

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC140544.4 - phase: 0 /pseudo
(305 letters)
Database: MTGI
36,976 sequences; 27,044,181 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
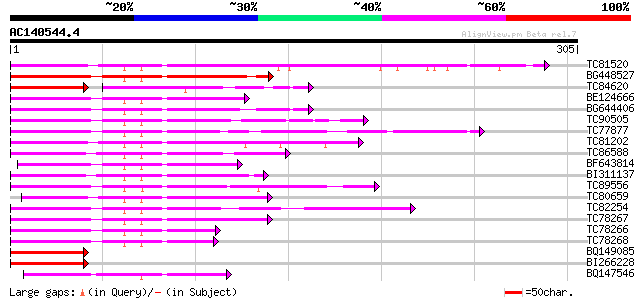
TC81520 similar to PIR|D86470|D86470 hypothetical protein AAD460... 139 1e-33
BG448527 similar to PIR|T51509|T51 probable transcription factor... 122 2e-28
TC84620 similar to PIR|JQ0956|JQ0956 myb-related protein 306 - g... 75 2e-24
BE124666 similar to PIR|JQ0957|JQ0 myb-related protein 330 - gar... 107 6e-24
BG644406 homologue to PIR|T07398|T07 myb-related transcription f... 105 3e-23
TC90505 similar to GP|3941528|gb|AAC83640.1| putative transcript... 104 5e-23
TC77877 similar to GP|5139806|dbj|BAA81732.1 GmMYB29A2 {Glycine ... 103 6e-23
TC81202 homologue to GP|17380966|gb|AAL36295.1 putative transcri... 102 2e-22
TC86588 similar to GP|5139814|dbj|BAA81736.1 GmMYB29B2 {Glycine ... 100 9e-22
BF643814 similar to GP|19386839|db putative myb-related protein ... 97 6e-21
BI311137 similar to GP|17380966|gb putative transcription factor... 96 1e-20
TC89556 similar to GP|18419456|gb|AAL69334.1 blind {Lycopersicon... 93 1e-19
TC80659 similar to PIR|T48607|T48607 probable transcription fact... 89 3e-18
TC82254 similar to GP|22266673|dbj|BAC07543. myb-related transcr... 85 4e-17
TC78267 weakly similar to PIR|B85064|B85064 MYB-like protein [im... 84 9e-17
TC78266 similar to GP|22266673|dbj|BAC07543. myb-related transcr... 80 1e-15
TC78268 weakly similar to PIR|B85064|B85064 MYB-like protein [im... 78 5e-15
BQ149085 similar to GP|9802570|gb|A F22O13.30 {Arabidopsis thali... 76 1e-14
BI266228 homologue to GP|10178078|dbj MYB96 transcription factor... 75 3e-14
BQ147546 similar to PIR|H96706|H96 probable transcription factor... 60 8e-10
>TC81520 similar to PIR|D86470|D86470 hypothetical protein AAD46010.1
[imported] - Arabidopsis thaliana, partial (39%)
Length = 1259
Score = 139 bits (351), Expect = 1e-33
Identities = 112/344 (32%), Positives = 171/344 (49%), Gaps = 54/344 (15%)
Frame = +2
Query: 1 MGRSPCCEKVGLKKGPWTSEEDQKLLSYIEEHGHGSWRSLPTKAVQDLRGVARAAD*DGL 60
MGRSPCC++ GLKKGPWT EEDQKL+ +I++HGHGSWR+LP L G+ R L
Sbjct: 56 MGRSPCCDESGLKKGPWTPEEDQKLVEHIQQHGHGSWRALPK-----LAGLNRCGKSCRL 220
Query: 61 ----II*GRILRG------ESLVCKKNKPLFNFMHF*ATATRLPKRTDNEIKNYWNTHLK 110
