

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
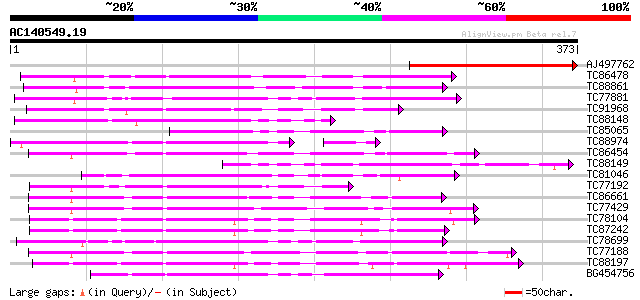
Query= AC140549.19 - phase: 0
(373 letters)
Database: MTGI
36,976 sequences; 27,044,181 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
AJ497762 weakly similar to GP|15528439|emb MEK map kinase kinsae... 226 1e-59
TC86478 similar to GP|3688193|emb|CAA08995.1 MAP3K alpha 1 prote... 192 2e-49
TC88861 weakly similar to GP|17027283|gb|AAL34137.1 putative pro... 167 5e-42
TC77881 homologue to GP|15528439|emb|CAC69137. MEK map kinase ki... 136 1e-32
TC91968 similar to PIR|T02550|T02550 NPK1-related protein kinase... 133 1e-31
TC88148 weakly similar to GP|17979279|gb|AAL49865.1 unknown prot... 130 1e-30
TC85065 similar to GP|1255448|dbj|BAA09057.1 mitogen-activated p... 129 1e-30
TC88974 similar to GP|3819699|emb|CAA08758.1 BnMAP4K alpha2 {Bra... 104 5e-26
TC86454 homologue to GP|4567091|gb|AAD23582.1| SNF-1-like serine... 108 3e-24
TC88149 similar to GP|10177595|dbj|BAB10942. protein kinase-like... 108 4e-24
TC81046 similar to GP|17529280|gb|AAL38867.1 putative protein ki... 106 1e-23
TC77192 similar to SP|P42818|KPK1_ARATH Serine/threonine-protein... 105 4e-23
TC86661 similar to GP|9798599|dbj|BAB11737.1 serine/threonine pr... 104 6e-23
TC77429 similar to GP|15215664|gb|AAK91377.1 AT5g25110/T11H3_120... 103 1e-22
TC78104 homologue to GP|19568098|gb|AAL89456.1 osmotic stress-ac... 102 3e-22
TC87242 homologue to PIR|S56716|S56716 protein kinase SPK-3 (EC ... 101 4e-22
TC78699 weakly similar to PIR|G96761|G96761 probable MAP kinase ... 101 4e-22
TC77188 similar to PIR|T04862|T04862 probable serine/threonine-s... 100 9e-22
TC88197 homologue to GP|6739629|gb|AAF27340.1| abscisic acid-act... 99 2e-21
BG454756 homologue to GP|12718822|db NQK1 MAPKK {Nicotiana tabac... 98 6e-21
>AJ497762 weakly similar to GP|15528439|emb MEK map kinase kinsae {Medicago
sativa subsp. x varia}, partial (5%)
Length = 498
Score = 226 bits (576), Expect = 1e-59
Identities = 107/110 (97%), Positives = 107/110 (97%)
Frame = +1
Query: 264 FLRRCLVRDPKKRSTALELLNHPFLVSTTLTHHKHYSASSPASVLEVHQFEDTYDDDDDD 323
FL LVRDPKKRSTALELLNHPFLVSTTLTHHKHYSASSPASVLEVHQFEDTYDDDDDD
Sbjct: 13 FLEEVLVRDPKKRSTALELLNHPFLVSTTLTHHKHYSASSPASVLEVHQFEDTYDDDDDD 192
Query: 324 DDSDDNDELISPAGGNHFFITKELLCPQGTTMWQLEDSASGNHWITIRSR 373
DDSDDNDELISPAGGNHFFITKELLCPQGTTMWQLEDSASGNHWITIRSR
Sbjct: 193 DDSDDNDELISPAGGNHFFITKELLCPQGTTMWQLEDSASGNHWITIRSR 342
>TC86478 similar to GP|3688193|emb|CAA08995.1 MAP3K alpha 1 protein kinase
{Brassica napus}, partial (52%)
Length = 2266
Score = 192 bits (487), Expect = 2e-49
Identities = 113/294 (38%), Positives = 166/294 (56%), Gaps = 7/294 (2%)
Frame = +2
Query: 8 SEWMKGKMVGCGSFGSVHLAMNKSTGGLFVVKKA-------HSEAGRDALENEVNILNTL 60
S+W KGK++G G+FG V+L N G + +K+ +S+ L E+++LN L
Sbjct: 965 SKWKKGKLLGRGTFGHVYLGFNSENGQMCAIKEVRVGCDDQNSKECLKQLHQEIDLLNQL 1144
Query: 61 NSSPSPYIVQCLGTDYDHQDNQLHVFMEYMSGGSLADVSHKFGGSLNEDVVRVYTRQIVH 120
+ P IVQ LG++ + L V++EY+SGGS+ + ++G E V++ YTRQIV
Sbjct: 1145 SH---PNIVQYLGSELGEES--LSVYLEYVSGGSIHKLLQEYG-PFKEPVIQNYTRQIVS 1306
Query: 121 GLYHLHQHGIVHCDLKCKNVLLASSGNVKLADFGCAKRVKKNMNMNKSSSCIIANGGTPL 180
