

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
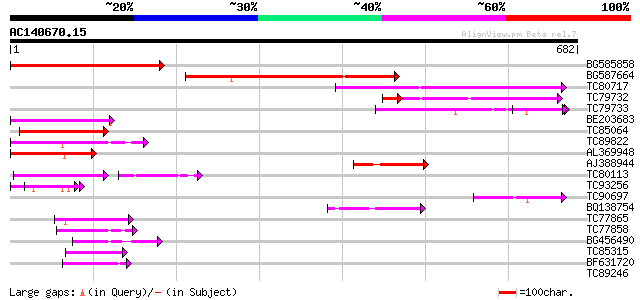
Query= AC140670.15 - phase: 0
(682 letters)
Database: MTGI
36,976 sequences; 27,044,181 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
BG585858 similar to GP|15983799|gb At2g05250/F5G3.15 {Arabidopsi... 379 e-105
BG587664 similar to GP|15983799|gb| At2g05250/F5G3.15 {Arabidops... 320 9e-88
TC80717 weakly similar to GP|6729021|gb|AAF27017.1| hypothetical... 158 6e-39
TC79732 weakly similar to GP|6729021|gb|AAF27017.1| hypothetical... 120 9e-34
TC79733 similar to PIR|T01797|T01797 hypothetical protein A_TM02... 121 8e-28
BE203683 weakly similar to GP|14587221|d contains ESTs AU108104(... 105 6e-23
TC85064 similar to GP|15529208|gb|AAK97698.1 At2g01710/T8O11.12 ... 86 5e-17
TC89822 similar to GP|15529208|gb|AAK97698.1 At2g01710/T8O11.12 ... 85 8e-17
AL369948 similar to GP|14587221|db contains ESTs AU108104(C12321... 81 2e-15
AJ388944 weakly similar to PIR|H84769|H847 probable DnaJ protein... 73 4e-13
TC80113 weakly similar to GP|21554000|gb|AAM63081.1 unknown {Ara... 66 5e-11
TC93256 60 4e-09
TC90697 homologue to GP|12322856|gb|AAG51418.1 hypothetical prot... 57 2e-08
BQ138754 weakly similar to GP|14587221|db contains ESTs AU108104... 54 2e-07
TC77865 similar to GP|17473565|gb|AAL38258.1 dnaJ-like protein {... 54 3e-07
TC77858 similar to GP|20197886|gb|AAD22362.2 putative DnaJ prote... 45 7e-05
BG456490 weakly similar to GP|22296361|dbj AHM1(AT hook-containi... 45 9e-05
TC85315 similar to GP|9758827|dbj|BAB09499.1 gene_id:MTH12.18~un... 43 3e-04
BF631720 weakly similar to GP|10798648|em putative DNAJ protein ... 42 8e-04
TC89246 homologue to PIR|T06594|T06594 heat shock protein dnaJ -... 40 0.002
>BG585858 similar to GP|15983799|gb At2g05250/F5G3.15 {Arabidopsis thaliana},
partial (19%)
Length = 851
Score = 379 bits (972), Expect = e-105
Identities = 184/186 (98%), Positives = 185/186 (98%)
Frame = +3
Query: 1 MEAKKEEALKAIENAEKRFSHRDFVGAKSYALKAKTLCPGLEGISQLVTTFEVYIASQVT 60
MEAKKEEALKAIENAEKRFSHRDFVGAKSYALKAKTLCPGLEGISQLVTTFEVYIASQVT
Sbjct: 288 MEAKKEEALKAIENAEKRFSHRDFVGAKSYALKAKTLCPGLEGISQLVTTFEVYIASQVT 467
Query: 61 CNGELDWYSIMGLNPSTNIEAVKKQYKKMAGLLHPDNNKCVGADGAFHLVSEAWARLSGS 120
CNGELDWYSIMGLNPSTNIEAVKKQYKKMAGLLHPDNNKCVGADGAFHLVSEAWARLSGS
