

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC140549.8 - phase: 0
(122 letters)
Database: MTGI
36,976 sequences; 27,044,181 total letters
Searching..................................................done
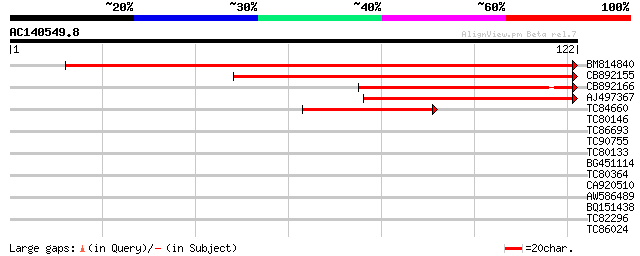
Score E
Sequences producing significant alignments: (bits) Value
BM814840 weakly similar to GP|15128241|db helicase-like protein ... 124 1e-29
CB892155 similar to PIR|D86481|D86 hypothetical protein AAG28292... 111 8e-26
CB892166 similar to GP|20197614|gb unknown protein {Arabidopsis ... 61 1e-10
AJ497367 similar to GP|14140286|gb putative helicase {Oryza sati... 60 2e-10
TC84660 similar to PIR|D86481|D86481 hypothetical protein AAG282... 40 2e-04
TC80146 37 0.001
TC86693 similar to SP|Q9SP35|IM17_ARATH Mitochondrial import inn... 27 2.5
TC90755 similar to SP|P53391|SUT1_STYHA High affinity sulphate t... 26 4.3
TC80133 similar to PIR|S52728|S52728 H+-transporting ATPase (EC ... 26 4.3
BG451114 similar to PIR|D96758|D967 hypothetical protein T18K17.... 26 4.3
TC80364 weakly similar to SP|P50165|TRNH_DATST Tropinone reducta... 26 4.3
CA920510 similar to GP|22094348|gb| putative transcription facto... 26 4.3
AW586489 25 7.3
BQ151438 similar to GP|18539492|gb ATP sulfurylase {Glycine max}... 25 7.3
TC82296 similar to PIR|B96716|B96716 probable serine/threonine k... 25 9.6
TC86024 similar to GP|16604703|gb|AAL24144.1 unknown protein {Ar... 25 9.6
>BM814840 weakly similar to GP|15128241|db helicase-like protein {Oryza
sativa (japonica cultivar-group)}, partial (6%)
Length = 733
Score = 124 bits (310), Expect = 1e-29
Identities = 60/110 (54%), Positives = 80/110 (72%)
Frame = +3
Query: 13 RVISGSNIDEKVFIPRLSLTPSDNRIPFKFKRRQFPISVSFAMIINKSQGQSLEHVGVYL 72
RVI G++ E +IPR++L PS + F+R QFP+ +SFAM INKSQGQ+L VG+YL
Sbjct: 78 RVIGGTHAGEVSYIPRMNLIPSGANVSITFERCQFPLVLSFAMTINKSQGQTLTSVGLYL 257
Query: 73 PSPIFSHGQLYVAISQVTSRGGLKILINDDDDDDIDVASNVVYREVFRNV 122
P P+F+HGQLYVA+S+V SR GLKILI D++ NVVY+EVF+ +
Sbjct: 258 PRPVFTHGQLYVAVSRVKSRSGLKILITDENGSPSSSTVNVVYQEVFQKI 407
>CB892155 similar to PIR|D86481|D86 hypothetical protein AAG28292.1
[imported] - Arabidopsis thaliana, partial (4%)
Length = 572
Score = 111 bits (277), Expect = 8e-26
Identities = 54/74 (72%), Positives = 64/74 (85%)
Frame = -2
Query: 49 ISVSFAMIINKSQGQSLEHVGVYLPSPIFSHGQLYVAISQVTSRGGLKILINDDDDDDID 108
+ V FAM INKSQGQSL+H+GVYLPS +FSHGQLYVA+S+VTSR GLKILI++DD +D
Sbjct: 370 VEVYFAMTINKSQGQSLKHIGVYLPSSVFSHGQLYVALSRVTSREGLKILISNDDGEDDC 191
Query: 109 VASNVVYREVFRNV 122
V SNVVYREVF N+
Sbjct: 190 VTSNVVYREVFHNL 149
>CB892166 similar to GP|20197614|gb unknown protein {Arabidopsis thaliana},
partial (3%)
Length = 748
Score = 60.8 bits (146), Expect = 1e-10
Identities = 31/47 (65%), Positives = 40/47 (84%)
Frame = -1
Query: 76 IFSHGQLYVAISQVTSRGGLKILINDDDDDDIDVASNVVYREVFRNV 122
+FSHGQLYVAIS+V+SR GLKIL+ D++ D ID +NVVY+ VF+NV