GL +LH VH D+K N+L+ +G +KLADFG AK + S++ +++ G+P
Sbjct: 1307 GLAYLHGRNTVHRDIKGANILVDPNGEIKLADFGMAKHI-------TSAASMLSFKGSPY 1465
Query: 181 WMAPEVLLMKNNSSINDESRVVDFAAADIWSLGCTVIEMATGRPPWVDDDLISISNPMAA 240
WMAPEV++ N S+ DIWSLGCT+IEMA +PPW S +AA
Sbjct: 1466 WMAPEVVMNTNGYSL----------PVDIWSLGCTLIEMAASKPPW------SQYEGVAA 1597
Query: 241 MFKIACGDGIPQFPIHFSQEGFDFLRRCLVRDPKKRSTALELLNHPFLVSTTLT 294
+FKI +P P H S + +F+ CL RDP R TA +LL HPF+ + T
Sbjct: 1598 IFKIGNSKDMPIIPEHLSNDAKNFIMLCLQRDPSARPTAQKLLEHPFIRDQSAT 1759
>TC88861 weakly similar to GP|17027283|gb|AAL34137.1 putative protein kinase
{Oryza sativa (japonica cultivar-group)}, partial (34%)
Length = 1847
Score = 167 bits (424), Expect = 5e-42
Identities = 109/286 (38%), Positives = 155/286 (54%), Gaps = 7/286 (2%)
Frame = +1
Query: 10 WMKGKMVGCGSFGSVHLAMNKSTGGLFVVKKAH-------SEAGRDALENEVNILNTLNS 62
W KGK++G GSFGSV+ A N TG +K+ S L+ E+ IL L+
Sbjct: 1024 WQKGKLIGRGSFGSVYHATNLETGASCALKEVDLVPDDPKSTDCIKQLDQEIRILGQLHH 1203
Query: 63 SPSPYIVQCLGTDYDHQDNQLHVFMEYMSGGSLADVSHKFGGSLNEDVVRVYTRQIVHGL 122
P IV+ G++ ++L ++MEY+ GSL G + E VVR +TR I+ GL
Sbjct: 1204 ---PNIVEYYGSEV--VGDRLCIYMEYVHPGSLQKFMQDHCGVMTESVVRNFTRHILSGL 1368
Query: 123 YHLHQHGIVHCDLKCKNVLLASSGNVKLADFGCAKRVKKNMNMNKSSSCIIANGGTPLWM 182
+LH +H D+K N+L+ +SG VKLADFG +K + + S ++ G+P WM
Sbjct: 1369 AYLHSTKTIHRDIKGANLLVDASGIVKLADFGVSKILTE-------KSYELSLKGSPYWM 1527
Query: 183 APEVLLMKNNSSINDESRVVDFAAADIWSLGCTVIEMATGRPPWVDDDLISISNPMAAMF 242
APE+++ +++ +E+ A DIWSLGCT+IEM TG+PPW S AMF
Sbjct: 1528 APELMM----AAMKNETNPTVAMAVDIWSLGCTIIEMLTGKPPW------SEFPGHQAMF 1677
Query: 243 KIACGDGIPQFPIHFSQEGFDFLRRCLVRDPKKRSTALELLNHPFL 288
K+ P P S EG DFL +C R+P R +A LL HPF+
Sbjct: 1678 KVL--HRSPDIPKTLSPEGQDFLEQCFQRNPADRPSAAVLLTHPFV 1809
>TC77881 homologue to GP|15528439|emb|CAC69137. MEK map kinase kinsae
{Medicago sativa subsp. x varia}, complete
Length = 1546
Score = 136 bits (343), Expect = 1e-32
Identities = 100/297 (33%), Positives = 151/297 (50%), Gaps = 3/297 (1%)
Frame = +2
Query: 4 IIKGSEWMKGKMVGCGSFGSVHLAMNKSTGGLFVVKKA---HSEAGRDALENEVNILNTL 60
+I SE + +G GS G+V+ +++ G + +K H E+ R + E+ IL +
Sbjct: 449 VIPFSELERLNRIGSGSGGTVYKVVHRINGRAYALKVIYGHHEESVRRQIHREIQILRDV 628
Query: 61 NSSPSPYIVQCLGTDYDHQDNQLHVFMEYMSGGSLADVSHKFGGSLNEDVVRVYTRQIVH 120
+ +V+C YDH + ++ V +EYM GGSL E+ + RQI+
Sbjct: 629 DDVN---VVKCHEM-YDH-NAEIQVLLEYMDGGSLEGKHIP-----QENQLADVARQILR 778
Query: 121 GLYHLHQHGIVHCDLKCKNVLLASSGNVKLADFGCAKRVKKNMNMNKSSSCIIANGGTPL 180
GL +LH+ IVH D+K N+L+ S VK+ADFG + + + M+ SS GT
Sbjct: 779 GLAYLHRRHIVHRDIKPSNLLINSRKQVKIADFGVGRILNQTMDPCNSSV------GTIA 940
Query: 181 WMAPEVLLMKNNSSINDESRVVDFAAADIWSLGCTVIEMATGRPPWVDDDLISISNPMAA 240
+M+PE + N+ IND D A DIWSLG +++E GR P+ + A+
Sbjct: 941 YMSPE----RINTDINDGQ--YDAYAGDIWSLGVSILEFYMGRFPFA----VGRQGDWAS 1090
Query: 241 MFKIACGDGIPQFPIHFSQEGFDFLRRCLVRDPKKRSTALELLNHPFLVSTTLTHHK 297
+ C P+ P S E DF+ RCL RDP +R TA LL+HPFLV H++
Sbjct: 1091LMCAICMSQPPEAPTTASPEFRDFVSRCLQRDPSRRWTASRLLSHPFLVRNGSNHNQ 1261
>TC91968 similar to PIR|T02550|T02550 NPK1-related protein kinase homolog
T26B15.7 - Arabidopsis thaliana, partial (20%)
Length = 662
Score = 133 bits (335), Expect = 1e-31
