

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC140721.10 + phase: 0 /pseudo
(329 letters)
Database: MTGI
36,976 sequences; 27,044,181 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
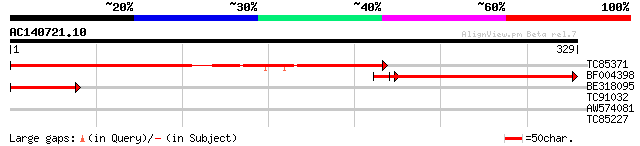
TC85371 similar to GP|15077144|gb|AAK83083.1 nuclear matrix prot... 252 1e-67
BF004398 similar to GP|21592356|gb nuclear matrix protein 1 {Ara... 216 4e-57
BE318095 similar to GP|21592356|gb| nuclear matrix protein 1 {Ar... 80 8e-16
TC91032 weakly similar to GP|20196995|gb|AAB91977.2 expressed pr... 29 2.2
AW574081 28 4.9
TC85227 similar to GP|15983773|gb|AAL10483.1 AT4g29060/F19B15_90... 28 6.4
>TC85371 similar to GP|15077144|gb|AAK83083.1 nuclear matrix protein 1
{Lycopersicon esculentum}, partial (66%)
Length = 738
Score = 252 bits (644), Expect = 1e-67
Identities = 151/232 (65%), Positives = 166/232 (71%), Gaps = 13/232 (5%)
Frame = +3
Query: 1 MASKQMEVIQKKLGMLNYPRANASAQSLLFAGMERYALFEWLFFRLLGDKSPFSQQNLQG 60
MAS QME IQKKLG+LNYPR+NASAQSLLFAGMERYAL EWLFFRLLGDKSPFSQQNLQG
Sbjct: 81 MASNQMEAIQKKLGILNYPRSNASAQSLLFAGMERYALLEWLFFRLLGDKSPFSQQNLQG 260
Query: 61 DALDRDEETARIQYLAEIAKFLGITTTVDTDAIQTHLLNDDLFFFL*LW*CRYGIGINLV 120
DALDRDEETARIQYLAE+AKFLGITTTVDTDAIQ H +D L L+
Sbjct: 261 DALDRDEETARIQYLAEMAKFLGITTTVDTDAIQGHGSYEDRTEML-----------RLI 407
Query: 121 CIFVSIVLTSR*PRTYT**ILLQKNRH----------KYFLKNVNCFP---QMFRFSPYI 167
V + + P ++ + K+ H + F + FP Q+ P +
Sbjct: 408 VDLVEATIYADNPE-WSVDEQVAKDIHLIDSIAEKQAQIFSEECKLFPADVQIQSIYP-L 581
Query: 168 P*VAELESKLTEQSKILLNLQQKVDDLASKHAYNPDEEYTEVESQLRAHLES 219
P VAELESKLTEQSKILLNLQQKVDDLASKHAYNPDEEYTEVESQLRAHL S
Sbjct: 582 PEVAELESKLTEQSKILLNLQQKVDDLASKHAYNPDEEYTEVESQLRAHLGS 737
>BF004398 similar to GP|21592356|gb nuclear matrix protein 1 {Arabidopsis
thaliana}, partial (34%)
Length = 605
Score = 216 bits (549), Expect(2) = 4e-57
Identities = 108/109 (99%), Positives = 108/109 (99%)
Frame = -1
Query: 221 LETARTFNMIYTKEIRPWTHMMEVPQLHGFGPAANRLLEAYKMLLKFLGNLQNLRDSHAA 280
LETAR FNMIYTKEIRPWTHMMEVPQLHGFGPAANRLLEAYKMLLKFLGNLQNLRDSHAA
Sbjct: 575 LETARPFNMIYTKEIRPWTHMMEVPQLHGFGPAANRLLEAYKMLLKFLGNLQNLRDSHAA 396
Query: 281 LAFGSSETSGGPSSVSRIISECESEMTVINRDLGILSASIAREHGEKMS 329
LAFGSSETSGGPSSVSRIISECESEMTVINRDLGILSASIAREHGEKMS
Sbjct: 395 LAFGSSETSGGPSSVSRIISECESEMTVINRDLGILSASIAREHGEKMS 249
Score = 23.5 bits (49), Expect(2) = 4e-57
Identities = 10/15 (66%), Positives = 12/15 (79%)
Frame = -3
Query: 212 QLRAHLESFLETART 226
Q+RAHLESFL + T
Sbjct: 603 QMRAHLESFLRNS*T 559
>BE318095 similar to GP|21592356|gb| nuclear matrix protein 1 {Arabidopsis