Sbjct: 274 VFSHGQLYVAISRVSSRNGLKILMIDENGDCIDNTTNVVYK-VFQNV 137
>AJ497367 similar to GP|14140286|gb putative helicase {Oryza sativa (japonica
cultivar-group)}, partial (1%)
Length = 543
Score = 60.1 bits (144), Expect = 2e-10
Identities = 28/46 (60%), Positives = 39/46 (83%)
Frame = +1
Query: 77 FSHGQLYVAISQVTSRGGLKILINDDDDDDIDVASNVVYREVFRNV 122
FS+G+LYVA+S+VTSR GLKIL+ +D + ++ SNVVY+EVFRN+
Sbjct: 19 FSNGKLYVAVSRVTSRKGLKILLAHEDGNCMNTTSNVVYKEVFRNL 156
>TC84660 similar to PIR|D86481|D86481 hypothetical protein AAG28292.1
[imported] - Arabidopsis thaliana, partial (1%)
Length = 1009
Score = 40.4 bits (93), Expect = 2e-04
Identities = 19/29 (65%), Positives = 23/29 (78%)
Frame = +2
Query: 64 SLEHVGVYLPSPIFSHGQLYVAISQVTSR 92
S+ G+YLP PIF HG LYVA+S+VTSR
Sbjct: 68 SMATKGMYLPQPIF*HG*LYVALSRVTSR 154
>TC80146
Length = 476
Score = 37.4 bits (85), Expect = 0.001
Identities = 16/35 (45%), Positives = 24/35 (67%)
Frame = -3
Query: 49 ISVSFAMIINKSQGQSLEHVGVYLPSPIFSHGQLY 83
I + + INKS+ QSL ++ +YL PIFSH ++Y
Sbjct: 471 IQIPYFKTINKSR*QSLSYMKIYLSRPIFSHEEMY 367
>TC86693 similar to SP|Q9SP35|IM17_ARATH Mitochondrial import inner membrane
translocase subunit TIM17. [Mouse-ear cress], partial
(62%)
Length = 1047
Score = 26.6 bits (57), Expect = 2.5
Identities = 18/50 (36%), Positives = 24/50 (48%), Gaps = 1/50 (2%)
Frame = -2
Query: 33 PSDNRIPFKFKRRQFPISVSFAMIINKSQGQSLE-HVGVYLPSPIFSHGQ 81
P+ N + FKF + +F S II +Q + E H PSP F H Q
Sbjct: 764 PNPNFLKFKFSKHKFTTS-KL*FIITYTQNSAQEAHQNSPKPSPHFHHHQ 618
>TC90755 similar to SP|P53391|SUT1_STYHA High affinity sulphate
transporter 1. {Stylosanthes hamata}, partial (24%)
Length = 847
Score = 25.8 bits (55), Expect = 4.3
Identities = 14/39 (35%), Positives = 22/39 (55%)
Frame = +2
Query: 49 ISVSFAMIINKSQGQSLEHVGVYLPSPIFSHGQLYVAIS 87
+++SFA I K QGQ L++ G + GQL++ S
Sbjct: 11 VAISFAKIFYK*QGQRLQY*GSF-------QGQLFIGTS 106
>TC80133 similar to PIR|S52728|S52728 H+-transporting ATPase (EC 3.6.1.35) -
kidney bean, partial (20%)
Length = 788
Score = 25.8 bits (55), Expect = 4.3
Identities = 11/25 (44%), Positives = 17/25 (68%)
Frame = -2
Query: 53 FAMIINKSQGQSLEHVGVYLPSPIF 77
F +++ +G S EHV + LPSP+F
Sbjct: 418 FLLVLKVHEG-SFEHVPIELPSPLF 347
>BG451114 similar to PIR|D96758|D967 hypothetical protein T18K17.10
[imported] - Arabidopsis thaliana, partial (55%)
Length = 614
Score = 25.8 bits (55), Expect = 4.3
Identities = 26/94 (27%), Positives = 48/94 (50%), Gaps = 6/94 (6%)
Frame = +2
Query: 26 IPRLSLTPSD-NRIPFKFKRRQFPISVSFAMIINKSQGQSLEHVGVYLPSPIFSHGQLYV 84
I ++L SD N + F + Q I+ + ++ SQ QS++ + LP + Q+
Sbjct: 206 IEEVNLFQSDGNVVHFLNPKVQASIAANTYVVSGPSQVQSIQEL---LPGIM---NQIGA 367
Query: 85 AISQV-----TSRGGLKILINDDDDDDIDVASNV 113
+SQ+ + +GG K+ +++DDD DV + V
Sbjct: 368 DMSQIQKLAESMQGGGKLKDQEEEDDDDDVPTLV 469
>TC80364 weakly similar to SP|P50165|TRNH_DATST Tropinone reductase homolog
(EC 1.1.1.-) (P29X). [Jimsonweed Common thornapple]
