

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC140550.8 + phase: 0 /pseudo
(496 letters)
Database: MTGI
36,976 sequences; 27,044,181 total letters
Searching..................................................done
Score E
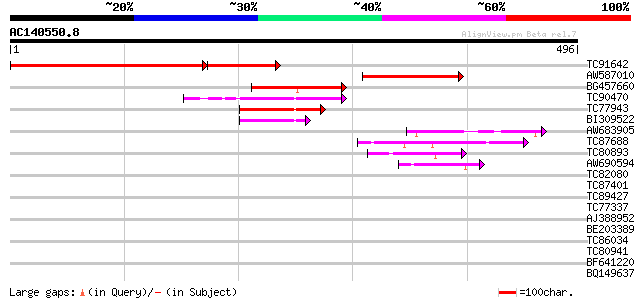
Sequences producing significant alignments: (bits) Value
TC91642 similar to PIR|E96613|E96613 hypothetical protein T15M6.... 345 e-129
AW587010 similar to PIR|E96613|E966 hypothetical protein T15M6.2... 181 6e-46
BG457660 similar to PIR|T00472|T00 probable RING3 protein [impor... 73 2e-13
TC90470 weakly similar to GP|9294219|dbj|BAB02121.1 gb|AAF01563.... 65 5e-11
TC77943 similar to GP|10179602|gb|AAG13810.1 PSTVd RNA-binding p... 62 4e-10
BI309522 similar to PIR|A86198|A86 hypothetical protein [importe... 52 5e-07
AW683905 weakly similar to GP|9757854|dbj| contains similarity t... 43 2e-04
TC87688 similar to GP|10177535|dbj|BAB10930. gene_id:K1F13.21~un... 43 3e-04
TC80893 similar to GP|4557063|gb|AAD22502.1| expressed protein {... 42 5e-04
AW690594 similar to GP|23498163|emb hypothetical protein {Plasmo... 42 5e-04
TC82080 similar to GP|9757854|dbj|BAB08488.1 contains similarity... 41 0.001
TC87401 similar to PIR|B84587|B84587 probable glutaredoxin [impo... 41 0.001
TC89427 weakly similar to GP|4557063|gb|AAD22502.1| expressed pr... 40 0.002
TC77337 weakly similar to GP|4019275|gb|AAC95573.1| orf 48 {Atel... 40 0.002
AJ388952 similar to PIR|T04526|T045 hypothetical protein F16A16.... 40 0.002
BE203389 weakly similar to PIR|T06754|T0 DNA-directed RNA polyme... 39 0.005
TC86034 similar to GP|6815067|dbj|BAA90354.1 ESTs AU082210(C5365... 38 0.008
TC80941 weakly similar to PIR|T39903|T39903 serine-rich protein ... 38 0.010
BF641220 weakly similar to PIR|G86203|G86 probable N-arginine di... 37 0.017
BQ149637 similar to PIR|D85135|D85 hypothetical protein AT4g1261... 33 0.19
>TC91642 similar to PIR|E96613|E96613 hypothetical protein T15M6.22
[imported] - Arabidopsis thaliana, partial (11%)
Length = 949
Score = 345 bits (884), Expect(2) = e-129
Identities = 173/173 (100%), Positives = 173/173 (100%)
Frame = +2
Query: 1 MKRKRGHKKGRPKGNTNPVSVTLTNEQEEEEEEEEETSAFDVGSKEDNNSGMEVDTPSST 60
MKRKRGHKKGRPKGNTNPVSVTLTNEQEEEEEEEEETSAFDVGSKEDNNSGMEVDTPSST