+ I RG E + + L N + A AT LP RTDNEIKN+WNTHLK
Sbjct: 221 RWTNYLRPDIKRGKFSQEEEQTILHLHSILGN--KWSAIATHLPGRTDNEIKNFWNTHLK 394
Query: 111 KRLTKMGIDPVTHKPKNETLLPENHSKNVANLS------HKAQWE----SARLEAEARLV 160
K+L +MG DP+TH+P+ + + + +AN++ +++ W+ +A +A V
Sbjct: 395 KKLIQMGFDPMTHQPRTDLVSTIPYLLALANMTEIIDHQNQSSWDDQQHAAMSSLQAEAV 574
Query: 161 RESKLRSQLQHQFGSNTSVFPSQSSSSSSNQVLNIKEE----------GEKEWKGYE--- 207
+KL+ SN+S+ + +S + + N+++E KE +
Sbjct: 575 HLAKLQCLQYLLQSSNSSININNNSYDQNAMITNMEQEQPLSLLNTISNVKENTIMDCSQ 754
Query: 208 -DSTHLLEFKDLMENSS----------MAFS--STLQHEM----TMINVAEEGFTNLLLD 250
DSTH F + + S ++FS S L +E T+ N A T+ ++
Sbjct: 755 LDSTHAASFSQPLHHQSVLPHFLDPQQVSFSSQSCLNNEQSQGGTITNFATVDETS-WIN 931
Query: 251 NSNSGDLSLSPE----SGGECNTCDGSGSGGGSDFNEDNKNYWN 290
+S +S+ P S G+ ++C S GGG +YW+
Sbjct: 932 VPSSAPISVPPNAIGISAGDASSCTSSYGGGG---GPSPVSYWS 1054
>BG448527 similar to PIR|T51509|T51 probable transcription factor (MYB9) -
Arabidopsis thaliana, partial (40%)
Length = 633
Score = 122 bits (305), Expect = 2e-28
Identities = 68/152 (44%), Positives = 93/152 (60%), Gaps = 10/152 (6%)
Frame = +1
Query: 1 MGRSPCCEKVGLKKGPWTSEEDQKLLSYIEEHGHGSWRSLPTKAVQDLRGVARAAD*DGL 60
MGRSPCC+++G+KKGPWT EED+KL+ YI +HGHG+W +L +A G+ R L
Sbjct: 40 MGRSPCCDEIGVKKGPWTQEEDEKLIDYINKHGHGNWGTLSKRA-----GLNRCGKSCRL 204
Query: 61 ----II*GRILRG------ESLVCKKNKPLFNFMHF*ATATRLPKRTDNEIKNYWNTHLK 110
+ I RG E ++ + L N + A LP RTDNEIKNYWNT+++
Sbjct: 205 RWTNYLRPDIKRGKFTDEEERVIINLHSVLGN--KWSKIAAHLPGRTDNEIKNYWNTNIR 378
Query: 111 KRLTKMGIDPVTHKPKNETLLPENHSKNVANL 142
K+L KMGIDP THKP+ + NH +++NL
Sbjct: 379 KKLLKMGIDPETHKPRTD----YNHLMSLSNL 462
>TC84620 similar to PIR|JQ0956|JQ0956 myb-related protein 306 - garden
snapdragon, partial (44%)
Length = 593
Score = 75.1 bits (183), Expect(2) = 2e-24
Identities = 30/42 (71%), Positives = 37/42 (87%)
Frame = +2
Query: 1 MGRSPCCEKVGLKKGPWTSEEDQKLLSYIEEHGHGSWRSLPT 42
MGR PCCEK+G+KKGPWT EED L+SYI++HG G+WRS+PT
Sbjct: 119 MGRPPCCEKLGIKKGPWTPEEDIILVSYIQQHGPGNWRSVPT 244
Score = 54.7 bits (130), Expect(2) = 2e-24
Identities = 40/118 (33%), Positives = 60/118 (49%), Gaps = 5/118 (4%)
Frame = +3
Query: 51 VARAAD*DGLII*GRILRGESLVCKKNKPLFNFMHF*ATATRL-----PKRTDNEIKNYW 105
V RAAD DG I ++ + + K K LF F AT L P+RTDN+IKNYW
Sbjct: 264 VVRAADLDGQTIFDQVSKEVTSQIMKRK*LFTSKLFWATDGLL*LHILPQRTDNDIKNYW 443
Query: 106 NTHLKKRLTKMGIDPVTHKPKNETLLPENHSKNVANLSHKAQWESARLEAEARLVRES 163