Sbjct: 468 CNGELDWYSIMGLNPSTNIEAVKKQYKKMAGLLHPDNNKCVGADGAFHLVSEAWARLSGS 647
Query: 121 YDMKRNAQLGAGNGVNHKGLSSVHASGGNQDTFWTICTSCKVQYEYLRKYVNKKLSCKNC 180
YDMKRNAQLGAGNGVNHKGLSSVHASGGNQDTFWTICTS +VQYEYLRKYVNKKLSCKNC
Sbjct: 648 YDMKRNAQLGAGNGVNHKGLSSVHASGGNQDTFWTICTSWQVQYEYLRKYVNKKLSCKNC 827
Query: 181 RGIFIA 186
RGIFIA
Sbjct: 828 RGIFIA 845
>BG587664 similar to GP|15983799|gb| At2g05250/F5G3.15 {Arabidopsis
thaliana}, partial (4%)
Length = 813
Score = 320 bits (821), Expect = 9e-88
Identities = 169/264 (64%), Positives = 201/264 (76%), Gaps = 6/264 (2%)
Frame = +2
Query: 212 HSYDGVTYVPTNGAYFNGNGVPGYHSKHGYEYVSNVPYQLGSAGYVNQNGSTTL----SA 267
+S+DGVTYVPTN AYFNGNGV GYHS HGY+YVSNV +QLGSAG ++QNGS T S
Sbjct: 11 YSFDGVTYVPTNAAYFNGNGVTGYHSGHGYDYVSNVSFQLGSAGLIHQNGSATTLPADSV 190
Query: 268 CQTNGKAKRGRPKVKSGADRKHCLTETVVNISSDVSFSRNEPQEVKPSRPEKKRKV-LGA 326
+ NG AKRGRPKVKSGA+ + + ETVVNI+S V FS N+PQEV P RP KKRKV +GA
Sbjct: 191 YRVNGNAKRGRPKVKSGANGRPPMAETVVNINSHVLFSCNKPQEVMPDRPYKKRKVTVGA 370
Query: 327 SLRNVHEGKGSKCASELALANGNGSVGHGQRISSTSEIPTKQYSMAPAFDARKLLIEKAR 386
S RN ++ KGSKCA E + GN ++G GQ++ +E+ TK M PAFDARKLLIEKAR
Sbjct: 371 SFRNGYDAKGSKCALEAVVPKGNDNIGPGQKVVVKNEVQTKHCFMPPAFDARKLLIEKAR 550
Query: 387 TEIRKKLEEMKLASETAAAVIEGKKSQADVGQVKGDICTKTALNVSDNQLE-HRKTVPVT 445
T IRKKLEE+KL+SE AA + E +K+Q DV QVK + C K +LNVS QLE H K P++
Sbjct: 551 TVIRKKLEEIKLSSE-AATLKEKEKAQVDVCQVKRETCRKASLNVSGLQLEPHGKAGPIS 727
Query: 446 ITVPDPDFHDFDKDRSEPCFKPKQ 469
ITVPD DFHDFDKDR+E CFKPKQ
Sbjct: 728 ITVPDSDFHDFDKDRTEECFKPKQ 799
>TC80717 weakly similar to GP|6729021|gb|AAF27017.1| hypothetical protein
{Arabidopsis thaliana}, partial (28%)
Length = 1198
Score = 158 bits (400), Expect = 6e-39
Identities = 85/278 (30%), Positives = 142/278 (50%), Gaps = 1/278 (0%)
Frame = +1
Query: 393 LEEMKLASETAAAVIEGKKSQADVGQVKGDICTKTALNVSDNQLEHRKTVPVTITVPDPD 452
+ +++L S A G+ + Q + + N + P I PDP+
Sbjct: 142 ITQVQLVSLADVAAQNGEMRNKENAQPEKTVSRNKMKTEQLNPQRKETSNPDIICCPDPE 321
Query: 453 FHDFDKDRSEPCFKPKQIWALYDEEDGMPRLYCLIREVVSVNPFKINISYLSSKTDSEFG 512
F DF+K R + CF Q WA+YD D MPR Y I++V S PF + ++L +
Sbjct: 322 FSDFEKVRKKDCFAVGQYWAVYDNTDCMPRFYARIKKVHS--PFGLEYTWLEPNPVRK-D 492
Query: 513 PVNWLVSGFTKSCGNFRAMTSDVVDQVNIFSHVLSRVKAGRGGCVRIYPKCGDVWAVYRN 572
++W +G +CG +R S + + +FSH + +K G +YP G+ WA++R+