Sbjct: 236 MKRKRGHKKGRPKGNTNPVSVTLTNEQEEEEEEEEETSAFDVGSKEDNNSGMEVDTPSST 415
Query: 61 GTDQHSNLANINPDGSIRKPVGRVKVKLKTSKMLDSQPNSSDAPSQSDTDKSSQQQHGLE 120
GTDQHSNLANINPDGSIRKPVGRVKVKLKTSKMLDSQPNSSDAPSQSDTDKSSQQQHGLE
Sbjct: 416 GTDQHSNLANINPDGSIRKPVGRVKVKLKTSKMLDSQPNSSDAPSQSDTDKSSQQQHGLE 595
Query: 121 RHGVNADRIEDSVTSFPDLKYGASYKKAGSIKIKSSKVLGLNADQTSKPLPIS 173
RHGVNADRIEDSVTSFPDLKYGASYKKAGSIKIKSSKVLGLNADQTSKPLPIS
Sbjct: 596 RHGVNADRIEDSVTSFPDLKYGASYKKAGSIKIKSSKVLGLNADQTSKPLPIS 754
Score = 135 bits (341), Expect(2) = e-129
Identities = 64/64 (100%), Positives = 64/64 (100%)
Frame = +3
Query: 174 SEIIHPKERKMPPLNPRYNKHELDTSLMIIRKVMKMDAAEPFNVPVNPEALGIPDYFDII 233
SEIIHPKERKMPPLNPRYNKHELDTSLMIIRKVMKMDAAEPFNVPVNPEALGIPDYFDII
Sbjct: 756 SEIIHPKERKMPPLNPRYNKHELDTSLMIIRKVMKMDAAEPFNVPVNPEALGIPDYFDII 935
Query: 234 DTPM 237
DTPM
Sbjct: 936 DTPM 947
>AW587010 similar to PIR|E96613|E966 hypothetical protein T15M6.22 [imported]
- Arabidopsis thaliana, partial (3%)
Length = 542
Score = 181 bits (459), Expect = 6e-46
Identities = 87/89 (97%), Positives = 87/89 (97%)
Frame = +2
Query: 309 DFKQEESMLVESPGGEDSLSNTDELLGTDGDVDDDKGEVTKMETSEKNCSPSDGRHEVNE 368
DFKQEESMLVESPGGEDSLSNTDELLGTDGDVDDDKGEVTKMETSEKNCSPSDGRHEVNE
Sbjct: 275 DFKQEESMLVESPGGEDSLSNTDELLGTDGDVDDDKGEVTKMETSEKNCSPSDGRHEVNE 454
Query: 369 VDDDGMEEENHGPEDVEEDEEEDGEGENE 397
VDDDGMEEENHGP DVEEDEEEDGEG NE
Sbjct: 455 VDDDGMEEENHGP*DVEEDEEEDGEGGNE 541
>BG457660 similar to PIR|T00472|T00 probable RING3 protein [imported] -
Arabidopsis thaliana, partial (34%)
Length = 654
Score = 73.2 bits (178), Expect = 2e-13
Identities = 37/85 (43%), Positives = 52/85 (60%), Gaps = 2/85 (2%)
Frame = +3
Query: 212 AEPFNVPVNPEALGIPDYFDIIDTPMDFGTICSNLEKND--KYMNSEDVYNDVRYIWENC 269
A PF PV+ E LG+ DY++IID PMDFGTI + +E D Y N ++Y DVR I++N
Sbjct: 114 AWPFLEPVDVEGLGLHDYYEIIDKPMDFGTIKNKMEAKDGTGYKNVREIYADVRLIFKNA 293
Query: 270 YKYNNKGDYIVDLMKRVKKNFMKYW 294
KYNN+ + + K + + F W
Sbjct: 294 MKYNNEKHDVHVMAKTLMEKFEDKW 368
>TC90470 weakly similar to GP|9294219|dbj|BAB02121.1
gb|AAF01563.1~gene_id:K17E12.8~similar to unknown
protein {Arabidopsis thaliana}, partial (11%)
Length = 772
Score = 65.5 bits (158), Expect = 5e-11
Identities = 47/142 (33%), Positives = 68/142 (47%)
Frame = +3
