

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC140720.4 + phase: 0
(653 letters)
Database: MTGI
36,976 sequences; 27,044,181 total letters
Searching..................................................done
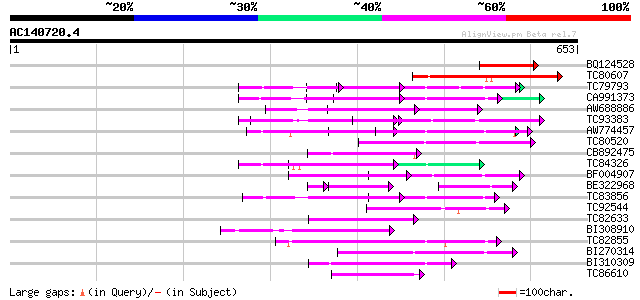
Score E
Sequences producing significant alignments: (bits) Value
BQ124528 weakly similar to GP|9757878|dbj| gb|AAD25748.1~gene_id... 140 2e-33
TC80607 weakly similar to PIR|D86431|D86431 protein T5I8.6 [impo... 132 4e-31
TC79793 weakly similar to GP|6630464|gb|AAF19552.1| F23N19.4 {Ar... 79 5e-15
CA991373 weakly similar to PIR|D86260|D86 protein T12C24.22 [imp... 70 3e-12
AW688886 similar to GP|3258568|gb|A Unknown protein {Arabidopsis... 67 2e-11
TC93383 similar to PIR|T47786|T47786 hypothetical protein F17J16... 67 2e-11
AW774457 weakly similar to GP|10176973|dbj gb|AAF19552.1~gene_id... 66 5e-11
TC80520 weakly similar to PIR|T02562|T02562 probable salt-induci... 59 8e-09
CB892475 weakly similar to GP|9280662|gb|A F21B7.18 {Arabidopsis... 58 1e-08
TC84326 weakly similar to PIR|F85225|F85225 hypothetical protein... 57 2e-08
BF004907 weakly similar to GP|8953393|emb putative protein {Arab... 56 5e-08
BE322968 similar to GP|5882738|gb| Contains 3 PF|01535 DUF17 dom... 55 5e-08
TC83856 weakly similar to PIR|B96656|B96656 unknown protein 419... 55 7e-08
TC92544 weakly similar to PIR|A86190|A86190 hypothetical protein... 54 2e-07
TC82633 similar to GP|15450347|gb|AAK96467.1 At1g20300/F14O10_8 ... 54 2e-07
BI308910 similar to GP|6682243|gb| hypothetical protein {Arabido... 53 4e-07
TC82855 weakly similar to PIR|T45829|T45829 hypothetical protein... 53 4e-07
BI270314 similar to GP|9759030|dbj contains similarity to salt-i... 52 7e-07
BI310309 similar to GP|15810431|gb unknown protein {Arabidopsis ... 52 9e-07
TC86610 similar to GP|15809886|gb|AAL06871.1 AT5g20890/F22D1_60 ... 51 1e-06
>BQ124528 weakly similar to GP|9757878|dbj|
gb|AAD25748.1~gene_id:K9I9.14~similar to unknown protein
{Arabidopsis thaliana}, partial (10%)
Length = 558
Score = 140 bits (353), Expect = 2e-33
Identities = 67/68 (98%), Positives = 68/68 (99%)
Frame = +2
Query: 542 VLFEEVKVAKSDLRPDAYTYNLMLEASSRGHQWEYFEHVYKEMILSGYHLDQNKHLPLLV 601
VLFEEVKVAKSDLRPDAYTYNLMLEASSRGHQWEYFEHVYKEMILSGYHLDQNKHLPLLV
Sbjct: 2 VLFEEVKVAKSDLRPDAYTYNLMLEASSRGHQWEYFEHVYKEMILSGYHLDQNKHLPLLV 181
Query: 602 KASRAGKV 609
KASRAGK+
Sbjct: 182 KASRAGKL 205
Score = 33.1 bits (74), Expect = 0.35
Identities = 20/82 (24%), Positives = 36/82 (43%)
Frame = +2
Query: 342 LEPDVVIYNAVLNACVPSKQWKGVSWVFQQMRKSSLKPNGATYGLAMEVMLQSGNYDLVH 401
L PD YN +L A QW+ V+++M S + + + ++G L+
Sbjct: 38 LRPDAYTYNLMLEASSRGHQWEYFEHVYKEMILSGYHLDQNKHLPLLVKASRAGKLHLLE 217
Query: 402 ELFEKMQRNGEVPEALTYKVMV 423
F+ + GE+P L + +V
Sbjct: 218 HAFDMVLEAGEIPHHLFFFELV 283
>TC80607 weakly similar to PIR|D86431|D86431 protein T5I8.6 [imported] -
Arabidopsis thaliana, partial (17%)
Length = 907
Score = 132 bits (332), Expect = 4e-31
Identities = 70/182 (38%), Positives = 115/182 (62%), Gaps = 10/182 (5%)
Frame = +1
Query: 465 GRWQDATLEVEKIKRLPHAKPLEVTFTGMIRSSMDGGHIDDCICIFEYMQDHCAPNVGTV 524
GR +A ++++KI ++ + KPL VT+TG++++S+D G+I D IFE M+D CAPN+ T
Sbjct: 4 GRSCEALMQIDKICKVAN-KPLVVTYTGLMQASLDSGNIQDGSYIFEKMKDICAPNLVTY 180
Query: 525 NTMLKVYSQNDMFSTAKVLFEEV-----KVAKSD-----LRPDAYTYNLMLEASSRGHQW 574
N MLK Y + MF AK LFE++ ++++D + PD YT+N ML+A + +W
Sbjct: 181 NIMLKAYVDHGMFREAKELFEQMLENTNHLSRNDDYKMRVIPDIYTFNTMLDACAAEKRW 360
Query: 575 EYFEHVYKEMILSGYHLDQNKHLPLLVKASRAGKVDNILVIYIRSQCLFYISAHSIISYK 634
+YF+HVY+ M+ GYH + +HL ++++ASRAGK + + + + I S+I +
Sbjct: 361 DYFDHVYQRMLYHGYHFNPKRHLRMILEASRAGKEEPLEITWNHLAATDRIPPVSLIKER 540
Query: 635 NC 636
C
Sbjct: 541 FC 546
Score = 35.0 bits (79), Expect = 0.091
Identities = 18/78 (23%), Positives = 36/78 (46%)
Frame = +1
Query: 295 PDMAAYHSIAVTLGQAGLLKELLNIVECMRQKPETLKYMYRKNWDPTLEPDVVIYNAVLN 354
P++ Y+ + G+ +E + E M + L ++ + PD+ +N +L+
Sbjct: 163 PNLVTYNIMLKAYVDHGMFREAKELFEQMLENTNHLSR--NDDYKMRVIPDIYTFNTMLD 336
Query: 355 ACVPSKQWKGVSWVFQQM 372
AC K+W V+Q+M
Sbjct: 337 ACAAEKRWDYFDHVYQRM 390
Score = 33.5 bits (75), Expect = 0.26
Identities = 22/69 (31%), Positives = 33/69 (46%)
Frame = +1
Query: 374 KSSLKPNGATYGLAMEVMLQSGNYDLVHELFEKMQRNGEVPEALTYKVMVRTFWKEGKVD 433
K + KP TY M+ L SGN +FEKM ++ P +TY +M++ + G
Sbjct: 46 KVANKPLVVTYTGLMQASLDSGNIQDGSYIFEKM-KDICAPNLVTYNIMLKAYVDHGMFR 222
Query: 434 EAVKAVRDM 442
EA + M
Sbjct: 223 EAKELFEQM 249
>TC79793 weakly similar to GP|6630464|gb|AAF19552.1| F23N19.4 {Arabidopsis
thaliana}, partial (7%)
Length = 1580
Score = 79.0 bits (193), Expect = 5e-15
Identities = 57/249 (22%), Positives = 114/249 (44%), Gaps = 2/249 (0%)
