

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
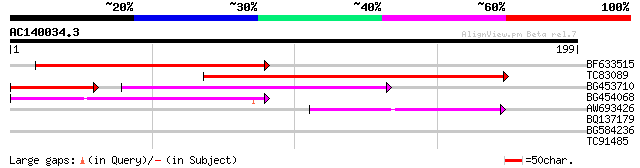
Query= AC140034.3 + phase: 0
(199 letters)
Database: MTGI
36,976 sequences; 27,044,181 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
BF633515 147 2e-36
TC83089 138 1e-33
BG453710 weakly similar to GP|21537057|gb| unknown {Arabidopsis ... 74 4e-21
BG454068 weakly similar to GP|22202804|dbj hypothetical protein~... 66 1e-11
AW693426 homologue to GP|23307580|dbj similar to serine/threonin... 43 9e-05
BQ137179 27 4.0
BG584236 weakly similar to GP|13446644|emb VIP3 protein {Zea may... 27 4.0
TC91485 similar to GP|12830362|emb|CAC29060. anthranilate syntha... 27 6.8
>BF633515
Length = 249
Score = 147 bits (372), Expect = 2e-36
Identities = 70/82 (85%), Positives = 74/82 (89%)
Frame = +2
Query: 10 LDDFACLTHLPIEGRMLDYEKKMPKHEGAALLMTYLGVSQHEAEKICNQEYGGYISYPRL 69
LDD ACLTHLPIEGRML + KKMPKHEGAALLM +LGVSQ EAEKIC QEYGGYISYPRL
Sbjct: 2 LDDVACLTHLPIEGRMLSHGKKMPKHEGAALLMRHLGVSQQEAEKICGQEYGGYISYPRL 181
Query: 70 RDFYTSYLGRANVLAGTEDPEE 91
RDFYT+YLGRAN+LA TEDP E
Sbjct: 182RDFYTTYLGRANLLASTEDPVE 247
>TC83089
Length = 887
Score = 138 bits (348), Expect = 1e-33
Identities = 66/107 (61%), Positives = 83/107 (76%)
Frame = +3
Query: 69 LRDFYTSYLGRANVLAGTEDPEELEELARVRTYCVRWYLMYLVGCLLFDDRSNKRIELIY 128
LR++Y YL A L+ ++ + +EL ++RT CV+WYL+YLVGCLLF D+SNKRIEL+Y
Sbjct: 264 LREYYEGYLDSATRLSDPQNLGDPQELEKIRTTCVKWYLLYLVGCLLFGDKSNKRIELVY 443
Query: 129 LTTMADGYAGMRNYSWGGMTLAYLYGELADACRPGHRALGGSVTLLT 175
LTTM DGYAGMRN+SWGGMTLAYLY L++ PG AL SVTL+T
Sbjct: 444 LTTMEDGYAGMRNHSWGGMTLAYLYHCLSEVSLPGGGALERSVTLVT 584
>BG453710 weakly similar to GP|21537057|gb| unknown {Arabidopsis thaliana},
partial (6%)
Length = 651
Score = 74.3 bits (181), Expect(2) = 4e-21
Identities = 42/95 (44%), Positives = 55/95 (57%)
Frame = -3
Query: 40 LLMTYLGVSQHEAEKICNQEYGGYISYPRLRDFYTSYLGRANVLAGTEDPEELEELARVR 99
L+ LG+S A E G+ISYP L+ Y +L A L EEL+E AR R
Sbjct: 298 LMTRLLGMSDANAMAEIRTESAGHISYPTLKRVYEDHLIEARRLEDPLTREELQERARRR 119
Query: 100 TYCVRWYLMYLVGCLLFDDRSNKRIELIYLTTMAD 134
+CVR L+YLV C+LF ++N+ I+LIYL MAD
Sbjct: 118 QWCVRSLLLYLVDCVLFTYKTNRHIDLIYLDCMAD 14
Score = 43.5 bits (101), Expect(2) = 4e-21
Identities = 18/31 (58%), Positives = 25/31 (80%)
Frame = -1
Query: 1 MPMGEMTVTLDDFACLTHLPIEGRMLDYEKK 31
+PMGEMT+TLDD L H+PI G MLD++++
Sbjct: 414 LPMGEMTITLDDVYNLLHIPIHGCMLDHDEQ 322
>BG454068 weakly similar to GP|22202804|dbj hypothetical protein~predicted by
GeneMark.hmm etc. {Oryza sativa (japonica
cultivar-group)}, partial (5%)
Length = 673
Score = 65.9 bits (159), Expect = 1e-11
Identities = 35/94 (37%), Positives = 51/94 (54%), Gaps = 3/94 (3%)