Query: 153 IKSSKVLGLNADQTSKPLPISSEIIHPKERKMPPLNPRYNKHELDTSLMIIRKVMKMDAA 212
IK S +LG N + S I PK ++ R H+ T I++ ++ +
Sbjct: 372 IKKSSLLGSNKREPS------GNIEGPKNKRQKL--DRKGSHKCAT---ILKCLISHPYS 518
Query: 213 EPFNVPVNPEALGIPDYFDIIDTPMDFGTICSNLEKNDKYMNSEDVYNDVRYIWENCYKY 272
F PV+P AL IPDYF +I PMD GTI L+KN Y + E+ DVR + N Y
Sbjct: 519 WVFKTPVDPVALNIPDYFTVISHPMDLGTIKFKLDKN-IYYSKEEFAADVRLTFSNAMTY 695
Query: 273 NNKGDYIVDLMKRVKKNFMKYW 294
N + + + K + K F + W
Sbjct: 696 NPPSNDVHLMAKELNKLFERKW 761
>TC77943 similar to GP|10179602|gb|AAG13810.1 PSTVd RNA-binding protein
Virp1a {Lycopersicon esculentum}, partial (43%)
Length = 1783
Score = 62.4 bits (150), Expect = 4e-10
Identities = 33/75 (44%), Positives = 45/75 (60%)
Frame = +3
Query: 202 IIRKVMKMDAAEPFNVPVNPEALGIPDYFDIIDTPMDFGTICSNLEKNDKYMNSEDVYND 261
I++K+MK F+ PV+P AL + DYFDII PMD GT+ S L KN Y + +D
Sbjct: 660 ILQKLMKTKIGWIFSSPVDPVALNLHDYFDIIKHPMDLGTVKSKLAKN-AYSTPAEFADD 836
Query: 262 VRYIWENCYKYNNKG 276
V+ ++N YN KG
Sbjct: 837 VKLTFKNALTYNPKG 881
>BI309522 similar to PIR|A86198|A86 hypothetical protein [imported] -
Arabidopsis thaliana, partial (17%)
Length = 729
Score = 52.0 bits (123), Expect = 5e-07
Identities = 28/62 (45%), Positives = 37/62 (59%)
Frame = +1
Query: 202 IIRKVMKMDAAEPFNVPVNPEALGIPDYFDIIDTPMDFGTICSNLEKNDKYMNSEDVYND 261
++ K+MK FN PV+ E LG+ DYF II PMD GT+ S L KN Y + ++ D
Sbjct: 538 LLEKLMKHKHGWVFNTPVDVERLGLHDYFIIISQPMDLGTVKSRLSKN-WYKSPKEFAED 714
Query: 262 VR 263
VR
Sbjct: 715 VR 720
>AW683905 weakly similar to GP|9757854|dbj| contains similarity to apoptosis
antagonizing transcription factor~gene_id:MFB13.10,
partial (12%)
Length = 492
Score = 43.1 bits (100), Expect = 2e-04
Identities = 38/128 (29%), Positives = 56/128 (43%), Gaps = 6/128 (4%)
Frame = +3
Query: 348 TKMETSE----KNCSPSDGRHEVNEVDDDGMEEENHGPEDVEEDEEEDGEGENEEEIEMN 403
TKM +S KN SD + + DDD +E + ++EE++EEDGE E EE
Sbjct: 72 TKMGSSSAKKSKNMQKSDSDLDEYDDDDDMSYDEVNDDSELEEEKEEDGEDEGEE----- 236
Query: 404 RVKRRMDETLRHGGTLAEESEVGDTAALHDEYKNTQQEGQAAVVQQQKKHKEPQD--KHQ 461
D+ G +E E L EYK+ + Q + KH + +D K Q
Sbjct: 237 ------DDGQEENGWRDDEME-----QLEKEYKDLHHQEQELNTLKNLKHHKDEDLLKGQ 383
Query: 462 KAKLLESL 469
K ++L
Sbjct: 384 AVKSQKAL 407
>TC87688 similar to GP|10177535|dbj|BAB10930. gene_id:K1F13.21~unknown