Sbjct: 493 EIDWHDAGLPVACGKYRLGHSQISRDIVMFSHEVHCIKGSGRGSYLVYPMKGETWAIFRH 672
Query: 573 WSTDWNRSTPDEVRHQYDMVEVLDDYSEELGLCVSPLIKLDGFKTVYKRNADKS-AIRYI 631
W W+ +Q++ VEVL D+ E G+ VS L K+ GF +++++ ++ I
Sbjct: 673 WDIGWSSEPEKNSEYQFEFVEVLSDFDESDGVKVSYLSKVKGFVSLFQQTVQNGISLCCI 852
Query: 632 PRREMLRFSHQVPSWLLKGEEASNLPDKCWDLDPAATP 669
P E+ RFSH+VPS+++ G+E +P ++LDPA P
Sbjct: 853 PPTELYRFSHRVPSFVMTGKEREGVPSGSYELDPAGLP 966
>TC79732 weakly similar to GP|6729021|gb|AAF27017.1| hypothetical protein
{Arabidopsis thaliana}, partial (19%)
Length = 654
Score = 120 bits (301), Expect(2) = 9e-34
Identities = 65/200 (32%), Positives = 105/200 (52%), Gaps = 1/200 (0%)
Frame = +1
Query: 467 PKQIWALYDEEDGMPRLYCLIREVVSVNPFKINISYLSSKTDSEFGPVNWLVSGFTKSCG 526
P + +YD+ DGMPR Y LI++V S FK+ I++L D E W+ +CG
Sbjct: 67 PGRYGPVYDDIDGMPRFYALIKKVFSTG-FKLQITWLEPDPDDE-EERRWVKEKLPSACG 240
Query: 527 NFRAMTSDVVDQVNIFSHVLSRVKAGRGGCVRIYPKCGDVWAVYRNWSTDWNRSTPDEVR 586
++ + +FSH++ K ++YP+ G+ WA+++NW W +
Sbjct: 241 KYQLGKTVTTKDQPMFSHLILYEKVR--STFKVYPRKGETWALFKNWDIKWYMDAESHQK 414
Query: 587 HQYDMVEVLDDYSEELGLCVSPLIKLDGFKTVYKRNADKSAIRY-IPRREMLRFSHQVPS 645
+ + VE+L DY E G+ VS L KL GF +++ R + IP E+ RFSH+VPS
Sbjct: 415 YDLEFVEILSDYVEGAGVFVSYLAKLKGFMSLFSRITKGGGCSFQIPPAELFRFSHRVPS 594
Query: 646 WLLKGEEASNLPDKCWDLDP 665
+ + G E + +P ++LDP
Sbjct: 595 FKMTGLERAGVPVGAFELDP 654
Score = 42.0 bits (97), Expect(2) = 9e-34
Identities = 16/24 (66%), Positives = 18/24 (74%)
Frame = +3
Query: 449 PDPDFHDFDKDRSEPCFKPKQIWA 472
PDP+F DFDKD+ E CF QIWA
Sbjct: 12 PDPEFSDFDKDKKEECFASGQIWA 83
>TC79733 similar to PIR|T01797|T01797 hypothetical protein A_TM021B04.9 -
Arabidopsis thaliana, partial (2%)
Length = 1253
Score = 121 bits (304), Expect = 8e-28
Identities = 81/240 (33%), Positives = 120/240 (49%), Gaps = 7/240 (2%)
Frame = +2
Query: 441 TVPVTITVPDPDFHDFDKDRSEPCFKPKQIWALYDEEDGMPRLYCLIREVVSV-NPFKIN 499
+V + VPDP F+ FD +RS F+ QIWA Y +ED +P+ Y I+ V + + ++
Sbjct: 338 SVAESFEVPDPSFNQFDAERSHEKFEAGQIWAFYGDEDELPKYYGQIKCVRRIDSKIELQ 517
Query: 500 ISYLSSKTDSEFGPVNWLVSGFTKSCGNFRAMTSD---VVDQVNIFSHVLSRVKAGRGGC 556
+ YL+ + + W SCG F+ S + N SH +
Sbjct: 518 VIYLTDCWVPK-KVIRWEDKDMIISCGRFKINPSGKLCTYNNTNSVSHQVHASAVRNNKE 694
Query: 557 VRIYPKCGDVWAVYRNWSTDWNRSTPDEVRHQYDMVEVLDDYSEELGLCVSPLIKLDGFK 616
IYP+ G++WA+YR W T RS D +YD+VEV +D ++ V L K+ G+