NTHLK+++ K +E + + S+ L +K QWE L+ + + +++
Sbjct: 444 NTHLKRKMNK------DQSSTDEGVDQXSRSQ----LPNKGQWERX-LQTDIHMAKQA 584
>BE124666 similar to PIR|JQ0957|JQ0 myb-related protein 330 - garden
snapdragon, partial (48%)
Length = 536
Score = 107 bits (267), Expect = 6e-24
Identities = 61/139 (43%), Positives = 80/139 (56%), Gaps = 10/139 (7%)
Frame = +3
Query: 1 MGRSPCCEKVGLKKGPWTSEEDQKLLSYIEEHGHGSWRSLPTKAVQDLRGVARAAD*DGL 60
MGRSPCCEK KG WT EED++L++YI+ HG G WRSLP A G+ R L
Sbjct: 27 MGRSPCCEKEHTNKGAWTKEEDERLINYIKLHGEGCWRSLPKAA-----GLLRCGKSCRL 191
Query: 61 ----II*GRILRG------ESLVCKKNKPLFNFMHF*ATATRLPKRTDNEIKNYWNTHLK 110
+ + RG + L+ + L N + A RLP RTDNEIKNYWNTH+K
Sbjct: 192 RWINYLRPDLKRGNFTEQEDDLIINLHSLLGN--KWSLIAARLPGRTDNEIKNYWNTHIK 365
Query: 111 KRLTKMGIDPVTHKPKNET 129
++L G+DP TH+ N++
Sbjct: 366 RKLYSRGVDPQTHRSLNDS 422
>BG644406 homologue to PIR|T07398|T07 myb-related transcription factor THM6 -
tomato, partial (42%)
Length = 762
Score = 105 bits (261), Expect = 3e-23
Identities = 64/173 (36%), Positives = 95/173 (53%), Gaps = 10/173 (5%)
Frame = +1
Query: 1 MGRSPCCEKVGLKKGPWTSEEDQKLLSYIEEHGHGSWRSLPTKAVQDLRGVARAAD*DGL 60
MGR PCC+K G+KKGPWT EED L+S+++EHG G+WR++PT G+ R + L
Sbjct: 145 MGRPPCCDKTGVKKGPWTPEEDIMLVSFVQEHGPGNWRTVPTHT-----GLRRCSKSCRL 309
Query: 61 ----II*GRILRG------ESLVCKKNKPLFNFMHF*ATATRLPKRTDNEIKNYWNTHLK 110
+ I RG E ++ + L N + A A+ LP+RTDN+IKNYWNTHLK
Sbjct: 310 RWTNYLRPGIKRGSFTDQEEKMIIQLQALLGN--KWAAIASYLPERTDNDIKNYWNTHLK 483
Query: 111 KRLTKMGIDPVTHKPKNETLLPENHSKNVANLSHKAQWESARLEAEARLVRES 163
K+L K+ + + H + N + + QWE L+A+ +++
Sbjct: 484 KKLKKLETSGLYSN--------DGHCLSTYNSTSRGQWERT-LQADINTAKQA 615
>TC90505 similar to GP|3941528|gb|AAC83640.1| putative transcription factor
{Arabidopsis thaliana}, partial (49%)
Length = 984
Score = 104 bits (259), Expect = 5e-23
Identities = 74/203 (36%), Positives = 109/203 (53%), Gaps = 10/203 (4%)
Frame = +1
Query: 1 MGRSPCCEKVGLKKGPWTSEEDQKLLSYIEEHGHGSWRSLPTKAVQDLRGVARAAD*DGL 60
MGR PCC+K G+KKGPWT EED L++YI+EHG G+WR++PTK G++R + L
Sbjct: 157 MGRPPCCDKEGVKKGPWTPEEDIILVTYIQEHGPGNWRAVPTKT-----GLSRCSKSCRL 321
Query: 61 ----II*GRILRG------ESLVCKKNKPLFNFMHF*ATATRLPKRTDNEIKNYWNTHLK 110
+ I RG E ++ L N + A A+ LP+RTDN+IKNYWNTHLK
Sbjct: 322 RWTNYLRPGIKRGNFTEQEEKMIIHLQDLLGN--RWAAIASYLPQRTDNDIKNYWNTHLK 495