protein {Arabidopsis thaliana}, partial (46%)
Length = 2077
Score = 42.7 bits (99), Expect = 3e-04
Identities = 37/157 (23%), Positives = 73/157 (45%), Gaps = 7/157 (4%)
Frame = +2
Query: 305 KGSTDFKQEESMLVESPGGEDSLSNTDELLGTDGDVDDDK----GEVTK-METSEKNCSP 359
+G DF++E+ + G ED DE G +DK E+T+ +E E N
Sbjct: 467 EGEDDFEEEDDE--DEEGSEDEDDEEDEKEKVKGGGIEDKFLKIDELTEYLEKEEDNYEK 640
Query: 360 SDGRHEVNE--VDDDGMEEENHGPEDVEEDEEEDGEGENEEEIEMNRVKRRMDETLRHGG 417
+ R E +E +DD +E+ D E+D+++D E E E++ R + G
Sbjct: 641 GEERDEADEDSEEDDELEKAGEFEMDDEDDDDDDEEDEEAEDMGNVRYEDFFGGKNEKGS 820
Query: 418 TLAEESEVGDTAALHDEYKNTQQEGQAAVVQQQKKHK 454
+ ++ + + D+ ++T+Q+ + A ++++ K
Sbjct: 821 --KRKDQLLEVSGDEDDMESTKQKKRTASTYEKQRGK 925
>TC80893 similar to GP|4557063|gb|AAD22502.1| expressed protein {Arabidopsis
thaliana}, partial (39%)
Length = 883
Score = 42.0 bits (97), Expect = 5e-04
Identities = 27/96 (28%), Positives = 50/96 (51%), Gaps = 10/96 (10%)
Frame = +3
Query: 314 ESMLVESPGGEDSLSNTDELLGTDGDVDDDKGEVTKMETSEKNCSPSDGRHEVNEVDD-- 371
E +L P +++ S+T++ + D DDD + + E++ S +G E + DD
Sbjct: 273 EHLLGRKPAYKENKSDTED---DEDDDDDDDVQDEDDDGEEEDYSGDEGEEEGDPEDDPE 443
Query: 372 --------DGMEEENHGPEDVEEDEEEDGEGENEEE 399
DG ++++ G E+ EED E++ + E+EEE
Sbjct: 444 ANGAGGSDDGEDDDDDGDEEDEEDGEDEEDEEDEEE 551
Score = 39.3 bits (90), Expect = 0.004
Identities = 25/90 (27%), Positives = 45/90 (49%)
Frame = +3
Query: 319 ESPGGEDSLSNTDELLGTDGDVDDDKGEVTKMETSEKNCSPSDGRHEVNEVDDDGMEEEN 378
E +D + + D+ G + D D+GE + + + G + + DDDG EE+
Sbjct: 333 EDDDDDDDVQDEDDD-GEEEDYSGDEGEEEGDPEDDPEANGAGGSDDGEDDDDDGDEEDE 509
Query: 379 HGPEDVEEDEEEDGEGENEEEIEMNRVKRR 408
ED E++E+E+ E E++E + KR+
Sbjct: 510 ---EDGEDEEDEEDEEEDDETPQPPAKKRK 590
Score = 28.9 bits (63), Expect = 4.7
Identities = 20/98 (20%), Positives = 44/98 (44%)
Frame = +3
Query: 368 EVDDDGMEEENHGPEDVEEDEEEDGEGENEEEIEMNRVKRRMDETLRHGGTLAEESEVGD 427
E D ++E+ +D +DE++DGE E+ E D+ +G +++ E D
Sbjct: 306 ENKSDTEDDEDDDDDDDVQDEDDDGEEEDYSGDEGEEEGDPEDDPEANGAGGSDDGEDDD 485
Query: 428 TAALHDEYKNTQQEGQAAVVQQQKKHKEPQDKHQKAKL 465
++ ++ + E ++ + +P K +K +L
Sbjct: 486 DDGDEEDEEDGEDEEDEEDEEEDDETPQPPAKKRK*EL 599
>AW690594 similar to GP|23498163|emb hypothetical protein {Plasmodium
falciparum 3D7}, partial (10%)
Length = 633