Identities = 84/250 (33%), Positives = 134/250 (53%), Gaps = 2/250 (0%)
Frame = +2
Query: 12 KGKMVGCGSFGSVHLAMNKSTGGLFVVKKAHSEAGRDALENEVNILNTLNSSPSPYIVQC 71
+G+++G GS +V++A +GG K+ + L+ E IL L P IV
Sbjct: 5 RGRIIGRGSTATVYVADRCHSGGEVFAVKSTELHRSELLKKEQQILTKLKC---PQIVSY 175
Query: 72 LGTD--YDHQDNQLHVFMEYMSGGSLADVSHKFGGSLNEDVVRVYTRQIVHGLYHLHQHG 129
G + +++ ++FMEY G+L+D G + E +V Y RQI+ GL +LH +G
Sbjct: 176 QGCEVTFENGVQLFNLFMEYAPKGTLSDAVRN-GNGMEEAMVGFYARQILLGLNYLHLNG 352
Query: 130 IVHCDLKCKNVLLASSGNVKLADFGCAKRVKKNMNMNKSSSCIIANGGTPLWMAPEVLLM 189
IVHCDLK +NVL+ G K++DFGCA+RV++ + ++ GTP +MAPEV
Sbjct: 353 IVHCDLKGQNVLVTEQG-AKISDFGCARRVEEELVIS----------GTPAFMAPEVARG 499
Query: 190 KNNSSINDESRVVDFAAADIWSLGCTVIEMATGRPPWVDDDLISISNPMAAMFKIACGDG 249
+ A+D+W+LGCT++EM TG+ PW S+P A +++I
Sbjct: 500 EEQG-----------YASDVWALGCTLLEMITGKMPWQ-----GFSDPAAVLYRIGFSGD 631
Query: 250 IPQFPIHFSQ 259
+P+ P + S+
Sbjct: 632 VPEIPDYVSE 661
>TC88148 weakly similar to GP|17979279|gb|AAL49865.1 unknown protein
{Arabidopsis thaliana}, partial (44%)
Length = 657
Score = 130 bits (326), Expect = 1e-30
Identities = 78/214 (36%), Positives = 120/214 (55%), Gaps = 3/214 (1%)
Frame = +1
Query: 4 IIKGSEWMKGKMVGCGSFGSVHLAMNKSTGGLFVVKKAHSEAGRDA-LENEVNILNTLNS 62
I+ +W++G+ VG GSF +V+L + K SE + L+NE ++L+ L S
Sbjct: 79 IVSSMDWVRGEAVGRGSFATVNLVIPKGNSYSTPTAVKTSEVSTSSSLKNEKHVLHQLGS 258
Query: 63 SPSPYIVQCLGTDYDHQDNQ--LHVFMEYMSGGSLADVSHKFGGSLNEDVVRVYTRQIVH 120
I+ C G DY ++ + ++F+EY S G+L+D GG + E +R YTR IV
Sbjct: 259 CQR--IIPCFGDDYTFENGKEYYNLFLEYASAGALSDQVKLNGGRIPEQHIRRYTRSIVE 432
Query: 121 GLYHLHQHGIVHCDLKCKNVLLASSGNVKLADFGCAKRVKKNMNMNKSSSCIIANGGTPL 180
GL H+H++G VHCD+K +N+L+ + G +K+ADFG AK+ +S C GTPL
Sbjct: 433 GLDHIHRNGFVHCDIKLQNILVFNDGEIKIADFGLAKKT----GGKQSLKC----RGTPL 588
Query: 181 WMAPEVLLMKNNSSINDESRVVDFAAADIWSLGC 214
+M+PE S+N+ + ADIW+LGC
Sbjct: 589 FMSPE--------SVNNGEHE---SPADIWALGC 657
>TC85065 similar to GP|1255448|dbj|BAA09057.1 mitogen-activated protein
kinase {Arabidopsis thaliana}, partial (26%)
Length = 789
Score = 129 bits (325), Expect = 1e-30
Identities = 77/184 (41%), Positives = 108/184 (57%), Gaps = 1/184 (0%)
Frame = +2
Query: 106 LNEDVVRVYTRQIVHGLYHLHQHGIVHCDLKCKNVLLASSGNVKLADFGCAKRVKKNMNM 165
L + V YTRQI+HGL +LH IVH D+KC N+L+ ++G+VK+ADFG + +K +
Sbjct: 11 LRDSQVSAYTRQILHGLKYLHDRNIVHRDIKCANILVDANGSVKVADFGLS*AIK----L 178
Query: 166 NKSSSCIIANGGTPLWMAPEVLLMKNNSSINDESRVVDFA-AADIWSLGCTVIEMATGRP 224
N SC GT WMAPEV+ +V + ADIWSLGCTV+EM TG+
Sbjct: 179 NDVKSC----QGTAFWMAPEVV----------RGKVKGYGLPADIWSLGCTVLEMLTGQV 316
Query: 225 PWVDDDLISISNPMAAMFKIACGDGIPQFPIHFSQEGFDFLRRCLVRDPKKRSTALELLN 284
P+ + IS AMF+I G+ +P P S++ DF+ +CL +P R TA +LL+
Sbjct: 317 PYAPMECIS------AMFRIGKGE-LPPVPDTLSRDARDFILQCLKVNPDDRPTAAQLLD 475
Query: 285 HPFL 288
H F+
Sbjct: 476 HKFV 487
>TC88974 similar to GP|3819699|emb|CAA08758.1 BnMAP4K alpha2 {Brassica
napus}, partial (34%)
Length = 851
Score = 104 bits (260), Expect(2) = 5e-26
Identities = 58/189 (30%), Positives = 105/189 (54%), Gaps = 2/189 (1%)
Frame = +3
Query: 1 MCGIIK--GSEWMKGKMVGCGSFGSVHLAMNKSTGGLFVVKKAHSEAGRDALENEVNILN 58
+ G+++ G+ + +++G GSFG V+ +K +K E D +++ ++
Sbjct: 156 LAGLVEATGTRFTSLELIGQGSFGDVYKGFDKELNKEVAIKVIDLEESEDDIDDIQKEIS 335