Frame = +3
Query: 1 MPMGEMTVTLDDFACLTHLPIEGRMLDYEKKMPKHEGAALLMTYLGVSQHEAEKICNQEY 60
+P+GEMT+ LDD +CL H+P+ G +L + + + H+G L+ YLG+ E +
Sbjct: 387 LPVGEMTIALDDISCLLHIPVGGNLL-FHESLSIHQGTEYLVNYLGLEFEEIAAETKRLK 563
Query: 61 GGYISYPRLRDFYTSYLGRANVLA---GTEDPEE 91
+I+Y L YTSYL A A G ED E
Sbjct: 564 SAHITYDTLLSIYTSYLTEAKSYANQPGEEDSME 665
>AW693426 homologue to GP|23307580|dbj similar to serine/threonine
phosphatase PP7 {Oryza sativa (japonica
cultivar-group)}, partial (2%)
Length = 209
Score = 42.7 bits (99), Expect = 9e-05
Identities = 24/69 (34%), Positives = 36/69 (51%)
Frame = +1
Query: 106 YLMYLVGCLLFDDRSNKRIELIYLTTMADGYAGMRNYSWGGMTLAYLYGELADACRPGHR 165
+L+ LVGC LF D+S + + YL D Y+WG LA+LY L A +
Sbjct: 1 FLLCLVGCTLFSDKSTFVVNVAYLEFFRD-LDSCGGYAWGVAALAHLYDNLRYASFHHMK 177
Query: 166 ALGGSVTLL 174
++ G +TL+
Sbjct: 178 SISGYLTLI 204
>BQ137179
Length = 920
Score = 27.3 bits (59), Expect = 4.0
Identities = 14/54 (25%), Positives = 25/54 (45%), Gaps = 2/54 (3%)
Frame = +3
Query: 98 VRTYCVRWYLMYLVGC--LLFDDRSNKRIELIYLTTMADGYAGMRNYSWGGMTL 149
VR +C+ WY+ + + LF+ N R +YL+ D + Y+ G +
Sbjct: 216 VRYFCIAWYIFHFIHV*F*LFNYNMNFRNSFLYLSECGDDTKAVHCYNEYGQII 377
>BG584236 weakly similar to GP|13446644|emb VIP3 protein {Zea mays}, partial
(38%)
Length = 589
Score = 27.3 bits (59), Expect = 4.0
Identities = 9/18 (50%), Positives = 14/18 (77%)
Frame = +1
Query: 176 VRKLKFYIYVNFIFLLYF 193
++K+ YIY+NF F LY+
Sbjct: 412 IKKINIYIYLNF*FFLYY 465
>TC91485 similar to GP|12830362|emb|CAC29060. anthranilate synthase alpha
subunit {Catharanthus roseus}, partial (52%)
Length = 1113
Score = 26.6 bits (57), Expect = 6.8
Identities = 9/17 (52%), Positives = 11/17 (63%)
Frame = -1
Query: 56 CNQEYGGYISYPRLRDF 72
C Q +G YI YPR + F
Sbjct: 384 CRQPWGNYIMYPRYKSF 334
Database: MTGI
Posted date: Oct 22, 2004 3:39 PM
Number of letters in database: 27,044,181
Number of sequences in database: 36,976
Lambda K H
0.325 0.142 0.440
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 6,512,775
Number of Sequences: 36976
Number of extensions: 89716
Number of successful extensions: 601
Number of sequences better than 10.0: 16
Number of HSP's better than 10.0 without gapping: 600
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 601
length of query: 199
length of database: 9,014,727
effective HSP length: 91
effective length of query: 108
effective length of database: 5,649,911
effective search space: 610190388
effective search space used: 610190388
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 15 ( 7.0 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 40 (21.6 bits)
S2: 56 (26.2 bits)
Medicago: description of AC140034.3