Frame = +1
Query: 342 LEPDVVIYNAVLNACVPSKQWKGVSWVFQQMRKSSLKPNGATYGLAMEVMLQSGNYDLVH 401
++P+ V YN++++ K+ +F M + + P+ +Y + + + +D
Sbjct: 544 IKPNFVTYNSLMDGYCLVKEVNKAKSIFNTMAQGGVNPDIQSYSIMINGFCKIKKFDEAM 723
Query: 402 ELFEKMQRNGEVPEALTYKVMVRTFWKEGKVDEAVKAVRDMERRGVMGTASVYYELACCL 461
LF++M R +P+ +TY ++ K G++ A++ V M RGV Y + L
Sbjct: 724 NLFKEMHRKNIIPDVVTYSSLIDGLSKSGRISYALQLVDQMHDRGVPPNICTYNSILDAL 903
Query: 462 CNCGRWQDATLEVEKIKRLPHAKPLEVTFTGMIRSSMDGGHIDDCICIFE--YMQDHCAP 519
C + A + K K +P T++ +I+ G ++D +FE ++ H
Sbjct: 904 CKTHQVDKAIALLTKFKD-KGFQPDISTYSILIKGLCQSGKLEDARKVFEGLLVKGHNL- 1077
Query: 520 NVGTVNTMLKVYSQNDMFSTAKVLFEEVKVAKSDLRPDAYTYNLMLEASSRGHQWEYFEH 579
NV T M++ + +F+ A L K+ + PDA TY +++ + + + + E
Sbjct: 1078NVDTYTIMIQGFCVEGLFNEALALLS--KMEDNGCIPDAKTYEIIILSLFKKDENDMAEK 1251
Query: 580 VYKEMILSG 588
+ +EMI G
Sbjct: 1252LLREMIARG 1278
Score = 64.3 bits (155), Expect = 1e-10
Identities = 44/191 (23%), Positives = 87/191 (45%)
Frame = +1
Query: 264 YTKLLAVLGKARRPKEALQIFNMMLGNIRVYPDMAAYHSIAVTLGQAGLLKELLNIVECM 323
Y+ ++ K ++ EA+ +F M + PD+ Y S+ L ++G + L +V+ M
Sbjct: 670 YSIMINGFCKIKKFDEAMNLFKEMHRK-NIIPDVVTYSSLIDGLSKSGRISYALQLVDQM 846
Query: 324 RQKPETLKYMYRKNWDPTLEPDVVIYNAVLNACVPSKQWKGVSWVFQQMRKSSLKPNGAT 383
D + P++ YN++L+A + Q + + + +P+ +T
Sbjct: 847 H--------------DRGVPPNICTYNSILDALCKTHQVDKAIALLTKFKDKGFQPDIST 984
Query: 384 YGLAMEVMLQSGNYDLVHELFEKMQRNGEVPEALTYKVMVRTFWKEGKVDEAVKAVRDME 443
Y + ++ + QSG + ++FE + G TY +M++ F EG +EA+ + ME
Sbjct: 985 YSILIKGLCQSGKLEDARKVFEGLLVKGHNLNVDTYTIMIQGFCVEGLFNEALALLSKME 1164
Query: 444 RRGVMGTASVY 454
G + A Y
Sbjct: 1165DNGCIPDAKTY 1197
Score = 57.4 bits (137), Expect(2) = 1e-12
Identities = 44/217 (20%), Positives = 86/217 (39%), Gaps = 1/217 (0%)
Frame = +1
Query: 377 LKPNGATYGLAMEVMLQSGNYDLVHELFEKMQRNGEVPEALTYKVMVRTFWKEGKVDEAV 436
+KPN TY M+ + +F M + G P+ +Y +M+ F K K DEA+
Sbjct: 544 IKPNFVTYNSLMDGYCLVKEVNKAKSIFNTMAQGGVNPDIQSYSIMINGFCKIKKFDEAM 723
Query: 437 KAVRDMERRGVMGTASVYYELACCLCNCGRWQDATLEVEKIKRLPHAKPLEVTFTGMIRS 496
++M R+ ++ P VT++ +I
Sbjct: 724 NLFKEMHRKNII------------------------------------PDVVTYSSLIDG 795
Query: 497 SMDGGHIDDCICIFEYMQDH-CAPNVGTVNTMLKVYSQNDMFSTAKVLFEEVKVAKSDLR 555
G I + + + M D PN+ T N++L + A L + K +
Sbjct: 796 LSKSGRISYALQLVDQMHDRGVPPNICTYNSILDALCKTHQVDKAIALLTKFK--DKGFQ 969
Query: 556 PDAYTYNLMLEASSRGHQWEYFEHVYKEMILSGYHLD 592
PD TY+++++ + + E V++ +++ G++L+
Sbjct: 970 PDISTYSILIKGLCQSGKLEDARKVFEGLLVKGHNLN 1080
Score = 34.7 bits (78), Expect = 0.12
Identities = 19/98 (19%), Positives = 44/98 (44%)
Frame = +2
Query: 345 DVVIYNAVLNACVPSKQWKGVSWVFQQMRKSSLKPNGATYGLAMEVMLQSGNYDLVHELF 404
D + Y +++ + + + Q++ + ++PN Y ++ M + + +LF
Sbjct: 131 DQISYGTLIHGLCKVGETRAALDLLQRVDGNLVQPNVVMYNTIIDSMCKVKLVNEAFDLF 310
Query: 405 EKMQRNGEVPEALTYKVMVRTFWKEGKVDEAVKAVRDM 442
+M G P+ +TY ++ F GK+ +A+ M
Sbjct: 311 SEMVSKGISPDVVTYSALISGFCILGKLKDAIDLFNKM 424
Score = 33.5 bits (75), Expect(2) = 1e-12
Identities = 28/121 (23%), Positives = 55/121 (45%)
Frame = +2
Query: 264 YTKLLAVLGKARRPKEALQIFNMMLGNIRVYPDMAAYHSIAVTLGQAGLLKELLNIVECM 323
Y L+ L K + AL + + GN+ V P++ Y++I ++ + L+ E ++ M
Sbjct: 143 YGTLIHGLCKVGETRAALDLLQRVDGNL-VQPNVVMYNTIIDSMCKVKLVNEAFDLFSEM 319
Query: 324 RQKPETLKYMYRKNWDPTLEPDVVIYNAVLNACVPSKQWKGVSWVFQQMRKSSLKPNGAT 383
K + PDVV Y+A+++ + K +F +M ++KP+ T
Sbjct: 320 VSKG--------------ISPDVVTYSALISGFCILGKLKDAIDLFNKMILENIKPDVYT 457
Query: 384 Y 384
+
Sbjct: 458 F 460
Score = 32.0 bits (71), Expect = 0.77
Identities = 21/95 (22%), Positives = 46/95 (48%)
Frame = +1
Query: 519 PNVGTVNTMLKVYSQNDMFSTAKVLFEEVKVAKSDLRPDAYTYNLMLEASSRGHQWEYFE 578
PN T N+++ Y + AK +F + A+ + PD +Y++M+ + +++
Sbjct: 550 PNFVTYNSLMDGYCLVKEVNKAKSIFNTM--AQGGVNPDIQSYSIMINGFCKIKKFDEAM 723
Query: 579 HVYKEMILSGYHLDQNKHLPLLVKASRAGKVDNIL 613
+++KEM D + L+ S++G++ L
Sbjct: 724 NLFKEMHRKNIIPDVVTYSSLIDGLSKSGRISYAL 828
>CA991373 weakly similar to PIR|D86260|D86 protein T12C24.22 [imported] -
Arabidopsis thaliana, partial (6%)
Length = 756
Score = 69.7 bits (169), Expect = 3e-12
Identities = 53/227 (23%), Positives = 99/227 (43%), Gaps = 1/227 (0%)
Frame = +2
Query: 342 LEPDVVIYNAVLNACVPSKQWKGVSWVFQQMRKSSLKPNGATYGLAMEVMLQSGNYDLVH 401
++P VV Y ++++ K+ + M + + P+ +Y + ++ + D
Sbjct: 14 IKPIVVTYCSLMDGYCLVKEVNKAKSILYTMSQRGVNPDIQSYNILIDGFCKIKKVDEAM 193
Query: 402 ELFEKMQRNGEVPEALTYKVMVRTFWKEGKVDEAVKAVRDMERRGVMGTASVYYELACCL 461
LF++M +P+ +TY ++ K GK+ A+K V +M RGV Y + L
Sbjct: 194 NLFKEMHHKHIIPDVVTYNSLIDGLCKLGKISYALKLVDEMHDRGVPPDIITYSSILDAL 373
Query: 462 CNCGRWQDATLEVEKIKRLPHAKPLEVTFTGMIRSSMDGGHIDDCICIFE-YMQDHCAPN 520
C + A + K+K +P T+T +I GG ++D IFE +