Query: 111 KRLTKMGIDPVTHKPKNETLLPENHSKNVANLSHKAQWESARLEAEARLVRESKLRSQLQ 170
K+L K E + E + + + QWE RL+ + ++ +++ L L
Sbjct: 496 KKLKKSQTG-----NNAECINFEEEKFSASQRIPRGQWE-RRLQTDIQMAKKA-LNEALS 654
Query: 171 HQFGSNTSVFPSQSSSSSSNQVL 193
Q + P+ S+S+ +N L
Sbjct: 655 PQ-----KLSPNFSTSNPTNSNL 708
>TC77877 similar to GP|5139806|dbj|BAA81732.1 GmMYB29A2 {Glycine max},
partial (78%)
Length = 1156
Score = 103 bits (258), Expect = 6e-23
Identities = 87/266 (32%), Positives = 125/266 (46%), Gaps = 11/266 (4%)
Frame = +3
Query: 1 MGRSPCCEKVGLKKGPWTSEEDQKLLSYIEEHGHGSWRSLPTKAVQDLRGVARAAD*DGL 60
M R+PCCEK GLKKGPWT EED+ L SYI +HGH +WR+LP A G+ R L
Sbjct: 66 MVRAPCCEKKGLKKGPWTLEEDEILTSYINKHGHSNWRALPKHA-----GLLRCGKSCRL 230
Query: 61 ----II*GRILRG------ESLVCKKNKPLFNFMHF*ATATRLPKRTDNEIKNYWNTHLK 110
+ I RG E + K ++ L N + A A +LP RTDNEIKN W+THLK
Sbjct: 231 RWINYLKPDIKRGNFTNEEEETIIKMHESLGN--RWSAIAAKLPGRTDNEIKNVWHTHLK 404
Query: 111 KRLTKMGIDPVTHKPKNETLLPENHSKNVANLSHK-AQWESARLEAEARLVRESKLRSQL 169
KRL ++ ++P + T K V+ K + S+ L + S SQ
Sbjct: 405 KRL----LNTNNNQPNSNT------KKRVSKQKIKRSDSNSSTLTTASNCTFSSDFSSQE 554
Query: 170 QHQFGSNTSVFPSQSSSSSSNQVLNIKEEGEKEWKGYEDSTHLLEFKDLMENSSMAFSST 229
++ +S + ++ + E E W ++ E M ++SM S+
Sbjct: 555 KN----------LDNSIICEDSLVTMPEIDESFW---SETVIDDEISSTMPSNSMTVSND 695
Query: 230 LQHEMTMINVAEEGFTNLLLDNSNSG 255
L + + N + E F N DN + G
Sbjct: 696 LPDQQCIFNNSVENFQN-PFDNDDDG 770
>TC81202 homologue to GP|17380966|gb|AAL36295.1 putative transcription
factor {Arabidopsis thaliana}, partial (40%)
Length = 1506
Score = 102 bits (253), Expect = 2e-22
Identities = 79/221 (35%), Positives = 106/221 (47%), Gaps = 31/221 (14%)
Frame = +2
Query: 1 MGRSPCCEKVGLKKGPWTSEEDQKLLSYIEEHGHGSWRSLPTKAVQDLRGVARAAD*DGL 60
+GR CC K L+KG W+ EED+KLL+YI +HGHG W S+P L G+ R L
Sbjct: 59 LGRHSCCYKQKLRKGLWSPEEDEKLLNYITKHGHGCWSSVPK-----LAGLQRCGKSCRL 223
Query: 61 ----II*GRILRG------ESLVCKKNKPLFNFMHF*ATATRLPKRTDNEIKNYWNTHLK 110
+ + RG E+L+ + + L N + A +LP RTDNEIKN WN+ LK
Sbjct: 224 RWINYLRPDLKRGAFSQQEENLIIELHAVLGN--RWSQIAAQLPGRTDNEIKNLWNSCLK 397
Query: 111 KRLTKMGIDPVTHKP------KNETLLPENHSKNVANLSH-----------KAQWESARL 153
KRL + GIDP THKP NE L N + ++S K E L
Sbjct: 398 KRLRQSGIDPNTHKPLSEVENDNEKSLTSNKTNQKGSVSSNEVMSLMIEPTKPSIEGYPL 577
Query: 154 EAEARLVRESKLRSQ----LQHQFGSNTSVFPSQSSSSSSN 190
E + S L +FGS++S F Q+ + SN
Sbjct: 578 EVSTTSKINNSSSSSHELFLDTRFGSSSSYFSFQNLNYGSN 700