Sbjct: 695 YEIYPRKGEIWALYRGWRTTLKRS--DLKNCEYDIVEVTED--ADMWTDVLFLEKVSGYS 862
Query: 617 TVYK---RNADKSAIRYIPRREMLRFSHQVPSWLLKGEEASNLPDKCWDLDPAATPDELL 673
+V+K N I R E+LRFSH++P++ L E SNL W+LDPAA P L
Sbjct: 863 SVFKGKLSNGGSKMTMTIDRTELLRFSHKIPAFKLTEEHGSNLRG-FWELDPAAVPHHYL 1039
Score = 42.0 bits (97), Expect = 8e-04
Identities = 24/67 (35%), Positives = 36/67 (52%), Gaps = 1/67 (1%)
Frame = +2
Query: 606 VSPLIKLDGFKTVYKRNADKSAIRY-IPRREMLRFSHQVPSWLLKGEEASNLPDKCWDLD 664
VS L K GF +++ R + IP E+ RFSH V S+ + G E + +P ++LD
Sbjct: 8 VSYLAKSKGFMSLFSRITKGGGCSFQIPPAELFRFSHTVTSFKMTGLERAGVPVGAFELD 187
Query: 665 PAATPDE 671
P + P E
Sbjct: 188 PISLPME 208
>BE203683 weakly similar to GP|14587221|d contains ESTs AU108104(C12321)
C26435(C12321)~similar to Arabidopsis thaliana
chromosome 3 F24P17., partial (12%)
Length = 565
Score = 105 bits (262), Expect = 6e-23
Identities = 59/132 (44%), Positives = 80/132 (59%), Gaps = 6/132 (4%)
Frame = +3
Query: 1 MEAKKEEALKAIENAEKRFSHRDFVGAKSYALKAKTLCPGLEGISQLVTTFEVYI-ASQV 59
M+ KEEAL+A E AEK+ +RDF GA+ +ALKA+ L P LE I+Q++ +V+ A Q
Sbjct: 159 MDCNKEEALRAKEIAEKKMENRDFAGARKFALKAQRLYPVLENIAQMLVVCDVHCSAEQK 338
Query: 60 TCNGELDWYSIMGLNPSTNIEAVKKQYKKMAGLLHPDNNKCVGADGAFHLVSEAWARLSG 119
E++WY I+ L + +KKQ++K A LHPD NK GA+ AF L+ EA LS
Sbjct: 339 VFGDEINWYGILQLERTAGDAMIKKQFRKFALQLHPDKNKFAGAEAAFKLIGEAQRVLSD 518
Query: 120 -----SYDMKRN 126
YDMK N
Sbjct: 519 REKRTRYDMKLN 554
>TC85064 similar to GP|15529208|gb|AAK97698.1 At2g01710/T8O11.12
{Arabidopsis thaliana}, partial (29%)
Length = 645
Score = 85.9 bits (211), Expect = 5e-17
Identities = 44/107 (41%), Positives = 70/107 (65%), Gaps = 1/107 (0%)
Frame = +1
Query: 13 ENAEKRFSHRDFVGAKSYALKAKTLCPGLEGISQLVTTFEVYIASQVTCNGELDWYSIMG 72
E+ + +RD +G+K +A+ A+ P LEG Q++ +V IAS+ N DWYSI+
Sbjct: 130 ESPKSLLQNRDLMGSKEFAILAQETEPLLEGSDQILAIIDVLIASEKRVNNNPDWYSILQ 309
Query: 73 LNP-STNIEAVKKQYKKMAGLLHPDNNKCVGADGAFHLVSEAWARLS 118
++ S +++ +KKQY+++A LLHPD ++ AD AF LV++AWA LS
Sbjct: 310 IDRRSDDLDLIKKQYRRLALLLHPDKSRFHFADHAFKLVADAWAVLS 450
>TC89822 similar to GP|15529208|gb|AAK97698.1 At2g01710/T8O11.12
{Arabidopsis thaliana}, partial (35%)
Length = 539
Score = 85.1 bits (209), Expect = 8e-17
Identities = 58/170 (34%), Positives = 82/170 (48%), Gaps = 4/170 (2%)
Frame = +2
Query: 1 MEAKKEEALKAIENAEKRFSHRDFVGAKSYALKAKTLCPGLEGISQLVTTFEVYIASQVT 60