Query: 59 TLNSSPSPYIVQCLGTDYDHQDNQLHVFMEYMSGGSLADVSHKFGGSLNEDVVRVYTRQI 118
L+ PYI + G+ + +L + MEYM+GGS+AD+ G L+E + R +
Sbjct: 336 VLSQCRCPYITEYYGSFLNQ--TKLWIIMEYMAGGSVADLLQS-GPPLDEMSIAYILRDL 506
Query: 119 VHGLYHLHQHGIVHCDLKCKNVLLASSGNVKLADFGCAKRVKKNMNMNKSSSCIIANGGT 178
+H + +LH G +H D+K N+LL+ +G+VK+ADFG + ++ + ++ K+ GT
Sbjct: 507 LHAVDYLHNEGKIHRDIKAANILLSENGDVKVADFGVSAQLTRTISRRKTFV------GT 668
Query: 179 PLWMAPEVL 187
P WMAPEV+
Sbjct: 669 PFWMAPEVI 695
Score = 30.8 bits (68), Expect(2) = 5e-26
Identities = 18/38 (47%), Positives = 20/38 (52%)
Frame = +1
Query: 207 ADIWSLGCTVIEMATGRPPWVDDDLISISNPMAAMFKI 244
ADIWSLG T IEMA G D +PM +F I
Sbjct: 724 ADIWSLGITAIEMAKGGASLAD------LHPMRVLFII 819
>TC86454 homologue to GP|4567091|gb|AAD23582.1| SNF-1-like serine/threonine
protein kinase {Glycine max}, partial (90%)
Length = 2194
Score = 108 bits (270), Expect = 3e-24
Identities = 88/302 (29%), Positives = 138/302 (45%), Gaps = 5/302 (1%)
Frame = +3
Query: 13 GKMVGCGSFGSVHLAMNKSTGGLFVVK-----KAHSEAGRDALENEVNILNTLNSSPSPY 67
GK +G GSFG V +A + TG +K K + + + E+ IL
Sbjct: 123 GKTLGIGSFGKVKIAEHVLTGHKVAIKILNRRKIKNMEMEEKVRREIKILRLFMHHHIIR 302
Query: 68 IVQCLGTDYDHQDNQLHVFMEYMSGGSLADVSHKFGGSLNEDVVRVYTRQIVHGLYHLHQ 127
+ + + T D ++V MEY+ G L D + G L ED R + +QI+ G+ + H+
Sbjct: 303 LYEVVETPTD-----IYVVMEYVKSGELFDYIVE-KGRLQEDEARSFFQQIISGVEYCHR 464
Query: 128 HGIVHCDLKCKNVLLASSGNVKLADFGCAKRVKKNMNMNKSSSCIIANGGTPLWMAPEVL 187
+ +VH DLK +N+LL S +VK+ADFG + N+ + + + G+P + APEV+
Sbjct: 465 NMVVHRDLKPENLLLDSKWSVKIADFGLS-------NIMRDGHFLKTSCGSPNYAAPEVI 623
Query: 188 LMKNNSSINDESRVVDFAAADIWSLGCTVIEMATGRPPWVDDDLISISNPMAAMFKIACG 247
++ D+WS G + + G P+ D+++ + +FK G
Sbjct: 624 ----------SGKLYAGPEVDVWSCGVILYALLCGTLPFDDENIPN-------LFKKIKG 752
Query: 248 DGIPQFPIHFSQEGFDFLRRCLVRDPKKRSTALELLNHPFLVSTTLTHHKHYSASSPASV 307
GI P H S D + R LV DP KR T E+ HP+ H Y A P
Sbjct: 753 -GIYTLPSHLSPGARDLIPRLLVVDPMKRMTIPEIRQHPWF----QLHLPRYLAVPPPDT 917
Query: 308 LE 309
L+
Sbjct: 918 LQ 923
>TC88149 similar to GP|10177595|dbj|BAB10942. protein kinase-like protein
{Arabidopsis thaliana}, partial (33%)
Length = 881
Score = 108 bits (269), Expect = 4e-24
Identities = 79/237 (33%), Positives = 114/237 (47%), Gaps = 6/237 (2%)
Frame = +2
Query: 141 LLASSGNVKLADFGCAKRVKKNMNMNKSSSCIIANGGTPLWMAPEVLLMKNNSSINDESR 200
L+ + G +K+ADFG AK+ +S C GTPL+M+PE S+N
Sbjct: 2 LVFNDGEIKIADFGLAKKT----GGKQSLKC----RGTPLFMSPE--------SVNHGEH 133
Query: 201 VVDFAAADIWSLGCTVIEMATGRPPWVDDDLISISNPMAAMFKIACGDGIPQFPIHFSQE 260
+ ADIW+LGC V+EM TG+P W +L SN + + +I G+ P P S+E
Sbjct: 134 E---SPADIWALGCAVVEMVTGKPAW---NLEKDSNMWSLLLRIGTGEESPVIPEELSKE 295
Query: 261 GFDFLRRCLVRDPKKRSTALELLNHPFLVSTTLTHHKHYSASSPASVLEVHQFEDTYDDD 320
G DF+ +C V+DP+KR TA LLNHPF+ K+ + SSP + + F D
Sbjct: 296 GKDFVEKCFVKDPRKRWTAEMLLNHPFVEEV-----KNVNESSPRNHFD---FNDWVFSA 451
Query: 321 DDDDDSDDNDELISPAGGNHFFITKELLCPQGTTMWQ------LEDSASGNHWITIR 371
D + E SP + F C +WQ L S+ + WI++R
Sbjct: 452 TDSVPNSPESEESSPWDFDSKF------CSAADRLWQLVTDERLVSSSELDSWISVR 604
>TC81046 similar to GP|17529280|gb|AAL38867.1 putative protein kinase
{Arabidopsis thaliana}, partial (44%)
Length = 921
Score = 106 bits (265), Expect = 1e-23