>TC86588 similar to GP|5139814|dbj|BAA81736.1 GmMYB29B2 {Glycine max},
partial (85%)
Length = 1389
Score = 100 bits (248), Expect = 9e-22
Identities = 65/161 (40%), Positives = 90/161 (55%), Gaps = 10/161 (6%)
Frame = +2
Query: 1 MGRSPCCEKVGLKKGPWTSEEDQKLLSYIEEHGHGSWRSLPTKAVQDLRGVARAAD*DGL 60
M R+PCCEK+GLK+GPW+ EED+ L SYI++HGHG+WR+LP L G+ R L
Sbjct: 107 MVRAPCCEKMGLKRGPWSLEEDEILTSYIQKHGHGNWRALP-----KLAGLLRCGKSCRL 271
Query: 61 ----II*GRILRG------ESLVCKKNKPLFNFMHF*ATATRLPKRTDNEIKNYWNTHLK 110
+ I RG E + K ++ L N + A A +LP RTDNEIKN W+THLK
Sbjct: 272 RWINYLRPDIKRGNFTNEEEENIIKLHEMLGN--RWSAIAAKLPGRTDNEIKNVWHTHLK 445
Query: 111 KRLTKMGIDPVTHKPKNETLLPENHSKNVANLSHKAQWESA 151
K+L + + K + + ++ +N S Q ESA
Sbjct: 446 KKLNQ-----TNSEAKKKAISKPKIKRSDSNSSTITQSESA 553
>BF643814 similar to GP|19386839|db putative myb-related protein {Oryza
sativa (japonica cultivar-group)}, partial (49%)
Length = 545
Score = 97.4 bits (241), Expect = 6e-21
Identities = 55/131 (41%), Positives = 75/131 (56%), Gaps = 10/131 (7%)
Frame = +1
Query: 5 PCCEKVGLKKGPWTSEEDQKLLSYIEEHGHGSWRSLPTKAVQDLRGVARAAD*DGL---- 60
PCCEK KG W+ EED++L++YI++ G G WRSLP A G+AR L
Sbjct: 1 PCCEKQHTNKGAWSKEEDERLINYIKQQGEGCWRSLPKAA-----GLARCGKSCRLRWIN 165
Query: 61 II*GRILRG------ESLVCKKNKPLFNFMHF*ATATRLPKRTDNEIKNYWNTHLKKRLT 114
+ + RG + L+ + + N + A +LP RTDNEIKNYWNTH+K++L
Sbjct: 166 YLRPDLKRGNFTHEEDELIISLHAMVGN--KWSQIAQKLPGRTDNEIKNYWNTHIKRKLY 339
Query: 115 KMGIDPVTHKP 125
GIDP TH+P
Sbjct: 340 SRGIDPTTHQP 372
>BI311137 similar to GP|17380966|gb putative transcription factor
{Arabidopsis thaliana}, partial (35%)
Length = 761
Score = 96.3 bits (238), Expect = 1e-20
Identities = 61/149 (40%), Positives = 84/149 (55%), Gaps = 10/149 (6%)
Frame = +1
Query: 1 MGRSPCCEKVGLKKGPWTSEEDQKLLSYIEEHGHGSWRSLPTKAVQDLRGVARAAD*DGL 60
MGR CC K L+KG W+ EED+KLL+YI +HGHG W S+P L G+ R L
Sbjct: 148 MGRHSCCYKQKLRKGLWSPEEDEKLLNYITKHGHGCWSSVP-----KLAGLQRCGKSCRL 312
Query: 61 ----II*GRILRG------ESLVCKKNKPLFNFMHF*ATATRLPKRTDNEIKNYWNTHLK 110
+ + RG E+ + + + L N + A +LP RTDNEIKN WN+ LK
Sbjct: 313 RWINYLRPDLKRGAFSEQEENSIIELHALLGN--RWSQIAAQLPGRTDNEIKNLWNSSLK 486
Query: 111 KRLTKMGIDPVTHKPKNETLLPENHSKNV 139
K+L + GIDP TH+P +E +N + N+
Sbjct: 487 KKLKQKGIDPNTHQPFSEN--NDNDTPNI 567
>TC89556 similar to GP|18419456|gb|AAL69334.1 blind {Lycopersicon
esculentum}, partial (37%)
Length = 990