Sbjct: 374 CKNHQVDKAIALLTKLKD-QGIRPNMYTYTILIDGLCKGGRLEDAHNIFEDLLVKGYNIT 550
Query: 521 VGTVNTMLKVYSQNDMFSTAKVLFEEVKVAKSDLRPDAYTYNLMLEA 567
V T M+ + +F A L ++K + PDA TY +++ +
Sbjct: 551 VNTYTVMIHGFCNKGLFDEALTLLSKMK--DNSCIPDAVTYEIIIRS 685
Score = 64.7 bits (156), Expect = 1e-10
Identities = 44/191 (23%), Positives = 86/191 (44%)
Frame = +2
Query: 264 YTKLLAVLGKARRPKEALQIFNMMLGNIRVYPDMAAYHSIAVTLGQAGLLKELLNIVECM 323
Y L+ K ++ EA+ +F M + + PD+ Y+S+ L + G + L +V+ M
Sbjct: 140 YNILIDGFCKIKKVDEAMNLFKEM-HHKHIIPDVVTYNSLIDGLCKLGKISYALKLVDEM 316
Query: 324 RQKPETLKYMYRKNWDPTLEPDVVIYNAVLNACVPSKQWKGVSWVFQQMRKSSLKPNGAT 383
D + PD++ Y+++L+A + Q + +++ ++PN T
Sbjct: 317 H--------------DRGVPPDIITYSSILDALCKNHQVDKAIALLTKLKDQGIRPNMYT 454
Query: 384 YGLAMEVMLQSGNYDLVHELFEKMQRNGEVPEALTYKVMVRTFWKEGKVDEAVKAVRDME 443
Y + ++ + + G + H +FE + G TY VM+ F +G DEA+ + M+
Sbjct: 455 YTILIDGLCKGGRLEDAHNIFEDLLVKGYNITVNTYTVMIHGFCNKGLFDEALTLLSKMK 634
Query: 444 RRGVMGTASVY 454
+ A Y
Sbjct: 635 DNSCIPDAVTY 667
Score = 59.3 bits (142), Expect = 5e-09
Identities = 47/243 (19%), Positives = 96/243 (39%), Gaps = 1/243 (0%)
Frame = +2
Query: 374 KSSLKPNGATYGLAMEVMLQSGNYDLVHELFEKMQRNGEVPEALTYKVMVRTFWKEGKVD 433
K +KP TY M+ + + M + G P+ +Y +++ F K KVD
Sbjct: 5 KQGIKPIVVTYCSLMDGYCLVKEVNKAKSILYTMSQRGVNPDIQSYNILIDGFCKIKKVD 184
Query: 434 EAVKAVRDMERRGVMGTASVYYELACCLCNCGRWQDATLEVEKIKRLPHAKPLEVTFTGM 493
EA+ ++M + H P VT+ +
Sbjct: 185 EAMNLFKEMHHK------------------------------------HIIPDVVTYNSL 256
Query: 494 IRSSMDGGHIDDCICIFEYMQDH-CAPNVGTVNTMLKVYSQNDMFSTAKVLFEEVKVAKS 552
I G I + + + M D P++ T +++L +N A L ++K
Sbjct: 257 IDGLCKLGKISYALKLVDEMHDRGVPPDIITYSSILDALCKNHQVDKAIALLTKLK--DQ 430
Query: 553 DLRPDAYTYNLMLEASSRGHQWEYFEHVYKEMILSGYHLDQNKHLPLLVKASRAGKVDNI 612
+RP+ YTY ++++ +G + E ++++++++ GY++ N + ++ G D
Sbjct: 431 GIRPNMYTYTILIDGLCKGGRLEDAHNIFEDLLVKGYNITVNTYTVMIHGFCNKGLFDEA 610
Query: 613 LVI 615
L +
Sbjct: 611 LTL 619
>AW688886 similar to GP|3258568|gb|A Unknown protein {Arabidopsis thaliana},
partial (11%)
Length = 660
Score = 67.4 bits (163), Expect = 2e-11
Identities = 43/182 (23%), Positives = 86/182 (46%), Gaps = 4/182 (2%)
Frame = +2
Query: 367 WVFQQMRKSSLKPNGATYGLAM-EVMLQSGNYDLVHELFEKMQRNGE--VPEALTYKVMV 423
+ F++M P+ TY A+ V ++ G + H L M + + P+ +TY ++
Sbjct: 5 YFFKEMTSFDCDPDVVTYNTALLMVCVEXGKIKVAHNLVNGMSKKCKDLSPDVVTYTTLI 184
Query: 424 RTFWKEGKVDEAVKAVRDMERRGVMGTASVYYELACCLCNCGRWQDATLEVEKIKRLPHA 483
R + ++ +VDEA+ + +M RG+ Y L LC +W +E++K +
Sbjct: 185 RGYCRKQEVDEALDILEEMNGRGLKPNIVTYNTLIKGLCEAQKWDKMKEILEQMKGDGGS 364
Query: 484 KPLEVTFTGMIRSSMDGGHIDDCICIFEYMQD-HCAPNVGTVNTMLKVYSQNDMFSTAKV 542
P TF +I S G++D+ +FE M+ + + + + +++ Q + A++
Sbjct: 365 IPDACTFNTLINSHCCAGNLDEAFKVFENMKKLEVSADSASYSVLIRTLCQKGDYGKAEM 544
Query: 543 LF 544
LF
Sbjct: 545 LF 550
Score = 65.9 bits (159), Expect = 5e-11
Identities = 43/180 (23%), Positives = 85/180 (46%), Gaps = 2/180 (1%)
Frame = +2
Query: 295 PDMAAYHS-IAVTLGQAGLLKELLNIVECMRQKPETLKYMYRKNWDPTLEPDVVIYNAVL 353
PD+ Y++ + + + G +K N+V M +K + L PDVV Y ++
Sbjct: 41 PDVVTYNTALLMVCVEXGKIKVAHNLVNGMSKKCKDLS------------PDVVTYTTLI 184
Query: 354 NACVPSKQWKGVSWVFQQMRKSSLKPNGATYGLAMEVMLQSGNYDLVHELFEKMQRN-GE 412
++ + ++M LKPN TY ++ + ++ +D + E+ E+M+ + G
Sbjct: 185 RGYCRKQEVDEALDILEEMNGRGLKPNIVTYNTLIKGLCEAQKWDKMKEILEQMKGDGGS 364
Query: 413 VPEALTYKVMVRTFWKEGKVDEAVKAVRDMERRGVMGTASVYYELACCLCNCGRWQDATL 472
+P+A T+ ++ + G +DEA K +M++ V ++ Y L LC G + A +
Sbjct: 365 IPDACTFNTLINSHCCAGNLDEAFKVFENMKKLEVSADSASYSVLIRTLCQKGDYGKAEM 544
Score = 40.8 bits (94), Expect = 0.002
Identities = 29/110 (26%), Positives = 52/110 (46%), Gaps = 5/110 (4%)
Frame = +2
Query: 478 KRLPHAKPLEVTFTGMIRSSMDGGHIDDCICIFEYMQDH-CAPNVGTVNTMLKVYSQNDM 536
K+ P VT+T +IR +D+ + I E M PN+ T NT++K +
Sbjct: 134 KKCKDLSPDVVTYTTLIRGYCRKQEVDEALDILEEMNGRGLKPNIVTYNTLIKGLCEAQK 313
Query: 537 FSTAKVLFEEVKVAKSDLRPDAYTYNLMLE----ASSRGHQWEYFEHVYK 582
+ K + E++K + PDA T+N ++ A + ++ FE++ K
Sbjct: 314 WDKMKEILEQMKGDGGSI-PDACTFNTLINSHCCAGNLDEAFKVFENMKK 460
Score = 34.3 bits (77), Expect = 0.16
Identities = 23/104 (22%), Positives = 48/104 (46%), Gaps = 4/104 (3%)
Frame = +2
Query: 485 PLEVTF-TGMIRSSMDGGHIDDCICIFEYMQDHC---APNVGTVNTMLKVYSQNDMFSTA 540
P VT+ T ++ ++ G I + M C +P+V T T+++ Y + A
Sbjct: 41 PDVVTYNTALLMVCVEXGKIKVAHNLVNGMSKKCKDLSPDVVTYTTLIRGYCRKQEVDEA 220
Query: 541 KVLFEEVKVAKSDLRPDAYTYNLMLEASSRGHQWEYFEHVYKEM 584
+ EE+ L+P+ TYN +++ +W+ + + ++M
Sbjct: 221 LDILEEMN--GRGLKPNIVTYNTLIKGLCEAQKWDKMKEILEQM 346
>TC93383 similar to PIR|T47786|T47786 hypothetical protein F17J16.90 -
Arabidopsis thaliana, partial (43%)
Length = 1038
Score = 67.4 bits (163), Expect = 2e-11