thaliana}, partial (12%)
Length = 332
Score = 80.5 bits (197), Expect = 8e-16
Identities = 39/41 (95%), Positives = 39/41 (95%)
Frame = +1
Query: 1 MASKQMEVIQKKLGMLNYPRANASAQSLLFAGMERYALFEW 41
MASKQME IQKKLGMLNYPRANASAQSLLFAGMERYAL EW
Sbjct: 208 MASKQMEAIQKKLGMLNYPRANASAQSLLFAGMERYALLEW 330
>TC91032 weakly similar to GP|20196995|gb|AAB91977.2 expressed protein
{Arabidopsis thaliana}, partial (37%)
Length = 1366
Score = 29.3 bits (64), Expect = 2.2
Identities = 40/147 (27%), Positives = 59/147 (39%), Gaps = 7/147 (4%)
Frame = -1
Query: 184 LLNLQQKVDDLASKH-----AYNPDEEYTEVESQ--LRAHLESFLETARTFNMIYTKEIR 236
LLNLQ +++ L+SKH NP + S L A SF +A TFN
Sbjct: 622 LLNLQTQIEFLSSKH*DHSFLNNPYTNGSRSSSSPILNATFNSFNPSAATFNFPLFNSSF 443
Query: 237 PWTHMMEVPQLHGFGPAANRLLEAYKMLLKFLGNLQNLRDSHAALAFGSSETSGGPSSVS 296
+++ P + P N L L F+ + +L + A G S+TS
Sbjct: 442 TLSNLSFNPTITPTNP-NNSLFTFNSSLFPFISSTTSLTSTSA----GRSDTS------L 296
Query: 297 RIISECESEMTVINRDLGILSASIARE 323
R +S +TV +AS+ RE
Sbjct: 295 RTVSRSSETLTVCE------TASLQRE 233
>AW574081
Length = 405
Score = 28.1 bits (61), Expect = 4.9
Identities = 14/38 (36%), Positives = 23/38 (59%)
Frame = -3
Query: 290 GGPSSVSRIISECESEMTVINRDLGILSASIAREHGEK 327
G P + R++++ +E+ VI L AS+ R+HGEK
Sbjct: 142 GIPGKIHRLLAQSRAELQVI------L*ASVTRKHGEK 47
>TC85227 similar to GP|15983773|gb|AAL10483.1 AT4g29060/F19B15_90
{Arabidopsis thaliana}, partial (63%)
Length = 3731
Score = 27.7 bits (60), Expect = 6.4
Identities = 15/43 (34%), Positives = 22/43 (50%)
Frame = -3
Query: 27 SLLFAGMERYALFEWLFFRLLGDKSPFSQQNLQGDALDRDEET 69
SL A +ER A E + LGD+SP + +GD+ + T
Sbjct: 609 SLFLARLERLAFAEPVLASDLGDESPLVPTSFEGDSPETSSAT 481
Database: MTGI
Posted date: Oct 22, 2004 3:39 PM
Number of letters in database: 27,044,181
Number of sequences in database: 36,976
Lambda K H
0.332 0.143 0.430
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 8,376,078
Number of Sequences: 36976
Number of extensions: 103650
Number of successful extensions: 585
Number of sequences better than 10.0: 13
Number of HSP's better than 10.0 without gapping: 582
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 585
length of query: 329
length of database: 9,014,727
effective HSP length: 96
effective length of query: 233
effective length of database: 5,465,031
effective search space: 1273352223
effective search space used: 1273352223
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 15 ( 7.2 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 40 (22.0 bits)
S2: 58 (26.9 bits)
Medicago: description of AC140721.10