M K EA + +E E+ RD G++ A + P LEG Q++ +V A++
Sbjct: 53 MNTSKAEAERLLEIGEELLQKRDLKGSREIANLVQETEPLLEGSDQILAIVDVLEAAEKP 232
Query: 61 CN---GELDWYSIMGLNP-STNIEAVKKQYKKMAGLLHPDNNKCVGADGAFHLVSEAWAR 116
N LDWY+++ ++ S ++ +KKQY+ +A LLHPD N A+ AF LV +AWA
Sbjct: 233 LNLNNHHLDWYAVLQIDRNSQDLNRIKKQYRTLALLLHPDKNPFSYAELAFKLVKDAWAV 412
Query: 117 LSGSYDMKRNAQLGAGNGVNHKGLSSVHASGGNQDTFWTICTSCKVQYEY 166
LS D + AQ G G G FWT C C YEY
Sbjct: 413 LS---DPVQKAQYDKGFEFELLG------KGNGNVNFWTACPYCYHMYEY 535
>AL369948 similar to GP|14587221|db contains ESTs AU108104(C12321)
C26435(C12321)~similar to Arabidopsis thaliana
chromosome 3 F24P17., partial (5%)
Length = 510
Score = 80.9 bits (198), Expect = 2e-15
Identities = 40/107 (37%), Positives = 67/107 (62%), Gaps = 3/107 (2%)
Frame = +2
Query: 1 MEAKKEEALKAIENAEKRFSHRDFVGAKSYALKAKTLCPGLEGISQLVTTFEVYIASQVT 60
M+ KEEAL+A + AEK+ +DF GA+++A KA+ L P LE I+Q++ +V+ +++
Sbjct: 185 MDCNKEEALRAKDIAEKKMESKDFTGARTFAHKAQKLYPDLENIAQMLVVCDVHCSAEQK 364
Query: 61 CNGE---LDWYSIMGLNPSTNIEAVKKQYKKMAGLLHPDNNKCVGAD 104
G +DWY ++ ++ + + + KQY+K A LH D NK GA+
Sbjct: 365 LLGNTNVVDWYKVLQIDRNDHDGIINKQYRKFALQLHTDKNKFAGAE 505
>AJ388944 weakly similar to PIR|H84769|H847 probable DnaJ protein [imported]
- Arabidopsis thaliana, partial (9%)
Length = 448
Score = 72.8 bits (177), Expect = 4e-13
Identities = 42/92 (45%), Positives = 56/92 (60%), Gaps = 2/92 (2%)
Frame = +2
Query: 414 ADVGQVKGD-ICTKTALNVSDNQLEHRKTVPVTITVPDPDFHDFDKDRSEPCFKPKQIWA 472
AD+ VK + ++ ++ S +LE + V D DF+DFDKDR E FK Q+WA
Sbjct: 182 ADLDSVKRKGLRSEKHMDTSGEELE-------VMAVADSDFYDFDKDRVERSFKKGQVWA 340
Query: 473 LYD-EEDGMPRLYCLIREVVSVNPFKINISYL 503
+YD ++DGMPR Y LI E VS NPF IS+L
Sbjct: 341 VYDGDDDGMPRQYVLIDETVSANPFNXMISWL 436
>TC80113 weakly similar to GP|21554000|gb|AAM63081.1 unknown {Arabidopsis
thaliana}, partial (17%)
Length = 1166
Score = 65.9 bits (159), Expect = 5e-11
Identities = 39/116 (33%), Positives = 63/116 (53%), Gaps = 2/116 (1%)
Frame = +2
Query: 5 KEEALKAIENAEKRFSHRDFVGAKSYALKAKTLCPGLEGISQLVTTFEVYIASQVTCNGE 64
+ EA + + A K S RD GA+S+A++A+ P + L+ + +A + N
Sbjct: 164 RAEAERWLYTANKLLSARDLHGARSFAIRARESDPTFDASELLLAVIDTLLAGESRINDH 343
Query: 65 -LDWYSIMG-LNPSTNIEAVKKQYKKMAGLLHPDNNKCVGADGAFHLVSEAWARLS 118
DWY I+ L +TNI+ + QY+++A LL P+ N + AF LV +AW+ LS
Sbjct: 344 HRDWYGILQILRYTTNIDHIANQYRRLALLLDPNRNPFAFSGHAFSLVHDAWSVLS 511