{Datura stramonium}, partial (66%)
Length = 811
Score = 25.8 bits (55), Expect = 4.3
Identities = 20/79 (25%), Positives = 37/79 (46%)
Frame = +2
Query: 36 NRIPFKFKRRQFPISVSFAMIINKSQGQSLEHVGVYLPSPIFSHGQLYVAISQVTSRGGL 95
N+ ++K + F ++ S I+ Q + L S IF HG+L + ++
Sbjct: 215 NKCLEEWKNKGFNVTGSVCDILFHEQRKRLMET----VSSIF-HGKLNILVNNAAKPTSK 379
Query: 96 KILINDDDDDDIDVASNVV 114
KI+ N D+D + + +N V
Sbjct: 380 KIIDNTDEDINTTLGTNFV 436
>CA920510 similar to GP|22094348|gb| putative transcription factor {Oryza
sativa (japonica cultivar-group)}, partial (55%)
Length = 573
Score = 25.8 bits (55), Expect = 4.3
Identities = 19/59 (32%), Positives = 28/59 (47%), Gaps = 5/59 (8%)
Frame = -2
Query: 66 EHVGVYLP---SPIFSH--GQLYVAISQVTSRGGLKILINDDDDDDIDVASNVVYREVF 119
+ V VY P SPI++ G+ A ++ G +DDDDD D ++V E F
Sbjct: 458 KEVAVYPPQYHSPIWAR*LGKSEEAS*AISEAGADATAAQEDDDDDDDAVPDLVPGETF 282
>AW586489
Length = 195
Score = 25.0 bits (53), Expect = 7.3
Identities = 9/17 (52%), Positives = 13/17 (75%)
Frame = +3
Query: 93 GGLKILINDDDDDDIDV 109
GG +++ DDDD D+DV
Sbjct: 102 GGSMLVVYDDDDVDVDV 152
>BQ151438 similar to GP|18539492|gb ATP sulfurylase {Glycine max}, partial
(29%)
Length = 643
Score = 25.0 bits (53), Expect = 7.3
Identities = 13/33 (39%), Positives = 16/33 (48%)
Frame = -3
Query: 36 NRIPFKFKRRQFPISVSFAMIINKSQGQSLEHV 68
NR+PF+ KR I S II SQ H+
Sbjct: 626 NRVPFESKRATL*IHRSDVXIIINSQNNRXRHI 528
>TC82296 similar to PIR|B96716|B96716 probable serine/threonine kinase
F23O10.20 [imported] - Arabidopsis thaliana, partial
(6%)
Length = 625
Score = 24.6 bits (52), Expect = 9.6
Identities = 15/35 (42%), Positives = 18/35 (50%)
Frame = +1
Query: 87 SQVTSRGGLKILINDDDDDDIDVASNVVYREVFRN 121
SQVT+ +ND+DDDD S V R RN
Sbjct: 454 SQVTA-------LNDEDDDDGGGFSTFVMRSTVRN 537
>TC86024 similar to GP|16604703|gb|AAL24144.1 unknown protein {Arabidopsis
thaliana}, partial (23%)
Length = 1431
Score = 24.6 bits (52), Expect = 9.6
Identities = 10/26 (38%), Positives = 16/26 (61%)
Frame = +1
Query: 46 QFPISVSFAMIINKSQGQSLEHVGVY 71
+FP S FA IINK + ++ + +Y
Sbjct: 883 RFPFSTLFAAIINKVPAEDMKLIHMY 960
Database: MTGI
Posted date: Oct 22, 2004 3:39 PM
Number of letters in database: 27,044,181
Number of sequences in database: 36,976
Lambda K H
0.320 0.140 0.393
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 2,986,128
Number of Sequences: 36976
Number of extensions: 35865
Number of successful extensions: 283
Number of sequences better than 10.0: 32
Number of HSP's better than 10.0 without gapping: 254
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 277
length of query: 122
length of database: 9,014,727
effective HSP length: 84
effective length of query: 38
effective length of database: 5,908,743
effective search space: 224532234
effective search space used: 224532234
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.8 bits)
S2: 52 (24.6 bits)
Medicago: description of AC140549.8