Identities = 48/185 (25%), Positives = 86/185 (45%)
Frame = +1
Query: 264 YTKLLAVLGKARRPKEALQIFNMMLGNIRVYPDMAAYHSIAVTLGQAGLLKELLNIVECM 323
Y L+ GKARR +EAL +F ML + V P AY+ + +G++++ + + M
Sbjct: 109 YALLINAYGKARREEEALAVFEEML-DAGVRPTRKAYNILLDAFSISGMVEQARIVFKSM 285
Query: 324 RQKPETLKYMYRKNWDPTLEPDVVIYNAVLNACVPSKQWKGVSWVFQQMRKSSLKPNGAT 383
R+ KYM PD+ Y +L+A V + +G F+++ + +PN T
Sbjct: 286 RRD----KYM----------PDLCSYTTMLSAYVNAPDMEGAEKFFKRLIQDGFEPNVVT 423
Query: 384 YGLAMEVMLQSGNYDLVHELFEKMQRNGEVPEALTYKVMVRTFWKEGKVDEAVKAVRDME 443
YG ++ ++ + + V E +E+M G ++ K G D AV ++M
Sbjct: 424 YGTLIKGYAKANDIEKVMEKYEEMLGRGIKANQTILTTIMDAHGKNGDFDSAVNWFKEMA 603
Query: 444 RRGVM 448
G++
Sbjct: 604 LNGLL 618
Score = 59.3 bits (142), Expect = 5e-09
Identities = 53/225 (23%), Positives = 109/225 (47%), Gaps = 4/225 (1%)
Frame = +1
Query: 396 NYDLVHELFEKMQRNGEVPEALTYKVMVRTFWKEGKVDEAVKAVRDMERRGVMGTASVYY 455
NY V ++++MQR P+ ++Y +++ + K + +EA+ +M GV T Y
Sbjct: 40 NYKEVSNIYDQMQRADLRPDVVSYALLINAYGKARREEEALAVFEEMLDAGVRPTRKAYN 219
Query: 456 ELACCLCNCGRWQDATLEVEKIKRLPHAKPLEVTFTGMIRSSMDGGHIDDCICIFEYM-Q 514
L G + A + + ++R + L ++T M+ + ++ ++ F+ + Q
Sbjct: 220 ILLDAFSISGMVEQARIVFKSMRRDKYMPDL-CSYTTMLSAYVNAPDMEGAEKFFKRLIQ 396
Query: 515 DHCAPNVGTVNTMLKVYSQ-NDMFSTAKVLFEEVKVAKSDLRPDAYTYNLMLEASSRGHQ 573
D PNV T T++K Y++ ND+ KV+ + ++ ++ + +++A +
Sbjct: 397 DGFEPNVVTYGTLIKGYAKANDI---EKVMEKYEEMLGRGIKANQTILTTIMDAHGKNGD 567
Query: 574 WEYFEHVYKEMILSGYHLDQN-KHLPL-LVKASRAGKVDNILVIY 616
++ + +KEM L+G DQ K++ L L K K N LV++
Sbjct: 568 FDSAVNWFKEMALNGLLPDQKAKNILLSLAKTEEDIKEANELVLH 702
Score = 48.1 bits (113), Expect = 1e-05
Identities = 35/176 (19%), Positives = 77/176 (42%)
Frame = +1
Query: 278 KEALQIFNMMLGNIRVYPDMAAYHSIAVTLGQAGLLKELLNIVECMRQKPETLKYMYRKN 337
KE I++ M + PD+ +Y + G+A +E L + E M
Sbjct: 46 KEVSNIYDQMQ-RADLRPDVVSYALLINAYGKARREEEALAVFEEML------------- 183
Query: 338 WDPTLEPDVVIYNAVLNACVPSKQWKGVSWVFQQMRKSSLKPNGATYGLAMEVMLQSGNY 397
D + P YN +L+A S + VF+ MR+ P+ +Y + + + +
Sbjct: 184 -DAGVRPTRKAYNILLDAFSISGMVEQARIVFKSMRRDKYMPDLCSYTTMLSAYVNAPDM 360
Query: 398 DLVHELFEKMQRNGEVPEALTYKVMVRTFWKEGKVDEAVKAVRDMERRGVMGTASV 453
+ + F+++ ++G P +TY +++ + K +++ ++ +M RG+ ++
Sbjct: 361 EGAEKFFKRLIQDGFEPNVVTYGTLIKGYAKANDIEKVMEKYEEMLGRGIKANQTI 528
>AW774457 weakly similar to GP|10176973|dbj
gb|AAF19552.1~gene_id:MTG13.9~strong similarity to
unknown protein {Arabidopsis thaliana}, partial (13%)
Length = 650
Score = 65.9 bits (159), Expect = 5e-11
Identities = 44/197 (22%), Positives = 86/197 (43%), Gaps = 21/197 (10%)
Frame = -1
Query: 273 KARRPKEALQIFNMMLGNIRVYPDMAAYHSIAVTLGQAGLLKELLNIVE----------- 321
K + EAL +FN M + PD Y+S+ L ++G + +V+
Sbjct: 632 KIKMVDEALSLFNDMQFK-GIAPDKVTYNSLIDGLCKSGRISYAWELVDEMHDNGQPANI 456
Query: 322 ----------CMRQKPETLKYMYRKNWDPTLEPDVVIYNAVLNACVPSKQWKGVSWVFQQ 371
C + + +K D ++PD+ +N ++ + K VFQ
Sbjct: 455 FTYNCLIDALCKNHHVDQAIALVKKIKDQGIQPDMYTFNILIYGLCKVGRLKNAQDVFQD 276
Query: 372 MRKSSLKPNGATYGLAMEVMLQSGNYDLVHELFEKMQRNGEVPEALTYKVMVRTFWKEGK 431
+ N TY + + + + G +D L KM NG +P+A+TY+ +++ + + +
Sbjct: 275 LLSKGYSVNAWTYNIMVNGLCKEGLFDEAEALLSKMDDNGIIPDAVTYETLIQALFHKDE 96
Query: 432 VDEAVKAVRDMERRGVM 448
++A K +R+M RG++
Sbjct: 95 NEKAEKLLREMIARGLL 45
Score = 58.9 bits (141), Expect = 6e-09
Identities = 47/221 (21%), Positives = 88/221 (39%)
Frame = -1
Query: 368 VFQQMRKSSLKPNGATYGLAMEVMLQSGNYDLVHELFEKMQRNGEVPEALTYKVMVRTFW 427
+F M+ + P+ TY ++ + +SG EL ++M NG+ TY ++
Sbjct: 602 LFNDMQFKGIAPDKVTYNSLIDGLCKSGRISYAWELVDEMHDNGQPANIFTYNCLIDALC 423
Query: 428 KEGKVDEAVKAVRDMERRGVMGTASVYYELACCLCNCGRWQDATLEVEKIKRLPHAKPLE 487
K VD+A+ V+ ++ +G+ + L LC GR ++A
Sbjct: 422 KNHHVDQAIALVKKIKDQGIQPDMYTFNILIYGLCKVGRLKNAQ---------------- 291
Query: 488 VTFTGMIRSSMDGGHIDDCICIFEYMQDHCAPNVGTVNTMLKVYSQNDMFSTAKVLFEEV 547
+ + + G+ + N T N M+ + +F A+ L
Sbjct: 290 ----DVFQDLLSKGY---------------SVNAWTYNIMVNGLCKEGLFDEAEALLS-- 174
Query: 548 KVAKSDLRPDAYTYNLMLEASSRGHQWEYFEHVYKEMILSG 588
K+ + + PDA TY +++A + E E + +EMI G
Sbjct: 173 KMDDNGIIPDAVTYETLIQALFHKDENEKAEKLLREMIARG 51
Score = 45.8 bits (107), Expect = 5e-05
Identities = 42/198 (21%), Positives = 87/198 (43%), Gaps = 17/198 (8%)
Frame = -1
Query: 422 MVRTFWKEGKVDEAVKAVRDMERRGVMGTASVYYELACCLCNCGRWQDATLEVEKIKRLP 481
M+ F K VDEA+ DM+ +G+ Y L LC GR A V+++
Sbjct: 650 MINGFCKIKMVDEALSLFNDMQFKGIAPDKVTYNSLIDGLCKSGRISYAWELVDEMH--D 477
Query: 482 HAKPLEV-TFTGMIRSSMDGGHIDDCICIFEYMQDH-CAPNVGTVNTMLKVYSQNDMFST 539
+ +P + T+ +I + H+D I + + ++D P++ T N ++ +
Sbjct: 476 NGQPANIFTYNCLIDALCKNHHVDQAIALVKKIKDQGIQPDMYTFNILIYGLCKVGRLKN 297
Query: 540 AKVLFEEVKVAKSDLRPDAYTYNLMLEASSRGHQWEYFEHV---------------YKEM 584
A+ +F++ + +A+TYN+M+ + ++ E + Y+ +