Score = 42.0 bits (97), Expect = 5e-04
Identities = 24/80 (30%), Positives = 45/80 (56%), Gaps = 5/80 (6%)
Frame = +3
Query: 341 DDDKGEVTKMETSEKNCSPSDGRHEVNEVDDDGMEEENHGPEDVEEDEEEDGEGENE--- 397
++D+GEV + E +++ + H+ E +++ EEE+ ++ EE+EEE+ E E +
Sbjct: 60 EEDEGEVEEEE--DEHDEEEEDEHDEEEEEEEEEEEEDEHDDEEEEEEEEEEEEEEDDDE 233
Query: 398 --EEIEMNRVKRRMDETLRH 415
EE E++R+ D L H
Sbjct: 234 EGEEDEIDRISTLPDPLLLH 293
Score = 37.4 bits (85), Expect = 0.013
Identities = 26/70 (37%), Positives = 30/70 (42%)
Frame = +3
Query: 365 EVNEVDDDGMEEENHGPEDVEEDEEEDGEGENEEEIEMNRVKRRMDETLRHGGTLAEESE 424
E NE +D+G EE D EE++E D E E EEE E DE EE E
Sbjct: 48 EHNEEEDEGEVEEEEDEHDEEEEDEHDEEEEEEEEEE--EEDEHDDEEEEEEEEEEEEEE 221
Query: 425 VGDTAALHDE 434
D DE
Sbjct: 222 DDDEEGEEDE 251
Score = 34.3 bits (77), Expect = 0.11
Identities = 21/60 (35%), Positives = 29/60 (48%), Gaps = 7/60 (11%)
Frame = +3
Query: 349 KMETSEKNCSPSDGRHEVNEVDDDGMEEENHGPEDVEEDEEEDG-------EGENEEEIE 401
+ + E N +G E E + D EE+ H E+ EE+EEE+ E E EEE E
Sbjct: 33 RRKVEEHNEEEDEGEVEEEEDEHDEEEEDEHDEEEEEEEEEEEEDEHDDEEEEEEEEEEE 212
Score = 30.4 bits (67), Expect = 1.6
Identities = 16/49 (32%), Positives = 23/49 (46%)
Frame = +3
Query: 26 EQEEEEEEEEETSAFDVGSKEDNNSGMEVDTPSSTGTDQHSNLANINPD 74
E+EEEEEEEEE D +E+ E + G + + + PD
Sbjct: 132 EEEEEEEEEEEDEHDDEEEEEEEEEEEEEEDDDEEGEEDEIDRISTLPD 278
>TC82080 similar to GP|9757854|dbj|BAB08488.1 contains similarity to
apoptosis antagonizing transcription
factor~gene_id:MFB13.10, partial (37%)
Length = 908
Score = 41.2 bits (95), Expect = 0.001
Identities = 35/123 (28%), Positives = 54/123 (43%), Gaps = 2/123 (1%)
Frame = +2
Query: 349 KMETSEKNCSPSDGRHEVNEVDDDGMEEENHGPEDVEEDEEEDGEGENEEEIEMNRVKRR 408
KM+ S+ + D NEV+DD +VEE+EEED EG +EEE + + + R
Sbjct: 47 KMQKSDSDLDEYDDDMSFNEVNDDS---------EVEEEEEEDDEGSDEEEEDERQEESR 199
Query: 409 MDETLRHGGTLAEESEVGDTAALHDEYKNTQQEGQAAVVQQQKKHKEPQD--KHQKAKLL 466
+ +E E L EY + + Q + KH + +D K Q K
Sbjct: 200 WKD---------DEME-----QLEKEYMDLHHQEQELNTLKNLKHHKDEDLLKGQAVKTQ 337
Query: 467 ESL 469
++L
Sbjct: 338 KAL 346
>TC87401 similar to PIR|B84587|B84587 probable glutaredoxin [imported] -
Arabidopsis thaliana, partial (63%)
Length = 914
Score = 40.8 bits (94), Expect = 0.001
Identities = 21/53 (39%), Positives = 31/53 (57%)
Frame = -2