Identities = 78/254 (30%), Positives = 122/254 (47%), Gaps = 5/254 (1%)
Frame = +2
Query: 48 DALENEVNILNTLNSSPSPYIVQCLGTDYDHQDNQLHVFMEYMSGGSLADV-SHKFGGSL 106
D + EV ++ ++ P+ C T + L V M +MSGGS + F
Sbjct: 17 DGIRREVQTMSLIDH-PNLLRAHCSFT----AGHSLWVVMPFMSGGSCLHIMKSSFPEGF 181
Query: 107 NEDVVRVYTRQIVHGLYHLHQHGIVHCDLKCKNVLLASSGNVKLADFGCAKRVKKNMNMN 166
+E V+ R+++ L +LH HG +H D+K N+LL ++G+VK+ADFG + + +
Sbjct: 182 DEPVIATVLREVLKALVYLHAHGHIHRDVKAGNILLDANGSVKMADFGVSACMFDTGDRQ 361
Query: 167 KSSSCIIANGGTPLWMAPEVLLMKNNSSINDESRVVDFAAADIWSLGCTVIEMATGRPPW 226
+S + + GTP WMAPEV+ + DF ADIWS G T +E+A G P+
Sbjct: 362 RSRNTFV---GTPCWMAPEVM---------QQLHGYDF-KADIWSFGITALELAHGHAPF 502
Query: 227 VDDDLISISNPMAAMFKIACGDGIPQFPI----HFSQEGFDFLRRCLVRDPKKRSTALEL 282
S PM + + + P FS+ + + CLV+DPKKR ++ +L
Sbjct: 503 ------SKYPPMKVLL-MTLQNAPPGLDYERDKRFSKSFKELVATCLVKDPKKRPSSEKL 661
Query: 283 LNHPFLVSTTLTHH 296
L H F T +
Sbjct: 662 LKHHFFKHARATEY 703
>TC77192 similar to SP|P42818|KPK1_ARATH Serine/threonine-protein kinase
AtPK1/AtPK6 (EC 2.7.1.-). [Mouse-ear cress] {Arabidopsis
thaliana}, partial (76%)
Length = 1992
Score = 105 bits (261), Expect = 4e-23
Identities = 72/220 (32%), Positives = 117/220 (52%), Gaps = 7/220 (3%)
Frame = +3
Query: 14 KMVGCGSFGSVHLAMNKSTGGLFVVK-----KAHSEAGRDALENEVNILNTLNSSPSPYI 68
K+VG G+F V+ K T ++ +K K + + ++ E IL + P++
Sbjct: 780 KVVGQGAFAKVYQVRKKGTSEIYAMKVMRKDKIMEKNHAEYMKAEREILTKIEH---PFV 950
Query: 69 VQCLGTDYDHQDN-QLHVFMEYMSGGSLA-DVSHKFGGSLNEDVVRVYTRQIVHGLYHLH 126
VQ Y Q +L++ +++++GG L + H+ G ED+ R+Y +IV + HLH
Sbjct: 951 VQLR---YSFQTKYRLYLVLDFVNGGHLFFQLYHQ--GLFREDLARIYAAEIVSAVSHLH 1115
Query: 127 QHGIVHCDLKCKNVLLASSGNVKLADFGCAKRVKKNMNMNKSSSCIIANGGTPLWMAPEV 186
GI+H DLK +N+L+ + G+V L DFG AK+ +++ N S C GT +MAPE+
Sbjct: 1116SKGIMHRDLKPENILMDADGHVMLTDFGLAKKFEESTRSN--SLC-----GTAEYMAPEI 1274
Query: 187 LLMKNNSSINDESRVVDFAAADIWSLGCTVIEMATGRPPW 226
+L K + AAD WS+G + EM TG+PP+
Sbjct: 1275ILGKGHDK-----------AADWWSVGILLFEMLTGKPPF 1361
>TC86661 similar to GP|9798599|dbj|BAB11737.1 serine/threonine protein
kinase {Arabidopsis thaliana}, partial (58%)
Length = 1840
Score = 104 bits (259), Expect = 6e-23
Identities = 83/281 (29%), Positives = 137/281 (48%), Gaps = 6/281 (2%)
Frame = +2
Query: 13 GKMVGCGSFGSVHLAMNKSTGGLFVVK-----KAHSEAGRDALENEVNILNTLNSSPSPY 67
GK++GCG+F V+ A N G VK K + ++ E++I++ L
Sbjct: 314 GKLLGCGAFAKVYHARNTENGQSVAVKVINKKKISATGLAGHVKREISIMSKLRHPNIVR 493
Query: 68 IVQCLGTDYDHQDNQLHVFMEYMSGGSL-ADVSHKFGGSLNEDVVRVYTRQIVHGLYHLH 126
+ + L T +++ ME+ GG L A +++K G +ED+ R +Q++ + + H
Sbjct: 494 LHEVLATK-----TKIYFVMEFAKGGELFAKIANK--GRFSEDLSRRLFQQLISAVGYCH 652
Query: 127 QHGIVHCDLKCKNVLLASSGNVKLADFGCAKRVKKNMNMNKSSSCIIANGGTPLWMAPEV 186
HG+ H DLK +N+LL GN+K++DFG + VK+ + ++ + GTP ++APE+
Sbjct: 653 SHGVFHRDLKPENLLLDDKGNLKVSDFGLS-AVKEQVRIDGMLHTLC---GTPAYVAPEI 820
Query: 187 LLMKNNSSINDESRVVDFAAADIWSLGCTVIEMATGRPPWVDDDLISISNPMAAMFKIAC 246
L R D A DIWS G + + G P+ D +L M KI
Sbjct: 821 L----------AKRGYDGAKVDIWSCGIILFVLVAGYLPFNDPNL------MVMYRKIYS 952
Query: 247 GDGIPQFPIHFSQEGFDFLRRCLVRDPKKRSTALELLNHPF 287
G+ + P FS + FL R + +P+ R T E+L P+
Sbjct: 953 GEF--KCPRWFSPDLKRFLSRLMDTNPETRITVDEILRDPW 1069