Sbjct: 296 AQDVFQD--LLSKGYSVNAWTYNIMVNGLCKEGLFDEAEALLSKMDDNGIIPDAVTYETL 123
Query: 585 ILSGYHLDQNKHLPLLVK 602
I + +H D+N+ L++
Sbjct: 122 IQALFHKDENEKAEKLLR 69
>TC80520 weakly similar to PIR|T02562|T02562 probable salt-inducible protein
[imported] - Arabidopsis thaliana, partial (6%)
Length = 1911
Score = 58.5 bits (140), Expect = 8e-09
Identities = 48/205 (23%), Positives = 92/205 (44%), Gaps = 1/205 (0%)
Frame = +3
Query: 402 ELFEKMQRNGEVPEALTYKVMVRTFWKEGKVDEAVKAVRDMERRGVMGTASVYYELACCL 461
+L++ M G ++Y M+ + GK DEA+ +M R+G+ Y L L
Sbjct: 1062 KLYDDMCGRGYAENVVSYSTMISGLYLHGKTDEALSLFHEMSRKGIARDLISYNSLIKGL 1241
Query: 462 CNCGRWQDATLEVEKIKRLPHAKPLEVTFTGMIRSSMDGGHIDDCICIFEYMQD-HCAPN 520
C G AT + K+ + +P +FT +I+ G + + + + M D H P
Sbjct: 1242 CQEGELAKATNLLNKL-LIQGLEPSVSSFTPLIKCLCKVGDTEGAMRLLKDMHDRHLEPI 1418
Query: 521 VGTVNTMLKVYSQNDMFSTAKVLFEEVKVAKSDLRPDAYTYNLMLEASSRGHQWEYFEHV 580
T + M+ S+ F A+ + +K+ L+P+ T+ +++ R ++ + V
Sbjct: 1419 ASTHDYMIIGLSKKGDF--AQGMEWLLKMLSWKLKPNMQTFEHLIDCLKRENRLDDILIV 1592
Query: 581 YKEMILSGYHLDQNKHLPLLVKASR 605
M GY L++++ L+ K S+
Sbjct: 1593 LDLMFREGYRLEKSRICSLVTKFSK 1667
Score = 37.7 bits (86), Expect = 0.014
Identities = 35/155 (22%), Positives = 64/155 (40%), Gaps = 4/155 (2%)
Frame = +1
Query: 345 DVVIYNAVLNA-CVPSKQWKGVSWVFQQMRKSSLKPNGATYGLAMEVMLQSGNYDLVHEL 403
DV ++ A C +K + G + +Q+ + L + + + + YD V E+
Sbjct: 577 DVETVGCLIKAFCAENKVFNGYE-LLRQVLEKGLCVDNTVFNALINGFCKQKQYDRVSEI 753
Query: 404 FEKMQRNGEVPEALTYKVMVRTFWKEGKVDEAVKAVRDMERRGVMGTASVYYELACCLCN 463
M P TY+ ++ K K DEA + D++ RG +Y + C+
Sbjct: 754 LHIMIAMKCNPSIYTYQEIINGLLKRRKNDEAFRVFNDLKDRGYFPDRVMYTTVIKGFCD 933
Query: 464 CGRWQDA-TLEVEKIKR--LPHAKPLEVTFTGMIR 495
G +A L E I++ +P+ V G++R
Sbjct: 934 MGLLAEARKLWFEMIQKGLVPNEYTYNVMIYGIVR 1038
>CB892475 weakly similar to GP|9280662|gb|A F21B7.18 {Arabidopsis thaliana},
partial (5%)
Length = 527
Score = 57.8 bits (138), Expect = 1e-08
Identities = 37/135 (27%), Positives = 66/135 (48%), Gaps = 4/135 (2%)
Frame = +1
Query: 344 PDVVIYNAVLNA-CVPSKQWKGVSWVFQQMRKSSLKPNGATYGLAMEVMLQSGNYDLVHE 402
PD++ N VLN C + + +S+V+ ++ N +Y + ++ Y H
Sbjct: 43 PDMISCNVVLNGFCKMGRLEEALSFVWM-IKNDGFSLNRNSYASLINAFFKARRYREAHA 219
Query: 403 LFEKMQRNGEVPEALTYKVMVRTFWKEGKVDEAVKAVRDMERRGVMGTASVYYELACCLC 462
+ KM + G VP+ + Y +M+R KEG+V EA K + +M + G+ + Y + LC
Sbjct: 220 CYTKMFKEGIVPDVVLYAIMIRGLSKEGRVGEAAKMLEEMTQIGLTPDSYCYNAVIQGLC 399
Query: 463 NC---GRWQDATLEV 474
+ R Q +LE+
Sbjct: 400 DVDLFDRAQSLSLEI 444
>TC84326 weakly similar to PIR|F85225|F85225 hypothetical protein AT4g19900
[imported] - Arabidopsis thaliana, partial (18%)
Length = 756
Score = 57.0 bits (136), Expect = 2e-08
Identities = 42/227 (18%), Positives = 89/227 (38%), Gaps = 1/227 (0%)
Frame = +1
Query: 322 CMRQKPETLKYMYRKNWDPTLEPDVVIYNAVLNACVPSKQWKGVSWVFQQMRKSSLKPNG 381
C K + + + + L P+ Y +++ + ++ + M PN
Sbjct: 25 CREDKLNRAEMLLSRMKEQGLVPNTNTYTTLIDGHCKAGNFERAYDLMNLMSSEGFSPNL 204
Query: 382 ATYGLAMEVMLQSGNYDLVHELFEKMQRNGEVPEALTYKVMVRTFWKEGKVDEAVKAVRD 441
TY + + + G +++ E +NG P+ TY +++ K+ + +A+
Sbjct: 205 CTYNAIVNGLCKRGRVQEAYKMLEDGFQNGLKPDKFTYNILMSEHCKQENIRQALALFNK 384
Query: 442 MERRGVMGTASVYYELACCLCNCGRWQDATLEVEKIKRLPHAKPLEVTFTGMIRSSMDGG 501
M + G+ Y L C R +++ + E+ R+ P T+T MI G
Sbjct: 385 MLKIGIQPDIHSYTTLIAVFCRENRMKESEMFFEEAVRI-GIIPTNKTYTSMICGYCREG 561
Query: 502 HIDDCICIFEYMQDH-CAPNVGTVNTMLKVYSQNDMFSTAKVLFEEV 547
++ + F + DH CAP T ++ + A+ L++ +
Sbjct: 562 NLTLAMKFFHRLSDHGCAPESITYGAIISGLCKQSKRDEARSLYDSM 702
Score = 51.2 bits (121), Expect = 1e-06
Identities = 44/206 (21%), Positives = 84/206 (40%), Gaps = 21/206 (10%)
Frame = +1
Query: 264 YTKLLAVLGKARRPKEALQIFNMMLGNIRVYPDMAAYHSIAVTLGQAGLLKELLNIVECM 323
YT L+ KA + A + N+M P++ Y++I L + G ++E ++E
Sbjct: 106 YTTLIDGHCKAGNFERAYDLMNLMSSE-GFSPNLCTYNAIVNGLCKRGRVQEAYKMLEDG 282
Query: 324 RQ---KPETLKY------------------MYRKNWDPTLEPDVVIYNAVLNACVPSKQW 362
Q KP+ Y ++ K ++PD+ Y ++ +
Sbjct: 283 FQNGLKPDKFTYNILMSEHCKQENIRQALALFNKMLKIGIQPDIHSYTTLIAVFCRENRM 462
Query: 363 KGVSWVFQQMRKSSLKPNGATYGLAMEVMLQSGNYDLVHELFEKMQRNGEVPEALTYKVM 422
K F++ + + P TY + + GN L + F ++ +G PE++TY +
Sbjct: 463 KESEMFFEEAVRIGIIPTNKTYTSMICGYCREGNLTLAMKFFHRLSDHGCAPESITYGAI 642
Query: 423 VRTFWKEGKVDEAVKAVRDMERRGVM 448
+ K+ K DEA M +G++
Sbjct: 643 ISGLCKQSKRDEARSLYDSMIEKGLV 720
Score = 32.0 bits (71), Expect = 0.77
Identities = 33/191 (17%), Positives = 72/191 (37%)
Frame = +1
Query: 419 YKVMVRTFWKEGKVDEAVKAVRDMERRGVMGTASVYYELACCLCNCGRWQDATLEVEKIK 478
Y M+R + +E K++ A + M+ +G++ + Y L C G ++ A
Sbjct: 1 YTAMIRGYCREDKLNRAEMLLSRMKEQGLVPNTNTYTTLIDGHCKAGNFERA-------- 156
Query: 479 RLPHAKPLEVTFTGMIRSSMDGGHIDDCICIFEYMQDHCAPNVGTVNTMLKVYSQNDMFS 538