Query: 374 MEEENHGPEDVEEDEEEDGEGENEEEIEMNRVKRRMDETLRHGGTLAEESEVG 426
M P++ EE+EEE+GEG+ EE++ +V R DE L + EE E+G
Sbjct: 292 MVSSRREPKEEEEEEEEEGEGDEEEDMARGKVGARRDEAL---VMMGEEVEMG 143
>TC89427 weakly similar to GP|4557063|gb|AAD22502.1| expressed protein
{Arabidopsis thaliana}, partial (50%)
Length = 753
Score = 40.0 bits (92), Expect = 0.002
Identities = 25/83 (30%), Positives = 41/83 (49%), Gaps = 8/83 (9%)
Frame = +1
Query: 319 ESPGGEDSLSNTDELLGTD-GDVDDDKGEVTKMETSEKNCSPSD-------GRHEVNEVD 370
E+ G ++ + DE D D DDD E + +++ P D G + ++ D
Sbjct: 286 ENKDGSETEDDDDEDDDDDVNDEDDDNDEDFSGDEDDEDADPEDDPVPNGAGGSDDDDED 465
Query: 371 DDGMEEENHGPEDVEEDEEEDGE 393
DD +++N ED +EDEEED +
Sbjct: 466 DDDDDDDNDDGEDEDEDEEEDDD 534
Score = 34.3 bits (77), Expect = 0.11
Identities = 24/88 (27%), Positives = 37/88 (41%), Gaps = 6/88 (6%)
Frame = +1
Query: 320 SPGGEDSLSNTDELLGTDGDVDDDKGEVTKMETSEKNCSPSDGRHEVNEVDDDGMEEENH 379
S G N D D D +DD +V + D E + +DD +
Sbjct: 262 SKGNHTCEENKDGSETEDDDDEDDDDDVNDEDDDNDEDFSGDEDDEDADPEDDPVPNGAG 441
Query: 380 GPEDVEEDEE------EDGEGENEEEIE 401
G +D +ED++ +DGE E+E+E E
Sbjct: 442 GSDDDDEDDDDDDDDNDDGEDEDEDEEE 525
Score = 28.9 bits (63), Expect = 4.7
Identities = 14/47 (29%), Positives = 26/47 (54%), Gaps = 1/47 (2%)
Frame = +1
Query: 358 SPSDGRHEVNEVDDDGME-EENHGPEDVEEDEEEDGEGENEEEIEMN 403
SP DGR ++++ EEN + E+D++ED + + +E + N
Sbjct: 226 SPMDGRFPLDQLSKGNHTCEENKDGSETEDDDDEDDDDDVNDEDDDN 366
>TC77337 weakly similar to GP|4019275|gb|AAC95573.1| orf 48 {Ateline
herpesvirus 3}, partial (8%)
Length = 986
Score = 40.0 bits (92), Expect = 0.002
Identities = 25/77 (32%), Positives = 40/77 (51%)
Frame = +3
Query: 323 GEDSLSNTDELLGTDGDVDDDKGEVTKMETSEKNCSPSDGRHEVNEVDDDGMEEENHGPE 382
GED LS+ D G G+ ++K K + +G E D+DG +++ E
Sbjct: 480 GEDELSSEDG--GGYGNNSNNKSNSKKAPEGGAGGADENGEEED---DEDGDDQDEDDDE 644
Query: 383 DVEEDEEEDGEGENEEE 399
D ++D+EE+G E+EEE
Sbjct: 645 DEDDDDEEEGGEEDEEE 695
>AJ388952 similar to PIR|T04526|T045 hypothetical protein F16A16.160 -
Arabidopsis thaliana, partial (24%)
Length = 591
Score = 40.0 bits (92), Expect = 0.002
Identities = 21/52 (40%), Positives = 31/52 (59%)
Frame = -3
Query: 375 EEENHGPEDVEEDEEEDGEGENEEEIEMNRVKRRMDETLRHGGTLAEESEVG 426
E + E+ EE+EEE+GEG+ EE++ +V R DE L + EE E+G