Score = 43.9 bits (102), Expect = 2e-04
Identities = 31/105 (29%), Positives = 48/105 (45%), Gaps = 3/105 (2%)
Frame = +2
Query: 131 AGNGVNHKGLSSVHASGGNQDTFWTICTSCKVQYEYLRKYVNKKLSCKNCRGIF--IALE 188
AG ++S + GN +FWT+C C V YEY ++Y + L C++CR F + +
Sbjct: 740 AGETTRQTRIASAAETVGNI-SFWTLCPYCYVHYEYPKEYEDCTLRCQSCRRGFHAVVIR 916
Query: 189 TAPANG-SFPYSPWSYGSSSGYGSHSYDGVTYVPTNGAYFNGNGV 232
+ P N + W + G+ S D NGA N N +
Sbjct: 917 SPPVNEIDSSFCTWGF-FPLGFSGDSKD------LNGASSNWNPI 1030
>TC93256
Length = 613
Score = 59.7 bits (143), Expect = 4e-09
Identities = 37/79 (46%), Positives = 48/79 (59%), Gaps = 6/79 (7%)
Frame = +3
Query: 18 RFSHRDFVGA-----KSYALKAKTLCPGLEGISQLVTTFEVYIASQVTCNGELD-WYSIM 71
R RDF+ K+YALKAK LCP LEGISQ+V+TF+V+IAS+ NGE+ +
Sbjct: 363 RMPRRDFLKEILQEQKNYALKAKELCPELEGISQMVSTFDVHIASEFRHNGEVRLLFGFS 542
Query: 72 GLNPSTNIEAVKKQYKKMA 90
N K+QYKK+A
Sbjct: 543 D*NLLRIKRLSKRQYKKLA 599
Score = 51.6 bits (122), Expect = 1e-06
Identities = 39/95 (41%), Positives = 50/95 (52%), Gaps = 11/95 (11%)
Frame = +2
Query: 1 MEAKKEEALKAIENAEKRFSHRDFVGAKSYALKAKTLCPGLEGISQLVTTFEVYIASQVT 60
MEA KEEALKAIENAEKRFS RDF GA+ + + G YI++ V
Sbjct: 326 MEANKEEALKAIENAEKRFSQRDFAGAEKLCFEG*RALS*VGG----------YISNGVN 475
Query: 61 --CN----GELDWYS-----IMGLNPSTNIEAVKK 84
C+ + W S ++GL P+ + EAVKK
Sbjct: 476 I*CSYCF*VQT*WRSKIIIRVLGLKPTADKEAVKK 580
>TC90697 homologue to GP|12322856|gb|AAG51418.1 hypothetical protein
contains DnaJ motif: prokaryotic heat shock protein
motif; 22764-26261, partial (1%)
Length = 630
Score = 57.4 bits (137), Expect = 2e-08
Identities = 36/114 (31%), Positives = 61/114 (52%), Gaps = 3/114 (2%)
Frame = +3
Query: 559 IYPKCGDVWAVYRNWSTDWNRSTPDEVRHQYDMVEVLDDYSEELGLCVSPLIKLDGFKTV 618
IYPK G++WA+Y+ + + S R + +VEVL D + + + V L++ + +
Sbjct: 45 IYPKKGEIWALYKEQNYELISSNQGRGRSECHIVEVLADSDKSIQVVV--LVRHSRSQPI 218
Query: 619 YKR---NADKSAIRYIPRREMLRFSHQVPSWLLKGEEASNLPDKCWDLDPAATP 669
+K K++I I R ++ RFSHQ+P + GE+ L CW DP++ P
Sbjct: 219 FKPPIIRRSKTSIIEILREDVGRFSHQIPVFKHNGEDDVQLRG-CWVADPSSIP 377
>BQ138754 weakly similar to GP|14587221|db contains ESTs AU108104(C12321)
C26435(C12321)~similar to Arabidopsis thaliana
chromosome 3 F24P17., partial (4%)
Length = 639
Score = 53.9 bits (128), Expect = 2e-07
Identities = 32/118 (27%), Positives = 60/118 (50%)