Score = 92.8 bits (229), Expect = 1e-19
Identities = 70/212 (33%), Positives = 106/212 (49%), Gaps = 13/212 (6%)
Frame = +3
Query: 1 MGRSPCCEKVGLKKGPWTSEEDQKLLSYIEEHG-HGSWRSLPTKAVQDLRGVARAAD*DG 59
MGRSPCC+K +KKGPW+ EED KL YIE+HG G+W +LP K G+ R
Sbjct: 18 MGRSPCCDKANVKKGPWSPEEDAKLKEYIEKHGTGGNWITLPKKV-----GLTRCGKSCR 182
Query: 60 L----II*GRILRGE------SLVCKKNKPLFNFMHF*ATATRLPKRTDNEIKNYWNTHL 109
L + I GE ++C + + + A++L RTDN++KNYWNT L
Sbjct: 183 LRWLNYLRPNIKHGEFSDSEDKIICTLFASIGS--RWSIIASKLEGRTDNDVKNYWNTKL 356
Query: 110 KKRLTKMGIDPVTHKPKNETLLP--ENHSKNVANLSHKAQWESARLEAEARLVRESKLRS 167
KK++ M V KP+ TLL +N +K+ +LS S + + L S +
Sbjct: 357 KKKIMSMN-HSVEMKPQQVTLLSILQNSTKSSPSLSFTNSSFSYSSTSSSLLSGNSSTSA 533
Query: 168 QLQHQFGSNTSVFPSQSSSSSSNQVLNIKEEG 199
+ + S S ++S NQ+ ++ ++G
Sbjct: 534 AQES--------YISPSKNNSKNQINHVIDQG 605
>TC80659 similar to PIR|T48607|T48607 probable transcription factor MYB40 -
Arabidopsis thaliana, partial (46%)
Length = 1040
Score = 88.6 bits (218), Expect = 3e-18
Identities = 55/145 (37%), Positives = 78/145 (52%), Gaps = 10/145 (6%)
Frame = +2
Query: 7 CEKVGLKKGPWTSEEDQKLLSYIEEHGHGSWRSLPTKAVQDLRGVARAAD*DGL----II 62
C+KVGLK+GPWT EED KL+++I +G WR +P L G+ R L +
Sbjct: 14 CDKVGLKRGPWTIEEDHKLVNFILNNGIHCWRMVPK-----LAGLLRCGKSCRLRWINYL 178
Query: 63 *GRILRG------ESLVCKKNKPLFNFMHF*ATATRLPKRTDNEIKNYWNTHLKKRLTKM 116
+ RG E + + + L N + A+ P RTDNEIKN+WNT +KK+L +
Sbjct: 179 RPDLKRGGFTEIEEDQIIQLHARLGN--RWSKIASHFPGRTDNEIKNHWNTKIKKKLKFL 352
Query: 117 GIDPVTHKPKNETLLPENHSKNVAN 141
G+DPV HKP + + KN+ N
Sbjct: 353 GLDPVNHKPIEQKQQTLDDDKNIIN 427
>TC82254 similar to GP|22266673|dbj|BAC07543. myb-related transcription
factor VlMYBB1-1 {Vitis labrusca x Vitis vinifera},
partial (46%)
Length = 754
Score = 84.7 bits (208), Expect = 4e-17
Identities = 67/228 (29%), Positives = 106/228 (46%), Gaps = 10/228 (4%)
Frame = +2
Query: 1 MGRSPCCEKVGLKKGPWTSEEDQKLLSYIEEHGHGSWRSLPTKAVQDLRGVARAAD*DGL 60
M R+P C+K GL+KG WT EED+KL++Y+ +G +WR LP A G+ R L
Sbjct: 98 MVRTPYCDKNGLRKGTWTPEEDKKLIAYVTRYGCWNWRQLPKFA-----GLERCGKSCRL 262
Query: 61 ----II*GRILRG------ESLVCKKNKPLFNFMHF*ATATRLPKRTDNEIKNYWNTHLK 110
+ I RG E + K ++ L N + + LP RTDNEIKNYW+T +K
Sbjct: 263 RWLNYLRPDIKRGSFSHEEEETIIKLHEKLGN--RWTMISANLPGRTDNEIKNYWHTTIK 436
Query: 111 KRLTKMGIDPVTHKPKNETLLPENHSKNVANLSHKAQWESARLEAEARLVRESKLRSQLQ 170
K L K K+ T +K+ + +H + + KL L+