Sbjct: 283 EPKEEEEEEEEEEEEEEGEGDEEEDMARGKVGARRDEAL---VMMGEEVEMG 137
>BE203389 weakly similar to PIR|T06754|T0 DNA-directed RNA polymerase I 190K
chain homolog F15B8.150 - Arabidopsis thaliana, partial
(6%)
Length = 652
Score = 38.9 bits (89), Expect = 0.005
Identities = 51/227 (22%), Positives = 88/227 (38%), Gaps = 9/227 (3%)
Frame = +2
Query: 186 PLNPRYNKHELDTSLMIIRKVMKMDAAEPFNVPVNPEALGIPDYFDIIDTPMDFGTICS- 244
PL P + + ++K+ D E V V P A+ G ICS
Sbjct: 29 PLRPNKSMEDAIRLADKMKKITVADIIESMKVSVVPVAV-------------KEGRICSI 169
Query: 245 -----NLEKNDKYMNSEDVYNDVRYIWENCYKYNNKGDYIVDLMKRVKKNFMKYWAAAGL 299
L K Y DV + WE + ++ +L ++ + +G+
Sbjct: 170 YKLTMKLHKPKHYPKYTDVTLED---WEETLRVG----FVRELEDAIENHISLLARISGI 328
Query: 300 YSEQPKGSTDFKQEESMLVESPGGEDSLSNTDELLGTDGDVDDDKGEVTKMETSEKNCSP 359
Q K ++ + ES ++ ++ D+ +G D ++D + K +
Sbjct: 329 KDFQGKSNSSNGLDNDHSNESASNQNGQTDDDDEVGDTEDAEEDGFDAQKSK-------- 484
Query: 360 SDGRHEVNEVD-DDGMEEENHGPEDVEEDE-EEDG-EGENEEEIEMN 403
+ +EVD DDG EEE H E E+ E EDG + E++ +E+N
Sbjct: 485 ---QRATDEVDYDDGPEEETHDGEKSEDVEVSEDGKDDEDDNGVEVN 616
>TC86034 similar to GP|6815067|dbj|BAA90354.1 ESTs AU082210(C53655)
AU056546(S20671) AU056545(S20671) correspond to a
region of the predicted, partial (42%)
Length = 2412
Score = 38.1 bits (87), Expect = 0.008
Identities = 37/136 (27%), Positives = 58/136 (42%), Gaps = 13/136 (9%)
Frame = +3
Query: 355 KNCSPSDGRHEVNEVDDDGMEEENHGPEDVEEDEEEDGEGENEEEI--EMNRVKRRMDET 412
K P + EV E + EEEN E+ + E +GE E EEE+ E ++ DET
Sbjct: 144 KQAEPEPQQEEVAEEEPTTQEEENTPIEEDQPQNEGEGEEEPEEEVAEEEEDDQQGEDET 323
Query: 413 LRH-----GGTLAEESEVGDTAALHD-----EYKNTQQEGQAAVVQQQ-KKHKEPQDKHQ 461
L + GG + + V + + E N ++E + + +K EP K Q
Sbjct: 324 LENQQQQDGGEASNNAAVSEQTVTVEANGGGENNNGEEEEDLELEDEPVEKLLEPFTKEQ 503
Query: 462 KAKLLESLCSENPMLS 477
L++ + P LS
Sbjct: 504 LHALVKQAVEKYPDLS 551
>TC80941 weakly similar to PIR|T39903|T39903 serine-rich protein - fission
yeast (Schizosaccharomyces pombe), partial (15%)
Length = 557
Score = 37.7 bits (86), Expect = 0.010
Identities = 30/139 (21%), Positives = 57/139 (40%)
Frame = +2
Query: 284 KRVKKNFMKYWAAAGLYSEQPKGSTDFKQEESMLVESPGGEDSLSNTDELLGTDGDVDDD 343
KR+ K + + + K +T+ K+E+ + E S+ +E D++
Sbjct: 26 KRMTKMLLNLRMRKWMLMXEVKETTEDKEEKEKIEAEKEAEGSIEEKEE------GNDEE 187
