

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC140772.6 + phase: 0 /partial
(40 letters)
Database: MTGI
36,976 sequences; 27,044,181 total letters
Searching..................................................done
Score E
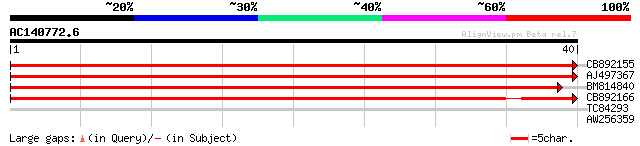
Sequences producing significant alignments: (bits) Value
CB892155 similar to PIR|D86481|D86 hypothetical protein AAG28292... 59 3e-10
AJ497367 similar to GP|14140286|gb putative helicase {Oryza sati... 54 1e-08
BM814840 weakly similar to GP|15128241|db helicase-like protein ... 47 2e-06
CB892166 similar to GP|20197614|gb unknown protein {Arabidopsis ... 44 2e-05
TC84293 similar to GP|23617010|dbj|BAC20706. contains ESTs D4714... 26 3.0
AW256359 25 6.6
>CB892155 similar to PIR|D86481|D86 hypothetical protein AAG28292.1
[imported] - Arabidopsis thaliana, partial (4%)
Length = 572
Score = 59.3 bits (142), Expect = 3e-10
Identities = 28/40 (70%), Positives = 34/40 (85%)
Frame = -2
Query: 1 LYVTISRVTSRGGLKILINDDDGDDTNVASSVIYREVFRN 40
LYV +SRVTSR GLKILI++DDG+D V S+V+YREVF N
Sbjct: 271 LYVALSRVTSREGLKILISNDDGEDDCVTSNVVYREVFHN 152
>AJ497367 similar to GP|14140286|gb putative helicase {Oryza sativa
(japonica cultivar-group)}, partial (1%)
Length = 543
Score = 54.3 bits (129), Expect = 1e-08
Identities = 25/40 (62%), Positives = 32/40 (79%)
Frame = +1
Query: 1 LYVTISRVTSRGGLKILINDDDGDDTNVASSVIYREVFRN 40
LYV +SRVTSR GLKIL+ +DG+ N S+V+Y+EVFRN
Sbjct: 34 LYVAVSRVTSRKGLKILLAHEDGNCMNTTSNVVYKEVFRN 153
>BM814840 weakly similar to GP|15128241|db helicase-like protein {Oryza
sativa (japonica cultivar-group)}, partial (6%)
Length = 733
Score = 47.0 bits (110), Expect = 2e-06
Identities = 21/39 (53%), Positives = 30/39 (76%)
Frame = +3
Query: 1 LYVTISRVTSRGGLKILINDDDGDDTNVASSVIYREVFR 39
LYV +SRV SR GLKILI D++G ++ +V+Y+EVF+
Sbjct: 285 LYVAVSRVKSRSGLKILITDENGSPSSSTVNVVYQEVFQ 401
>CB892166 similar to GP|20197614|gb unknown protein {Arabidopsis thaliana},
partial (3%)
Length = 748
Score = 43.5 bits (101), Expect = 2e-05
Identities = 22/40 (55%), Positives = 32/40 (80%)
Frame = -1
Query: 1 LYVTISRVTSRGGLKILINDDDGDDTNVASSVIYREVFRN 40
LYV ISRV+SR GLKIL+ D++GD + ++V+Y+ VF+N
Sbjct: 256 LYVAISRVSSRNGLKILMIDENGDCIDNTTNVVYK-VFQN 140
>TC84293 similar to GP|23617010|dbj|BAC20706. contains ESTs D47144(S12288)
AU032626(S12288)~similar to Arabidopsis thaliana
chromosome 5, partial (25%)
Length = 830
Score = 26.2 bits (56), Expect = 3.0
Identities = 10/20 (50%), Positives = 15/20 (75%)
Frame = +3
Query: 5 ISRVTSRGGLKILINDDDGD 24
+ TSRG LK+ +++DDGD
Sbjct: 141 VYNTTSRGILKVALSNDDGD 200
>AW256359
Length = 411
Score = 25.0 bits (53), Expect = 6.6
Identities = 10/23 (43%), Positives = 16/23 (69%)
Frame = -2
Query: 1 LYVTISRVTSRGGLKILINDDDG 23
+YV IS + GG ++ +ND+DG
Sbjct: 356 VYVFIS*INLLGGTELPVNDEDG 288
Database: MTGI
Posted date: Oct 22, 2004 3:39 PM
Number of letters in database: 27,044,181
Number of sequences in database: 36,976
Lambda K H
0.318 0.139 0.377
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 753,550
Number of Sequences: 36976
Number of extensions: 4009
Number of successful extensions: 35
Number of sequences better than 10.0: 12
Number of HSP's better than 10.0 without gapping: 35
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 35
length of query: 40
length of database: 9,014,727
effective HSP length: 16
effective length of query: 24
effective length of database: 8,423,111
effective search space: 202154664
effective search space used: 202154664
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.7 bits)
S2: 52 (24.6 bits)
Medicago: description of AC140772.6