Sbjct: 437 KTLV---------KNKSNTKRGTKKAKDSNSKNHPT------------MEKPKKLGEMLE 553
Query: 171 HQFGSNTSVFPSQSSSSSSNQVLNIKEEGEKEWKGYEDSTHLLEFKDL 218
+ N + PS S+S++ +++ + YE+S+ ++ DL
Sbjct: 554 NNSDFNITSPPSSQPSTSASSCISLMDTAATTTISYENSSLFDDYDDL 697
>TC78267 weakly similar to PIR|B85064|B85064 MYB-like protein [imported] -
Arabidopsis thaliana, partial (35%)
Length = 869
Score = 83.6 bits (205), Expect = 9e-17
Identities = 52/153 (33%), Positives = 85/153 (54%), Gaps = 12/153 (7%)
Frame = +2
Query: 1 MGRSPCCEKV-GLKKGPWTSEEDQKLLSYIEEHGHGSWRSLPTKAVQDLRGVARAAD*DG 59
M R+P C+K+ G++KG WT+EED+KL++Y+ +G +WR LP A G++R
Sbjct: 92 MARTPSCDKISGMRKGTWTAEEDRKLIAYVTRYGFWNWRQLPKFA-----GLSRCGKSCR 256
Query: 60 L----II*GRILRG------ESLVCKKNKPLFNFMHF*ATATRLPKRTDNEIKNYWNTHL 109
L + I RG E L+ + +K L N + A LP RTDNE+KN+W+T L
Sbjct: 257 LRWLNYLRPNIRRGNFTKDEEELIIRMHKKLGN--KWSTIAAELPGRTDNEVKNHWHTSL 430
Query: 110 KKR-LTKMGIDPVTHKPKNETLLPENHSKNVAN 141
KKR + + + T K++ ++ ++++N
Sbjct: 431 KKRAIDNIVTNEETKSTKSKDIIESTQGRDISN 529
>TC78266 similar to GP|22266673|dbj|BAC07543. myb-related transcription
factor VlMYBB1-1 {Vitis labrusca x Vitis vinifera},
partial (46%)
Length = 665
Score = 79.7 bits (195), Expect = 1e-15
Identities = 50/124 (40%), Positives = 71/124 (56%), Gaps = 11/124 (8%)
Frame = +1
Query: 1 MGRSPCCEKV-GLKKGPWTSEEDQKLLSYIEEHGHGSWRSLPTKAVQDLRGVARAAD*DG 59
M R+P C+K G++KG WT+EED+KL++Y+ +G +WR LP A G++R
Sbjct: 85 MARTPSCDKKSGMRKGTWTAEEDRKLIAYVTRYGCWNWRQLPKFA-----GLSRCGKSCR 249
Query: 60 L----II*GRILRG------ESLVCKKNKPLFNFMHF*ATATRLPKRTDNEIKNYWNTHL 109
L + I RG E L+ + +K L N + A LP RTDNE+KNYW+T L
Sbjct: 250 LRWLNYLRPNIKRGNFTQEEEELIIRMHKNLGN--RWSTIAAELPGRTDNEVKNYWHTSL 423
Query: 110 KKRL 113
KKR+
Sbjct: 424 KKRV 435
>TC78268 weakly similar to PIR|B85064|B85064 MYB-like protein [imported] -
Arabidopsis thaliana, partial (35%)
Length = 888
Score = 77.8 bits (190), Expect = 5e-15
Identities = 49/123 (39%), Positives = 70/123 (56%), Gaps = 11/123 (8%)
Frame = +1
Query: 1 MGRSPCCEKV-GLKKGPWTSEEDQKLLSYIEEHGHGSWRSLPTKAVQDLRGVARAAD*DG 59
M R+P C+K G++KG WT+EED+KL++Y+ +G +WR LP A G++R
Sbjct: 82 MARTPSCDKKSGMRKGTWTAEEDRKLIAYVTRYGCWNWRQLPKFA-----GLSRCGKSCR 246
Query: 60 L----II*GRILRG------ESLVCKKNKPLFNFMHF*ATATRLPKRTDNEIKNYWNTHL 109
L + I RG E L+ + +K L N + A LP RTDNE+KN+W+T L
Sbjct: 247 LRWLNYLRPNIKRGNFTQEEEELIIRMHKKLGN--RWSTIAAELPGRTDNEVKNHWHTSL 420