>TC77429 similar to GP|15215664|gb|AAK91377.1 AT5g25110/T11H3_120
{Arabidopsis thaliana}, partial (79%)
Length = 1869
Score = 103 bits (256), Expect = 1e-22
Identities = 80/306 (26%), Positives = 142/306 (46%), Gaps = 10/306 (3%)
Frame = +3
Query: 13 GKMVGCGSFGSVHLAMNKSTGGLFVVK-----KAHSEAGRDALENEVNILNTLNSSPSPY 67
G+++G G+F V+ A STG +K K E + ++ E++++ +
Sbjct: 279 GRVLGKGTFAKVYYAKEISTGEGVAIKVVDKDKVKKEGMMEQIKREISVMRLVKHPNIVN 458
Query: 68 IVQCLGTDYDHQDNQLHVFMEYMSGGSLADVSHKFGGSLNEDVVRVYTRQIVHGLYHLHQ 127
+ + + T ++ MEY GG L + K G E++ R Y +Q++ + + H
Sbjct: 459 LKEVMATK-----TKILFVMEYARGGELFEKVAK--GKFKEELARKYFQQLISAVDYCHS 617
Query: 128 HGIVHCDLKCKNVLLASSGNVKLADFGCAKRVKKNMNMNKSSSCIIANGGTPLWMAPEVL 187
G+ H DLK +N+LL + N+K++DFG + + + + GTP ++APEV+
Sbjct: 618 RGVSHRDLKPENLLLDENENLKVSDFGLSALPEH----LRQDGLLHTQCGTPAYVAPEVV 785
Query: 188 LMKNNSSINDESRVVDFAAADIWSLGCTVIEMATGRPPWVDDDLISISNPMAAMFKIACG 247
+ S AD WS G + + G P+ ++LIS+ N +FK
Sbjct: 786 RKRGYSGFK----------ADTWSCGVILYALLAGFLPFQHENLISMYN---KVFKEEY- 923
Query: 248 DGIPQFPIHFSQEGFDFLRRCLVRDPKKRSTALELLNHPFL-----VSTTLTHHKHYSAS 302
QFP FS E + + LV DP++R T ++N P+ +STT T + +
Sbjct: 924 ----QFPPWFSPESKRLISKILVADPERRITISSIMNVPWFQKGLCLSTTTTANDDFELE 1091
Query: 303 SPASVL 308
S +++
Sbjct: 1092SKVNLI 1109
>TC78104 homologue to GP|19568098|gb|AAL89456.1 osmotic stress-activated
protein kinase {Nicotiana tabacum}, partial (95%)
Length = 1434
Score = 102 bits (253), Expect = 3e-22
Identities = 91/309 (29%), Positives = 137/309 (43%), Gaps = 13/309 (4%)
Frame = +3
Query: 14 KMVGCGSFGSVHLAMNKSTGGLFVVKKAHSEAGRDALENEVNILNTLNSSPSPYIVQCLG 73
K +G G+FG L +K T L +K + E G EN + S P I++
Sbjct: 174 KDIGSGNFGVARLMRHKDTKQLVAMK--YIERGHKIDENVAREIINHRSLRHPNIIRF-- 341
Query: 74 TDYDHQDNQLHVFMEYMSGGSLADVSHKFGGSLNEDVVRVYTRQIVHGLYHLHQHGIVHC 133
+ L + MEY +GG L D G +ED R + +Q++ G+ + H I H
Sbjct: 342 KEVVLTPTHLGIVMEYAAGGELFDRICS-AGRFSEDEARYFFQQLISGVSYCHSMQICHR 518
Query: 134 DLKCKNVLLASSG--NVKLADFGCAKRVKKNMNMNKSSSCIIANGGTPLWMAPEVLLMKN 191
DLK +N LL S +K+ DFG +K ++ ++ S + GTP ++APEVL
Sbjct: 519 DLKLENTLLDGSPAPRLKICDFGYSK---SSLLHSRPKSTV----GTPAYIAPEVL---- 665
Query: 192 NSSINDESRVVDFAAADIWSLGCTVIEMATGRPPWVDDD-----LISISNPMAAMFKIAC 246
R D AD+WS G T+ M G P+ D D +I+ MA +KI
Sbjct: 666 ------SRREYDGKLADVWSCGVTLYVMLVGAYPFEDQDDPKNFRKTINRIMAVQYKI-- 821
Query: 247 GDGIPQFPIHFSQEGFDFLRRCLVRDPKKRSTALELLNHPFLVS------TTLTHHKHYS 300
P + +H SQE L R V P +R T E+ +HP+ + T + +Y
Sbjct: 822 ----PDY-VHISQECRHLLSRIFVASPARRITIKEIKSHPWFLKNLPRELTEMAQAVYYR 986
Query: 301 ASSPASVLE 309
+P L+
Sbjct: 987 KENPTYSLQ 1013
>TC87242 homologue to PIR|S56716|S56716 protein kinase SPK-3 (EC 2.7.1.-) -
soybean, complete
Length = 1659
Score = 101 bits (252), Expect = 4e-22
Identities = 86/283 (30%), Positives = 131/283 (45%), Gaps = 7/283 (2%)
Frame = +1
Query: 14 KMVGCGSFGSVHLAMNKSTGGLFVVKKAHSEAGRDALENEVNILNTLNSSPSPYIVQCLG 73
K +G G+FG L NK T L +K + E G+ EN + S P I++
Sbjct: 490 KDLGAGNFGVARLLRNKETKELVAMK--YIERGQKIDENVAREIINHRSLRHPNIIRF-- 657
Query: 74 TDYDHQDNQLHVFMEYMSGGSLADVSHKFGGSLNEDVVRVYTRQIVHGLYHLHQHGIVHC 133
+ L + MEY +GG L + G +ED R + +Q++ G+++ H I H
Sbjct: 658 KEVVLTPTHLAIVMEYAAGGELFERICN-AGRFSEDEARYFFQQLISGVHYCHAMQICHR 834