Frame = -2
Query: 383 EKARTEIRKKLEEMKLASETAAAVIEGKKSQADVGQVKGDICTKTALNVSDNQLEHRKTV 442
+ ART R K +++E ++++ K +++ +VK + + S +
Sbjct: 467 KNARTSKRSK---PSMSAEDIVSILKVKVETSNLTEVKDSLDDMDDCHAS-------AST 318
Query: 443 PVTITVPDPDFHDFDKDRSEPCFKPKQIWALYDEEDGMPRLYCLIREVVSVNPFKINI 500
P +PD F +F+ RS F+ QIWA Y +EDGMP+ Y I++VV+ ++++
Sbjct: 317 PEAFEIPDAQFFNFETGRSLDKFQVGQIWAFYSDEDGMPKYYGQIKKVVTGPTIELHV 144
>TC77865 similar to GP|17473565|gb|AAL38258.1 dnaJ-like protein {Arabidopsis
thaliana}, partial (70%)
Length = 1393
Score = 53.5 bits (127), Expect = 3e-07
Identities = 32/100 (32%), Positives = 53/100 (53%), Gaps = 4/100 (4%)
Frame = +1
Query: 54 YIASQVTCNGEL----DWYSIMGLNPSTNIEAVKKQYKKMAGLLHPDNNKCVGADGAFHL 109
Y QV+ E+ ++Y I+G+ S ++ V+K Y+K++ +HPD NK GA+ AF L
Sbjct: 427 YTEEQVSIIREIKRKKNYYDILGVEKSCTVDDVRKSYRKLSLKVHPDKNKAPGAEEAFKL 606
Query: 110 VSEAWARLSGSYDMKRNAQLGAGNGVNHKGLSSVHASGGN 149
VS+A+ LS ++ G V + ++ A G N
Sbjct: 607 VSKAFQCLSNEESKRKYDVSGEDEVVYERRAAARPARGFN 726
>TC77858 similar to GP|20197886|gb|AAD22362.2 putative DnaJ protein
{Arabidopsis thaliana}, partial (78%)
Length = 1869
Score = 45.4 bits (106), Expect = 7e-05
Identities = 29/97 (29%), Positives = 47/97 (47%)
Frame = +3
Query: 57 SQVTCNGELDWYSIMGLNPSTNIEAVKKQYKKMAGLLHPDNNKCVGADGAFHLVSEAWAR 116
S++ + D+Y+++G++ ++ +K Y+K+A HPD NK GA+ F +S A+
Sbjct: 471 SRLIVRADADYYTVLGVSKNSTKSEIKTAYRKLARNYHPDVNKDPGAEEKFKEISNAYEV 650
Query: 117 LSGSYDMKRNAQLGAGNGVNHKGLSSVHASGGNQDTF 153
LS D KR+ G GL GG F
Sbjct: 651 LSD--DEKRSIYDKYGEA----GLKGSQGFGGGMGGF 743
>BG456490 weakly similar to GP|22296361|dbj AHM1(AT hook-containing MAR
binding protein1)-like protein {Oryza sativa (japonica
cultivar-group)}, partial (6%)
Length = 531
Score = 45.1 bits (105), Expect = 9e-05
Identities = 34/110 (30%), Positives = 45/110 (40%), Gaps = 1/110 (0%)
Frame = +3
Query: 76 STNIEAVKKQYKKMAGLLHPDNN-KCVGADGAFHLVSEAWARLSGSYDMKRNAQLGAGNG 134
S N + ++ Y K A LL P ++ K D A V EAW LS D + G G
Sbjct: 6 SVNRDLARQHYAKFAILLDPTSSEKFPFQDEALARVREAWHVLS---DPGKRTVYDRGGG 176
Query: 135 VNHKGLSSVHASGGNQDTFWTICTSCKVQYEYLRKYVNKKLSCKNCRGIF 184
V + FWT C C Y+Y +KY + L C+ C F
Sbjct: 177 V----------TAATTAAFWTACPYCWNLYQYEKKYEDCSLMCQTCMKTF 296
>TC85315 similar to GP|9758827|dbj|BAB09499.1 gene_id:MTH12.18~unknown
protein {Arabidopsis thaliana}, partial (39%)
Length = 713
