT+F
Sbjct: 287 LIDELFDNLQGSKWFFKIDLRLG*HQHRVIGEDVPKTAF 403
>BG644740 similar to PIR|A84460|A84 probable retroelement pol polyprotein
[imported] - Arabidopsis thaliana, partial (4%)
Length = 754
Score = 48.5 bits (114), Expect = 8e-06
Identities = 22/49 (44%), Positives = 30/49 (60%)
Frame = -1
Query: 106 ANIVPVPKKDGKVRMCVDFRDLNKASPKDNFPLPHIDVLVDNTAQSKVF 154
A ++ V KKDG RMC+D+R NK + K+ +PLP ID L D + F
Sbjct: 238 AALLFVRKKDGYFRMCIDYRQFNKVTTKNKYPLPRIDNLFDKIQEDCYF 92
>BF003557 weakly similar to GP|21554257|gb| unknown {Arabidopsis thaliana},
partial (49%)
Length = 464
Score = 31.6 bits (70), Expect = 1.00
Identities = 24/63 (38%), Positives = 30/63 (47%), Gaps = 10/63 (15%)
Frame = -2
Query: 567 WPNTKRVSLGSMKQLT*ESN----------TSTSMEIRHLSSTR*RVNGRPTMPS*FLTV 616
WPN S + LT E+N TST ++IR + T R G PTM S FL +
Sbjct: 328 WPNNHAPSKLNDPGLTMETNPRTFS*KATVTSTDLQIREI*FTDFRRKGFPTMASIFLIL 149
Query: 617 IMR 619
I R
Sbjct: 148 IRR 140
>AL366988
Length = 405
Score = 30.0 bits (66), Expect = 2.9
Identities = 15/40 (37%), Positives = 22/40 (54%), Gaps = 1/40 (2%)
Frame = -3
Query: 343 IASRITCCNHLSLSHPWKEGL*LCICQCLMNLW-DAYLVN 381
I + I CC HL + HP+ + C+C + L AYL+N
Sbjct: 271 IIAYIRCCTHLDIKHPF-----ILRCRCYIELI*HAYLIN 167
Database: MTGI
Posted date: Oct 22, 2004 3:39 PM
Number of letters in database: 27,044,181
Number of sequences in database: 36,976
Lambda K H
0.347 0.151 0.514
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 21,744,537
Number of Sequences: 36976
Number of extensions: 327620
Number of successful extensions: 2849
Number of sequences better than 10.0: 16
Number of HSP's better than 10.0 without gapping: 1533
Number of HSP's successfully gapped in prelim test: 123
Number of HSP's that attempted gapping in prelim test: 1288
Number of HSP's gapped (non-prelim): 1718
length of query: 650
length of database: 9,014,727
effective HSP length: 102
effective length of query: 548
effective length of database: 5,243,175
effective search space: 2873259900
effective search space used: 2873259900
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 15 ( 7.5 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 38 (21.7 bits)
S2: 61 (28.1 bits)
Medicago: description of AC140850.3