Query: 134 DLKCKNVLLASSG--NVKLADFGCAKRVKKNMNMNKSSSCIIANGGTPLWMAPEVLLMKN 191
DLK +N LL S +K+ DFG +K ++ ++ S + GTP ++APEVL
Sbjct: 835 DLKLENTLLDGSPAPRLKICDFGYSK---SSLLHSRPKSTV----GTPAYIAPEVL---- 981
Query: 192 NSSINDESRVVDFAAADIWSLGCTVIEMATGRPPWVDDD-----LISISNPMAAMFKIAC 246
R D AD+WS G T+ M G P+ D D +I MA +K
Sbjct: 982 ------SRREYDGKLADVWSCGVTLYVMLVGAYPFEDQDDPRNFRKTIQRIMAVQYK--- 1134
Query: 247 GDGIPQFPIHFSQEGFDFLRRCLVRDPKKRSTALELLNHPFLV 289
IP + +H SQ+ L R V +P +R + E+ NHP+ +
Sbjct: 1135---IPDY-VHISQDCKHLLSRIFVANPLRRISLKEIKNHPWFL 1251
>TC78699 weakly similar to PIR|G96761|G96761 probable MAP kinase T9L24.32
[imported] - Arabidopsis thaliana, partial (71%)
Length = 1119
Score = 101 bits (252), Expect = 4e-22
Identities = 86/289 (29%), Positives = 143/289 (48%), Gaps = 5/289 (1%)
Frame = +3
Query: 5 IKGSEWMKGKMVGCGSFGSVHLAMNKSTGGLFVVKKAHSEAG----RDALENEVNILNTL 60
I ++ K ++G G+ G+V+ +K T ++ +K H ++ R AL EVNIL
Sbjct: 186 ISAGDFEKLSVLGHGNGGTVYKVRHKLTSIIYALKINHYDSDPTTRRRAL-TEVNILRRA 362
Query: 61 NSSPSPYIVQCLGTDYDHQDNQLHVFMEYMSGGSLADVSHKFGGSLNEDVVRVYTRQIVH 120
+ +V+ G+ ++ + + MEYM GSL + + K G+ +E + R I++
Sbjct: 363 TDCTN--VVKYHGS-FEKPTGDVCILMEYMDSGSL-ETALKTTGTFSESKLSTVARDILN 530
Query: 121 GLYHLHQHGIVHCDLKCKNVLLASSGNVKLADFGCAKRVKKNMNMNKSSSCIIANGGTPL 180
GL +LH I H D+K N+L+ VK+ADFG +K + + + S GT
Sbjct: 531 GLTYLHARNIAHRDIKPSNILVNIKNEVKIADFGVSKFMGRTLEACNSYV------GTCA 692
Query: 181 WMAPEVLLMKNNSSINDESRVVDFAAADIWSLGCTVIEMATGRPPWVDDDLISISNP-MA 239
+M+PE + + + + + +ADIWSLG T+ E+ G P+ L S P A
Sbjct: 693 YMSPE----RFDPEVYGGN--YNGFSADIWSLGLTLFELYVGYFPF----LQSGQRPDWA 842
Query: 240 AMFKIACGDGIPQFPIHFSQEGFDFLRRCLVRDPKKRSTALELLNHPFL 288
++ C P P S E +F+ CL ++ +R +A +LL HPFL
Sbjct: 843 SLMCAICFSDPPSLPETASSEFRNFVECCLKKESGERWSAAQLLTHPFL 989
>TC77188 similar to PIR|T04862|T04862 probable serine/threonine-specific
protein kinase (EC 2.7.1.-) F28A21.110 - Arabidopsis
thaliana, partial (79%)
Length = 1881
Score = 100 bits (249), Expect = 9e-22
Identities = 94/332 (28%), Positives = 146/332 (43%), Gaps = 11/332 (3%)
Frame = +1
Query: 13 GKMVGCGSFGSVHLAMNKSTGGLFVVK-----KAHSEAGRDALENEVNILNTLNSSPSPY 67
GK++G G+F VHLA N TG +K K ++ E++IL +
Sbjct: 199 GKLLGHGTFAKVHLAKNLKTGESVAIKIISKDKILKSGLVSHIKREISILRRVRHPNIVQ 378
Query: 68 IVQCLGTDYDHQDNQLHVFMEYMSGGSLADVSHKFGGSLNEDVVRVYTRQIVHGLYHLHQ 127
+ + + T +++ MEY+ GG L + K G L E+V R Y +Q++ + H
Sbjct: 379 LFEVMATK-----TKIYFVMEYVRGGELFNKVAK--GRLKEEVARKYFQQLICAVGFCHA 537
Query: 128 HGIVHCDLKCKNVLLASSGNVKLADFGCAKRVKKNMNMNKSSSCIIANGGTPLWMAPEVL 187
G+ H DLK +N+LL GN+K++DFG + + K GTP ++APEVL
Sbjct: 538 RGVFHRDLKPENLLLDEKGNLKVSDFG----LSAVSDEIKQDGLFHTFCGTPAYVAPEVL 705
Query: 188 LMKNNSSINDESRVVDFAAADIWSLGCTVIEMATGRPPWVDDDLISISNPMAAMFKIACG 247
K D A DIWS G + + G P+ D +N MA KI G
Sbjct: 706 SRKG----------YDGAKVDIWSCGVVLFVLMAGYLPFHDP-----NNVMAMYKKIYKG 840
Query: 248 DGIPQFPIHFSQEGFDFLRRCLVRDPKKRSTALELLNHPFLVSTTLTHHKHYSASSPASV 307
+ + P FS E L R L P+ R + E++ + + H K Y
Sbjct: 841 EF--RCPRWFSPELVSLLTRLLDIKPQTRISIPEIMENRWF-KIGFKHIKFYVEDDVVCD 1011
Query: 308 LEVHQFEDTYDDDDDDDDS------DDNDELI 333
L D+ D D +D+++ D +DE++
Sbjct: 1012L------DSLDLDGEDNNNKVVKVDDHHDEVL 1089
>TC88197 homologue to GP|6739629|gb|AAF27340.1| abscisic acid-activated