Query: 344 KGEVTKMETSEKNCSPSDGRHEVNEVDDDGMEEENHGPEDVEEDEEEDGEGENEEEIEMN 403
K EV + E + D++ EEE E+ +E E+ + E E EEE + +
Sbjct: 188 KTEVEEKEEGD---------------DNEKSEEETEEKEEGDEKEKTEAETEEEEEADED 322
Query: 404 RVKRRMDETLRHGGTLAEE 422
+ + +E G +E
Sbjct: 323 TIDKSKEEDKAEGSKGGKE 379
>BF641220 weakly similar to PIR|G86203|G86 probable N-arginine dibasic
convertase [imported] - Arabidopsis thaliana, partial
(5%)
Length = 634
Score = 37.0 bits (84), Expect = 0.017
Identities = 19/44 (43%), Positives = 29/44 (65%)
Frame = +2
Query: 358 SPSDGRHEVNEVDDDGMEEENHGPEDVEEDEEEDGEGENEEEIE 401
+P DG +D+D EE++ ED E+DEE+D EGE++E+ E
Sbjct: 338 APKDG-----SIDEDDEEEDD---EDEEDDEEDDDEGEDDEDEE 445
>BQ149637 similar to PIR|D85135|D85 hypothetical protein AT4g12610 [imported]
- Arabidopsis thaliana, partial (15%)
Length = 538
Score = 33.5 bits (75), Expect = 0.19
Identities = 34/126 (26%), Positives = 57/126 (44%), Gaps = 3/126 (2%)
Frame = +3
Query: 305 KGSTDFKQEESMLVESPGGEDSLSNTDELLGTDGDVDDDKGEVTKMETSEKNCSPSDGRH 364
KG D + EE + P +++ + DE GD D+D +V K + E +D
Sbjct: 15 KGEED-EDEEVDRKKRPNKKNTFDD-DEEGPRGGDHDEDDYDVEKGDDWEHEEIFTDDDE 188
Query: 365 EVN---EVDDDGMEEENHGPEDVEEDEEEDGEGENEEEIEMNRVKRRMDETLRHGGTLAE 421
V E +D E PE ++DEEE+ + NEE +++ + + + L G L+E
Sbjct: 189 AVGNDPEEREDLAPEVPAPPEIKQDDEEEEEDEXNEEGGGLSKSGKELKKLLGRTGGLSE 368
Query: 422 ESEVGD 427
+ D
Sbjct: 369 SDDDDD 386
Database: MTGI
Posted date: Oct 22, 2004 3:39 PM
Number of letters in database: 27,044,181
Number of sequences in database: 36,976
Lambda K H
0.305 0.127 0.356
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 13,238,133
Number of Sequences: 36976
Number of extensions: 176316
Number of successful extensions: 2299
Number of sequences better than 10.0: 184
Number of HSP's better than 10.0 without gapping: 1292
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 1795
length of query: 496
length of database: 9,014,727
effective HSP length: 100
effective length of query: 396
effective length of database: 5,317,127
effective search space: 2105582292
effective search space used: 2105582292
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.1 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 43 (21.9 bits)
S2: 60 (27.7 bits)
Medicago: description of AC140550.8