Score = 43.1 bits (100), Expect = 3e-04
Identities = 22/74 (29%), Positives = 37/74 (49%)
Frame = +2
Query: 68 YSIMGLNPSTNIEAVKKQYKKMAGLLHPDNNKCVGADGAFHLVSEAWARLSGSYDMKRNA 127
Y ++GL+PS ++ +KK Y+K+A HPD NK A F + A+ L S ++
Sbjct: 332 YEVLGLSPSASVNEIKKAYRKLALKYHPDVNKEDNAQEKFLRIKHAYNTLLNSSSRRKYD 511
Query: 128 QLGAGNGVNHKGLS 141
G+ + + S
Sbjct: 512 SGNRGSNSSQRSQS 553
>BF631720 weakly similar to GP|10798648|em putative DNAJ protein {Nicotiana
tabacum}, partial (25%)
Length = 630
Score = 42.0 bits (97), Expect = 8e-04
Identities = 26/84 (30%), Positives = 45/84 (52%), Gaps = 1/84 (1%)
Frame = +3
Query: 64 ELDWYSIMGLNPSTNIEAVKKQYKKMAGLLHPDNNK-CVGADGAFHLVSEAWARLSGSYD 122
E ++Y+ +G+ P+ + + +KK Y+K+A HPD N A+ F +SEA+ LS
Sbjct: 57 ETEYYTRLGVQPNASQDEIKKAYRKLAVKYHPDKNPGNKEAEEMFKKISEAYEVLSDEQK 236
Query: 123 MKRNAQLGAGNGVNHKGLSSVHAS 146
+ Q G +G+ + G + AS
Sbjct: 237 RQMYDQYGE-DGLKNSGYNPGDAS 305
>TC89246 homologue to PIR|T06594|T06594 heat shock protein dnaJ - garden
pea, partial (50%)
Length = 1119
Score = 40.4 bits (93), Expect = 0.002
Identities = 28/82 (34%), Positives = 41/82 (49%), Gaps = 1/82 (1%)
Frame = +1
Query: 66 DWYSIMGLNPSTNIEAVKKQYKKMAGLLHPDNNKCVGADGAFHLVSEAWARLSGSYDMKR 125
D+YS +G+ S + +K Y+++A HPD NK GA F +S A+ LS
Sbjct: 469 DYYSTLGVPKSATGKEIKAAYRRLARQYHPDVNKEPGATDKFKEISNAYEVLSDDKKRAL 648
Query: 126 NAQLG-AGNGVNHKGLSSVHAS 146
Q G AG + G SS +A+
Sbjct: 649 YDQYGEAGVKSSVGGGSSAYAT 714
Database: MTGI
Posted date: Oct 22, 2004 3:39 PM
Number of letters in database: 27,044,181
Number of sequences in database: 36,976
Lambda K H
0.315 0.132 0.397
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 21,952,085
Number of Sequences: 36976
Number of extensions: 332867
Number of successful extensions: 1293
Number of sequences better than 10.0: 84
Number of HSP's better than 10.0 without gapping: 1245
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 1273
length of query: 682
length of database: 9,014,727
effective HSP length: 103
effective length of query: 579
effective length of database: 5,206,199
effective search space: 3014389221
effective search space used: 3014389221
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.3 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 42 (22.0 bits)
S2: 62 (28.5 bits)
Medicago: description of AC140670.15