protein kinase {Vicia faba}, partial (96%)
Length = 1158
Score = 99.4 bits (246), Expect = 2e-21
Identities = 101/354 (28%), Positives = 151/354 (42%), Gaps = 31/354 (8%)
Frame = +1
Query: 16 VGCGSFGSVHLAMNKSTGGLFVVKKAHSEAGRDALENEVNILNTLNSSPSPYIVQCLGTD 75
+G G+FG L +K T L VK + E G EN + S P IV+ +
Sbjct: 145 IGSGNFGVARLMRDKHTKELVAVK--YIERGDKIDENVKREIINHRSLRHPNIVRF--KE 312
Query: 76 YDHQDNQLHVFMEYMSGGSLADVSHKFGGSLNEDVVRVYTRQIVHGLYHLHQHGIVHCDL 135
L + MEY SGG + D + G ED R + +Q++ G+ + H + H DL
Sbjct: 313 VILTPTHLAIVMEYASGGEMFDRISR-AGRFTEDEARFFFQQLISGVSYCHSMQVCHRDL 489
Query: 136 KCKNVLLASSG--NVKLADFGCAKRVKKNMNMNKSSSCIIANGGTPLWMAPEVLLMKNNS 193
K +N LL ++K+ DFG +K ++ ++ S + GTP ++APEVLL +
Sbjct: 490 KLENTLLDGDPALHLKICDFGYSK---SSVLHSQPKSTV----GTPAYIAPEVLLKQE-- 642
Query: 194 SINDESRVVDFAAADIWSLGCTVIEMATGRPPWVDDDLISISNP----MAAMFKIACGDG 249
D AD+WS G T+ M G P+ D D NP ++
Sbjct: 643 --------YDGKVADVWSCGVTLYVMLVGSYPFEDPD-----NPKDFRKTIQRVLSVQYS 783
Query: 250 IPQFPIHFSQEGFDFLRRCLVRDPKKRSTALELLNHPF--------LVSTTLTHHKH--- 298
+P F + S E + + R V DP KR T E+L + + LV+ LT ++
Sbjct: 784 VPDF-VQISPECREVISRIFVFDPAKRITIPEILKNEWFLKNLPADLVNEKLTDNQFEEP 960
Query: 299 --------------YSASSPASVLEVHQFEDTYDDDDDDDDSDDNDELISPAGG 338
A+ PA+ + QF D DDD DD D + EL + G
Sbjct: 961 DQPMQSMDTIMQIISEATVPAAGSYLDQFMPDNDTDDDMDDMDSDYELDVDSSG 1122
>BG454756 homologue to GP|12718822|db NQK1 MAPKK {Nicotiana tabacum}, partial
(60%)
Length = 648
Score = 97.8 bits (242), Expect = 6e-21
Identities = 71/234 (30%), Positives = 115/234 (48%), Gaps = 2/234 (0%)
Frame = +2
Query: 54 VNILNTLNSSPSPYIVQCLGTDYDHQDNQLHVFMEYMSGGSLADVSHKFGGSLNEDVVRV 113
V L +S P++V C + Y++ + + +EYM GSL DV + ++ E + V
Sbjct: 5 VQELKINQASQCPHVVVCYHSFYNN--GVISLVLEYMDRGSLVDVIRQV-NTILEPYLAV 175
Query: 114 YTRQIVHGLYHLH-QHGIVHCDLKCKNVLLASSGNVKLADFGCAKRVKKNMNMNKSSSCI 172
+Q++ GL +LH + ++H D+K N+L+ G VK+ DFG + + M +
Sbjct: 176 VCKQVLQGLVYLHNERHVIHRDIKPSNLLVNHKGEVKITDFGVSAMLASTMGQRDTFV-- 349
Query: 173 IANGGTPLWMAPEVLLMKNNSSINDESRVVDFAAADIWSLGCTVIEMATGRPPWVDDDLI 232
GT +M+PE + + S D S DIWSLG V+E A GR P++ +
Sbjct: 350 ----GTYNYMSPE----RISGSTYDYS-------CDIWSLGMVVLECAIGRFPYIQSEDQ 484
Query: 233 SISNPMAAMFKIACGDGIPQF-PIHFSQEGFDFLRRCLVRDPKKRSTALELLNH 285
+ + P P FS E F+ C+ +DP++RST+LELL+H
Sbjct: 485 QAWPSFYELLQAIVESPPPSAPPDQFSPEFCSFVSSCIKKDPRERSTSLELLDH 646
Database: MTGI
Posted date: Oct 22, 2004 3:39 PM
Number of letters in database: 27,044,181
Number of sequences in database: 36,976
Lambda K H
0.319 0.135 0.418
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 13,364,222
Number of Sequences: 36976
Number of extensions: 203628
Number of successful extensions: 3367
Number of sequences better than 10.0: 542
Number of HSP's better than 10.0 without gapping: 1979
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 2534
length of query: 373
length of database: 9,014,727
effective HSP length: 98
effective length of query: 275
effective length of database: 5,391,079
effective search space: 1482546725
effective search space used: 1482546725
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.8 bits)
S2: 59 (27.3 bits)
Medicago: description of AC140549.19