+ M S +G +PN+ T N ++ +
Sbjct: 157 -----------YDLMNLMSSEG----------------FSPNLCTYNAIVNGLCKRGRVQ 255
Query: 539 TAKVLFEEVKVAKSDLRPDAYTYNLMLEASSRGHQWEYFEHVYKEMILSGYHLDQNKHLP 598
A + E+ ++ L+PD +TYN+++ + ++ +M+ G D + +
Sbjct: 256 EAYKMLEDG--FQNGLKPDKFTYNILMSEHCKQENIRQALALFNKMLKIGIQPDIHSYTT 429
Query: 599 LLVKASRAGKV 609
L+ R ++
Sbjct: 430 LIAVFCRENRM 462
>BF004907 weakly similar to GP|8953393|emb putative protein {Arabidopsis
thaliana}, partial (34%)
Length = 665
Score = 55.8 bits (133), Expect = 5e-08
Identities = 44/180 (24%), Positives = 79/180 (43%), Gaps = 1/180 (0%)
Frame = +3
Query: 414 PEALTYKVMVRTFWKEGKVDEAVKAVRDMERRGVMGTASVYYELACCLCNCGRWQDATLE 473
P +TY +V + + +V++A++ V +M + G+ A VY + L GR+++A
Sbjct: 21 PSVVTYGTLVEGYCRMRRVEKALEMVGEMTKEGIKPNAIVYNPIIDALAEAGRFKEALGM 200
Query: 474 VEKIKRLPHAKPLEVTFTGMIRSSMDGGHIDDCICIFEYMQDH-CAPNVGTVNTMLKVYS 532
+E+ L L T+ +++ G I+ I + M P T N + +S
Sbjct: 201 MERFHVLQIGPTLS-TYNSLVKGFCKAGDIEGASKILKKMISRGFLPIPTTYNYFFRYFS 377
Query: 533 QNDMFSTAKVLFEEVKVAKSDLRPDAYTYNLMLEASSRGHQWEYFEHVYKEMILSGYHLD 592
+ L+ K+ +S PD TY+L+L+ + E V EM GY +D
Sbjct: 378 RCGKVDEGMNLY--TKMIESGHNPDRLTYHLVLKMLCEEEKLELAVQVSMEMRHKGYDMD 551
Score = 50.4 bits (119), Expect = 2e-06
Identities = 30/141 (21%), Positives = 63/141 (44%)
Frame = +3
Query: 322 CMRQKPETLKYMYRKNWDPTLEPDVVIYNAVLNACVPSKQWKGVSWVFQQMRKSSLKPNG 381
C ++ E M + ++P+ ++YN +++A + ++K + ++ + P
Sbjct: 60 CRMRRVEKALEMVGEMTKEGIKPNAIVYNPIIDALAEAGRFKEALGMMERFHVLQIGPTL 239
Query: 382 ATYGLAMEVMLQSGNYDLVHELFEKMQRNGEVPEALTYKVMVRTFWKEGKVDEAVKAVRD 441
+TY ++ ++G+ + ++ +KM G +P TY R F + GKVDE +
Sbjct: 240 STYNSLVKGFCKAGDIEGASKILKKMISRGFLPIPTTYNYFFRYFSRCGKVDEGMNLYTK 419
Query: 442 MERRGVMGTASVYYELACCLC 462
M G Y+ + LC
Sbjct: 420 MIESGHNPDRLTYHLVLKMLC 482
>BE322968 similar to GP|5882738|gb| Contains 3 PF|01535 DUF17 domains.
{Arabidopsis thaliana}, partial (13%)
Length = 369
Score = 54.7 bits (130), Expect(2) = 5e-08
Identities = 26/75 (34%), Positives = 44/75 (58%)
Frame = +3
Query: 368 VFQQMRKSSLKPNGATYGLAMEVMLQSGNYDLVHELFEKMQRNGEVPEALTYKVMVRTFW 427
VF+QM+++ PN ATY + + + + G YD V +LF +M+ + P+A TY ++++ F
Sbjct: 120 VFRQMQEAGCVPNSATYSILLNLYGKHGRYDDVRDLFLEMKVSNTDPDAGTYNILIQVFG 299
Query: 428 KEGKVDEAVKAVRDM 442
+ G E V DM
Sbjct: 300 EGGYFKEVVTLFHDM 344
Score = 52.8 bits (125), Expect = 4e-07
Identities = 28/92 (30%), Positives = 51/92 (55%), Gaps = 1/92 (1%)
Frame = +3
Query: 495 RSSMDGGHIDDCICIFEYMQDH-CAPNVGTVNTMLKVYSQNDMFSTAKVLFEEVKVAKSD 553
R D G I + I +F MQ+ C PN T + +L +Y ++ + + LF E+KV+ +D
Sbjct: 78 RLMRDMGFIKESIGVFRQMQEAGCVPNSATYSILLNLYGKHGRYDDVRDLFLEMKVSNTD 257
Query: 554 LRPDAYTYNLMLEASSRGHQWEYFEHVYKEMI 585
PDA TYN++++ G ++ ++ +M+
Sbjct: 258 --PDAGTYNILIQVFGEGGYFKEVVTLFHDMV 347
Score = 20.8 bits (42), Expect(2) = 5e-08
Identities = 10/24 (41%), Positives = 12/24 (49%)
Frame = +2
Query: 344 PDVVIYNAVLNACVPSKQWKGVSW 367
PDV YN +L A + GV W
Sbjct: 47 PDVSSYNVLLEAYARYGVY*GVYW 118
>TC83856 weakly similar to PIR|B96656|B96656 unknown protein 41955-40111
[imported] - Arabidopsis thaliana, partial (12%)
Length = 662
Score = 55.5 bits (132), Expect = 7e-08
Identities = 41/186 (22%), Positives = 81/186 (43%)
Frame = +2
Query: 269 AVLGKARRPKEALQIFNMMLGNIRVYPDMAAYHSIAVTLGQAGLLKELLNIVECMRQKPE 328
AV G + + + L NI+ PD+ ++ + ++G + L +V+ M + +
Sbjct: 2 AVFGTRAERLQLICLIK*YLENIK--PDVYTFNILVDVFCKSGKISYALKLVDEMHDRGQ 175
Query: 329 TLKYMYRKNWDPTLEPDVVIYNAVLNACVPSKQWKGVSWVFQQMRKSSLKPNGATYGLAM 388
P++V Y+++L+A + + + +++ ++PN TY + +
Sbjct: 176 P--------------PNIVTYSSILDALCKTHRVDKAVALLTKLKDQGIRPNMHTYTILI 313
Query: 389 EVMLQSGNYDLVHELFEKMQRNGEVPEALTYKVMVRTFWKEGKVDEAVKAVRDMERRGVM 448
+ + SG + +FE + G +TY VM F K+G DEA + ME G +
Sbjct: 314 DGLCTSGKLEDARNIFEDLLVKGYDITVVTYIVMFYGFCKKGLFDEASALLSKMEENGCI 493
Query: 449 GTASVY 454
A Y
Sbjct: 494 PDAKTY 511
Score = 51.2 bits (121), Expect = 1e-06
Identities = 42/152 (27%), Positives = 65/152 (42%), Gaps = 1/152 (0%)
Frame = +2
Query: 414 PEALTYKVMVRTFWKEGKVDEAVKAVRDMERRGVMGTASVYYELACCLCNCGRWQDATLE 473
P+ T+ ++V F K GK+ A+K V +M RG Y + LC R A
Sbjct: 74 PDVYTFNILVDVFCKSGKISYALKLVDEMHDRGQPPNIVTYSSILDALCKTHRVDKAVAL 253
Query: 474 VEKIKRLPHAKPLEVTFTGMIRSSMDGGHIDDCICIFE-YMQDHCAPNVGTVNTMLKVYS 532
+ K+K +P T+T +I G ++D IFE + V T M +
Sbjct: 254 LTKLKD-QGIRPNMHTYTILIDGLCTSGKLEDARNIFEDLLVKGYDITVVTYIVMFYGFC 430
Query: 533 QNDMFSTAKVLFEEVKVAKSDLRPDAYTYNLM 564
+ +F A L K+ ++ PDA TY L+
Sbjct: 431 KKGLFDEASALLS--KMEENGCIPDAKTYELI 520
Score = 38.1 bits (87), Expect = 0.011
Identities = 25/132 (18%), Positives = 59/132 (43%), Gaps = 1/132 (0%)
Frame = +2
Query: 480 LPHAKPLEVTFTGMIRSSMDGGHIDDCICIFEYMQDHCAP-NVGTVNTMLKVYSQNDMFS 538