Query: 110 KKR 112
KKR
Sbjct: 421 KKR 429
>BQ149085 similar to GP|9802570|gb|A F22O13.30 {Arabidopsis thaliana},
partial (16%)
Length = 464
Score = 76.3 bits (186), Expect = 1e-14
Identities = 31/42 (73%), Positives = 37/42 (87%)
Frame = +1
Query: 1 MGRSPCCEKVGLKKGPWTSEEDQKLLSYIEEHGHGSWRSLPT 42
MGR PCCEKVG+KKGPWT EED L+SYI+EHG G+WR++PT
Sbjct: 133 MGRPPCCEKVGIKKGPWTPEEDIILVSYIQEHGPGNWRTVPT 258
>BI266228 homologue to GP|10178078|dbj MYB96 transcription factor-like
protein {Arabidopsis thaliana}, partial (24%)
Length = 384
Score = 75.1 bits (183), Expect = 3e-14
Identities = 30/42 (71%), Positives = 37/42 (87%)
Frame = +1
Query: 1 MGRSPCCEKVGLKKGPWTSEEDQKLLSYIEEHGHGSWRSLPT 42
MGR PCCEK+G+KKGPWT EED L+SYI++HG G+WRS+PT
Sbjct: 55 MGRPPCCEKLGIKKGPWTPEEDIILVSYIQQHGPGNWRSVPT 180
>BQ147546 similar to PIR|H96706|H96 probable transcription factor T22E19.5
[imported] - Arabidopsis thaliana, partial (48%)
Length = 690
Score = 60.5 bits (145), Expect = 8e-10
Identities = 41/119 (34%), Positives = 60/119 (49%), Gaps = 7/119 (5%)
Frame = +3
Query: 8 EKVGLKKGPWTSEEDQKLLSYIEEHGHGSWRSLPTKAVQDLRGVARAAD*DGL-II*GRI 66
E+ L++GPWT EED L+ YI HG G W L A L+ ++ L + I
Sbjct: 150 EENELRRGPWTLEEDSLLIHYIARHGEGRWNMLAKSA--GLKRTGKSCRLRWLNYLKPDI 323
Query: 67 LRG------ESLVCKKNKPLFNFMHF*ATATRLPKRTDNEIKNYWNTHLKKRLTKMGID 119
RG + L+ + + N + A LP RTDNEIKNYW T ++K+ ++ I+
Sbjct: 324 KRGNLTPQEQLLILELHSKXGN--RWSKIAQHLPGRTDNEIKNYWRTRVQKQARQLNIE 494
Database: MTGI
Posted date: Oct 22, 2004 3:39 PM
Number of letters in database: 27,044,181
Number of sequences in database: 36,976
Lambda K H
0.316 0.131 0.392
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 10,433,220
Number of Sequences: 36976
Number of extensions: 148704
Number of successful extensions: 1189
Number of sequences better than 10.0: 129
Number of HSP's better than 10.0 without gapping: 1112
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 1177
length of query: 305
length of database: 9,014,727
effective HSP length: 96
effective length of query: 209
effective length of database: 5,465,031
effective search space: 1142191479
effective search space used: 1142191479
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.3 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.6 bits)
S2: 58 (26.9 bits)
Medicago: description of AC140544.4