L + KP TF ++ G I + + + M D P N+ T +++L +
Sbjct: 59 LENIKPDVYTFNILVDVFCKSGKISYALKLVDEMHDRGQPPNIVTYSSILDALCKTHRVD 238
Query: 539 TAKVLFEEVKVAKSDLRPDAYTYNLMLEASSRGHQWEYFEHVYKEMILSGYHLDQNKHLP 598
A L ++K +RP+ +TY ++++ + E ++++++++ GY + ++
Sbjct: 239 KAVALLTKLK--DQGIRPNMHTYTILIDGLCTSGKLEDARNIFEDLLVKGYDITVVTYIV 412
Query: 599 LLVKASRAGKVD 610
+ + G D
Sbjct: 413 MFYGFCKKGLFD 448
>TC92544 weakly similar to PIR|A86190|A86190 hypothetical protein [imported]
- Arabidopsis thaliana, partial (19%)
Length = 967
Score = 53.9 bits (128), Expect = 2e-07
Identities = 41/171 (23%), Positives = 77/171 (44%), Gaps = 6/171 (3%)
Frame = +2
Query: 411 GEVPEALTYKVMVRTFWKEGKVDEAVKAVRDMERRGVMGTASVYYELACCLCNCGRWQDA 470
G VP +Y M ++EGK+DEA K + +M+ R S++ LC G+ +A
Sbjct: 200 GSVPSTASYTSMAVDLYEEGKIDEADKVIIEMKDRRFKPKHSIFEAKVAALCKVGKVDEA 379
Query: 471 TLEVEKIKRLPHAKPLEVTFTGMIRSSMDGGHIDDCICIFEYMQD-----HCAPNVGTVN 525
+E+ + P +T ++++ G + + + E + +C + T
Sbjct: 380 IKVIEEDMVEVNCLPNATVYTILLKNL---GGVGNSTSVLESLNKMSKKVNCMADKETYC 550
Query: 526 TMLKVYSQNDMFSTAKVLFEEVKVAKSDLRPDAYTYNLMLEA-SSRGHQWE 575
+L++ + + A L E++ + P A TYNL++E S G Q+E
Sbjct: 551 ILLEMLCRERKYLEASQLLEQMSI--KSYWPCANTYNLLIEGLCSLGRQYE 697
Score = 31.2 bits (69), Expect = 1.3
Identities = 42/211 (19%), Positives = 78/211 (36%), Gaps = 42/211 (19%)
Frame = +2
Query: 421 VMVRTFWK----EGKVDEAVKAVRDMERRGVMGTASVYYELACCLCNCG-------RWQD 469
+++ +W GK DEAV+ + + R+G+ Y L C G RW
Sbjct: 5 ILLELYWMLYVIMGKFDEAVEILGKILRKGLKAPKRCYNRLDISQCGDGKDAEVTKRWIH 184
Query: 470 ATL-------------------------EVEKI---KRLPHAKPLEVTFTGMIRSSMDGG 501
L E +K+ + KP F + + G
Sbjct: 185 EALVRGSVPSTASYTSMAVDLYEEGKIDEADKVIIEMKDRRFKPKHSIFEAKVAALCKVG 364
Query: 502 HIDDCICIFE--YMQDHCAPNVGTVNTMLKVYSQNDMFSTAKVLFEEVKVAKS-DLRPDA 558
+D+ I + E ++ +C PN +LK + + ++ VL K++K + D
Sbjct: 365 KVDEAIKVIEEDMVEVNCLPNATVYTILLK--NLGGVGNSTSVLESLNKMSKKVNCMADK 538
Query: 559 YTYNLMLEASSRGHQWEYFEHVYKEMILSGY 589
TY ++LE R ++ + ++M + Y
Sbjct: 539 ETYCILLEMLCRERKYLEASQLLEQMSIKSY 631
>TC82633 similar to GP|15450347|gb|AAK96467.1 At1g20300/F14O10_8
{Arabidopsis thaliana}, partial (34%)
Length = 830
Score = 53.5 bits (127), Expect = 2e-07
Identities = 28/126 (22%), Positives = 54/126 (42%)
Frame = +1
Query: 345 DVVIYNAVLNACVPSKQWKGVSWVFQQMRKSSLKPNGATYGLAMEVMLQSGNYDLVHELF 404
D + YN ++ + + V M K + PN +T+ + + + + H ++
Sbjct: 52 DTISYNFLIESHCKDENLDEAVKVLDTMVKKGVAPNASTFNSIFGCIAELHDVNGAHRMY 231
Query: 405 EKMQRNGEVPEALTYKVMVRTFWKEGKVDEAVKAVRDMERRGVMGTASVYYELACCLCNC 464
KM+ +P LTY +++R F +D +K ++M+ V + Y L C
Sbjct: 232 AKMKELKCMPNTLTYNILMRMFADSKSIDMVLKLKKEMDESEVEPNVNTYRILILMFCEK 411
Query: 465 GRWQDA 470
G W +A
Sbjct: 412 GHWNNA 429
Score = 35.0 bits (79), Expect = 0.091
Identities = 32/136 (23%), Positives = 54/136 (39%), Gaps = 6/136 (4%)
Frame = +1
Query: 284 FNMMLGNIRVYPDMAAYHSIAVTLGQAGLLKELLNIVECMRQ----KPETLKYMYRKNWD 339
FN + G I D+ H + + + + L MR K + +K D
Sbjct: 169 FNSIFGCIAELHDVNGAHRMYAKMKELKCMPNTLTYNILMRMFADSKSIDMVLKLKKEMD 348
Query: 340 PT-LEPDVVIYNAVLNACVPSKQWKGVSWVFQQM-RKSSLKPNGATYGLAMEVMLQSGNY 397
+ +EP+V Y ++ W + ++M + LKPN + Y +E++ +G
Sbjct: 349 ESEVEPNVNTYRILILMFCEKGHWNNAYNLMKEMVEEKCLKPNLSIYETVLELLRNAGQL 528
Query: 398 DLVHELFEKMQRNGEV 413
EL EKM G V
Sbjct: 529 KKHEELVEKMVARGFV 576
>BI308910 similar to GP|6682243|gb| hypothetical protein {Arabidopsis
thaliana}, partial (23%)
Length = 591
Score = 52.8 bits (125), Expect = 4e-07
Identities = 47/202 (23%), Positives = 87/202 (42%), Gaps = 1/202 (0%)
Frame = +1
Query: 243 LSVVQWVYNSKDHRKFQSRFVYTKLLAVLGKARRPKEALQIFNMMLGNIRVYPDMAAYHS 302
L +++ +S H S YT +++ L KA A Q+ M I V + Y+
Sbjct: 37 LEAMRYQEDSSSHPDHVS---YTTVVSALVKAGFMDRAHQVLAEMT-RIGVPANRITYNI 204
Query: 303 IAVTLGQAGLLKELLNIVECMRQKPETLKYMYRKNWDPTLEPDVVIYNAVLNACVPSKQW 362
+ G K+L Q + ++ + D +EPDVV YN +++ C+
Sbjct: 205 LL-----KGYCKQL--------QMDKAMELLQEMAEDIGIEPDVVSYNILIDGCILVDDS 345
Query: 363 KGVSWVFQQMRKSSLKPNGATYGLAMEVMLQSGNYDLVHELFEKMQRNGEV-PEALTYKV 421
G F +MR + P +Y M+ SG L +F++M + V + + + +
Sbjct: 346 AGALSFFNEMRAKGIAPTKVSYTTLMKAFALSGQPKLAQRVFDEMVTDARVRVDLIAWNM 525
Query: 422 MVRTFWKEGKVDEAVKAVRDME 443
+V + + G V+E K ++ M+
Sbjct: 526 LVEGYNRLGLVEEGKKVIQKMK 591
>TC82855 weakly similar to PIR|T45829|T45829 hypothetical protein F2K15.100
- Arabidopsis thaliana, partial (63%)
Length = 1447
Score = 52.8 bits (125), Expect = 4e-07
Identities = 53/276 (19%), Positives = 120/276 (43%), Gaps = 16/276 (5%)
Frame = +3
Query: 307 LGQAGLLKELLNI-------VECMRQKPETLKYMYRKNW-DPTLEPDVVIYNAVLNACVP 358
L QAG++ ++ ++C +KP+T +++ D P + ++ V
Sbjct: 33 LTQAGVVPNIITYNLIFQTYLDC--RKPDTALENFKQFIKDAPFNPSPTTFRILIKGLVD 206
Query: 359 SKQWKGVSWVFQQMRKSSLKPNGATYGLAMEVMLQSGNYDLVHELFEKMQRN--GEVPEA 416
+ + + QM S P+ Y M +S + D V ++E+++ G V +
Sbjct: 207 NNRLDRAMDIKDQMNASGFAPDPLVYHYLMLGHARSSDGDGVLRVYEELKEKLGGVVEDG 386
Query: 417 LTYKVMVRTFWKEGKVDEAVKAVRDMERRGVMGTASVYYELACCLCNCGRWQDATLEVEK 476
+ +++ ++ +G EA++ +++ G + Y + L G++ +A ++
Sbjct: 387 VVLGCLMKGYFLKGMEKEAMECYQEVFVEGKKMSDIAYNSVLDALAKNGKFDEAMRLFDR 566
Query: 477 IKRLPHAKPLEVTFT-GMIRSSMDG----GHIDDCICIFEYM-QDHCAPNVGTVNTMLKV 530
+ + H P ++ G +DG G + I +F M + C P+ + N +++
Sbjct: 567 MIK-EHNPPAKLAVNLGSFNVMVDGYCAEGKFKEAIEVFRSMGESRCKPDTLSFNNLIEQ 743
Query: 531 YSQNDMFSTAKVLFEEVKVAKSDLRPDAYTYNLMLE 566
N M A+ ++ E++ + PD YTY L+++
Sbjct: 744 LCNNGMILEAEEVYGEME--GKGVNPDEYTYGLLMD 845
>BI270314 similar to GP|9759030|dbj contains similarity to salt-inducible
protein~gene_id:K6A12.14 {Arabidopsis thaliana}, partial
(30%)
Length = 669
Score = 52.0 bits (123), Expect = 7e-07
Identities = 46/209 (22%), Positives = 88/209 (42%), Gaps = 2/209 (0%)
Frame = +3
Query: 378 KPNGATYGLAMEVMLQSGNYDLVHELFEKMQRNGEVPEALTYKVMVRTFWKEGKV-DEAV 436
KP T+ + M + +V L +M+ G P A +Y ++ + ++ K+ D A
Sbjct: 3 KPTAVTFNILMYAYSRRMQPKIVESLLAEMKDFGLKPNANSYTCLISAYGRQKKMSDMAA 182
Query: 437 KAVRDMERRGVMGTASVYYELACCLCNCGRWQDATLEVEKIKRLPHAKPLEVTFTGMIRS 496
A M++ G+ T+ Y + G W + V + KP T+T ++ +
Sbjct: 183 DAFLKMKKVGIKPTSHSYTAMIHAYSVSG-WHEKAYAVFENMIREGIKPSIETYTTLLDA 359
Query: 497 SMDGGHIDDCICIFEYMQDHCAPNVG-TVNTMLKVYSQNDMFSTAKVLFEEVKVAKSDLR 555
G + + I++ M T N ++ +++ +F A+ + E K L+
Sbjct: 360 FRRVGDTETLMKIWKLMMSEKVKGTQVTFNILVDGFAKQGLFMEARDVISEF--GKIGLQ 533
Query: 556 PDAYTYNLMLEASSRGHQWEYFEHVYKEM 584
P TYN+++ A +RG + KEM
Sbjct: 534 PTVMTYNMLINAYARGGLDSNIPQLLKEM 620
>BI310309 similar to GP|15810431|gb unknown protein {Arabidopsis thaliana},
partial (37%)
Length = 634
Score = 51.6 bits (122), Expect = 9e-07
Identities = 40/171 (23%), Positives = 77/171 (44%), Gaps = 1/171 (0%)
Frame = +1
Query: 345 DVVIYNAVLNA-CVPSKQWKGVSWVFQQMRKSSLKPNGATYGLAMEVMLQSGNYDLVHEL 403
+ VIY +++ A C K + + V ++M + + Y M+ GN + +
Sbjct: 40 NAVIYGSIIYAHCQACKMGRAEALV-REMEEQGIDAPIDIYHTMMDGYTMIGNEEKCLIV 216
Query: 404 FEKMQRNGEVPEALTYKVMVRTFWKEGKVDEAVKAVRDMERRGVMGTASVYYELACCLCN 463
FE+++ G P ++Y ++ + K GKV +A++ R M+ G+ Y L
Sbjct: 217 FERLKECGFSPSIVSYGCLINLYTKIGKVSKALEISRVMKTVGIKHNMKTYSMLFNGFVK 396
Query: 464 CGRWQDATLEVEKIKRLPHAKPLEVTFTGMIRSSMDGGHIDDCICIFEYMQ 514
W +A E I + KP + + ++++ G++D ICI + MQ
Sbjct: 397 LKDWANAFSVFEDITK-DGLKPDVILYNNIVKAFCGMGNMDRAICIVKQMQ 546
Score = 39.7 bits (91), Expect = 0.004
Identities = 22/89 (24%), Positives = 39/89 (43%)
Frame = +1
Query: 349 YNAVLNACVPSKQWKGVSWVFQQMRKSSLKPNGATYGLAMEVMLQSGNYDLVHELFEKMQ 408
Y+ + N V K W VF+ + K LKP+ Y ++ GN D + ++MQ
Sbjct: 367 YSMLFNGFVKLKDWANAFSVFEDITKDGLKPDVILYNNIVKAFCGMGNMDRAICIVKQMQ 546
Query: 409 RNGEVPEALTYKVMVRTFWKEGKVDEAVK 437
R T+ ++ F + G+ A++
Sbjct: 547 RERHRATTRTFLPIIHGFARAGETRRALE 633
>TC86610 similar to GP|15809886|gb|AAL06871.1 AT5g20890/F22D1_60
{Arabidopsis thaliana}, partial (42%)
Length = 1425
Score = 51.2 bits (121), Expect = 1e-06
Identities = 26/107 (24%), Positives = 56/107 (52%)
Frame = +3
Query: 371 QMRKSSLKPNGATYGLAMEVMLQSGNYDLVHELFEKMQRNGEVPEALTYKVMVRTFWKEG 430
+M +L P+ T+ + + L+ G D L+E+M G VP+A+ + +++ + +G
Sbjct: 63 EMLNMNLVPDNITFSILINRFLKLGQLDEAASLYERMVSCGHVPDAVLFDSLLKGYSLKG 242
Query: 431 KVDEAVKAVRDMERRGVMGTASVYYELACCLCNCGRWQDATLEVEKI 477
K ++ V ++ M + V+ + + + CLCN + +++EKI
Sbjct: 243 KTEKVVSMLQQMADKDVVLDSKLTSTILACLCNMSK----DVDIEKI 371
Score = 33.9 bits (76), Expect = 0.20
Identities = 18/66 (27%), Positives = 34/66 (51%)
Frame = +3
Query: 392 LQSGNYDLVHELFEKMQRNGEVPEALTYKVMVRTFWKEGKVDEAVKAVRDMERRGVMGTA 451
L++G+ + EL +M VP+ +T+ +++ F K G++DEA M G + A
Sbjct: 21 LKAGDVESAKELLLEMLNMNLVPDNITFSILINRFLKLGQLDEAASLYERMVSCGHVPDA 200
Query: 452 SVYYEL 457
++ L
Sbjct: 201 VLFDSL 218
Database: MTGI
Posted date: Oct 22, 2004 3:39 PM
Number of letters in database: 27,044,181
Number of sequences in database: 36,976
Lambda K H
0.321 0.136 0.408
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 20,468,463
Number of Sequences: 36976
Number of extensions: 282815
Number of successful extensions: 1966
Number of sequences better than 10.0: 115
Number of HSP's better than 10.0 without gapping: 1828
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 1937
length of query: 653
length of database: 9,014,727
effective HSP length: 102
effective length of query: 551
effective length of database: 5,243,175
effective search space: 2888989425
effective search space used: 2888989425
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.9 bits)
S2: 62 (28.5 bits)
Medicago: description of AC140720.4


