

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC140034.13 + phase: 0 /partial
(433 letters)
Database: MTGI
36,976 sequences; 27,044,181 total letters
Searching..................................................done
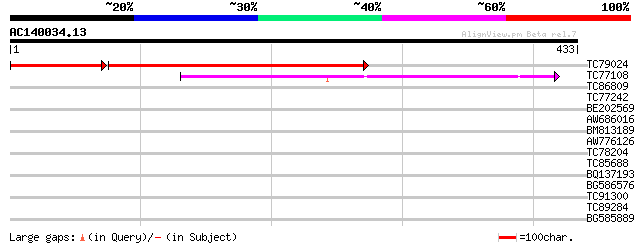
Score E
Sequences producing significant alignments: (bits) Value
TC79024 similar to PIR|A84716|A84716 probable GTP-binding protei... 391 e-152
TC77108 similar to GP|9758030|dbj|BAB08691.1 GTP-binding protein... 206 2e-53
TC86809 homologue to PIR|S35701|S35701 translation elongation fa... 40 0.001
TC77242 similar to GP|22655123|gb|AAM98152.1 putative protein {A... 31 1.1
BE202569 30 1.8
AW686016 weakly similar to GP|3581899|emb hypothetical serine-ri... 30 2.4
BM813189 weakly similar to GP|17104669|gb| unknown protein {Arab... 30 2.4
AW776126 similar to GP|12643044|gb| putative AT-Hook DNA-binding... 29 3.1
TC78204 similar to GP|20197010|gb|AAM14872.1 Expressed protein {... 29 3.1
TC85688 similar to SP|P50346|RLA0_SOYBN 60S acidic ribosomal pro... 29 4.0
BQ137193 28 5.3
BG586576 similar to GP|10129885|db mitochondrial elongation fact... 28 5.3
TC91300 similar to PIR|T33564|T33564 hypothetical protein R160.4... 28 6.9
TC89284 similar to GP|1890154|emb|CAA72432.1 alpha-mannosidase p... 28 9.0
BG585889 weakly similar to SP|Q40287|UFO5_ Flavonol 3-O-glucosyl... 28 9.0
>TC79024 similar to PIR|A84716|A84716 probable GTP-binding protein [imported]
- Arabidopsis thaliana, partial (67%)
Length = 1513
Score = 391 bits (1005), Expect(2) = e-152
Identities = 199/199 (100%), Positives = 199/199 (100%)
Frame = +2
Query: 76 LRNKDSGAEKIEDGKVVKVMKKKGTTMVVTDCAGAGDIISIAGLSSPSIGHTVTTVEIMS 135
LRNKDSGAEKIEDGKVVKVMKKKGTTMVVTDCAGAGDIISIAGLSSPSIGHTVTTVEIMS
Sbjct: 917 LRNKDSGAEKIEDGKVVKVMKKKGTTMVVTDCAGAGDIISIAGLSSPSIGHTVTTVEIMS 1096
Query: 136 ALPTVELDPPTISMTFGVNDSPLAGRDGTHLTGGKIGDRLMAESETNLAINVLPGMSETF 195
ALPTVELDPPTISMTFGVNDSPLAGRDGTHLTGGKIGDRLMAESETNLAINVLPGMSETF
Sbjct: 1097 ALPTVELDPPTISMTFGVNDSPLAGRDGTHLTGGKIGDRLMAESETNLAINVLPGMSETF 1276
Query: 196 EVQGRGELQLGILIENMRREGFELSVSPPKVMYKTDKGQKLEPIEEVTIEVNDEHVGFVM 255
EVQGRGELQLGILIENMRREGFELSVSPPKVMYKTDKGQKLEPIEEVTIEVNDEHVGFVM
Sbjct: 1277 EVQGRGELQLGILIENMRREGFELSVSPPKVMYKTDKGQKLEPIEEVTIEVNDEHVGFVM 1456
Query: 256 EALSHRRAEITDMGPVAGT 274
EALSHRRAEITDMGPVAGT
Sbjct: 1457 EALSHRRAEITDMGPVAGT 1513
Score = 164 bits (415), Expect(2) = e-152
Identities = 74/74 (100%), Positives = 74/74 (100%)
Frame = +3
Query: 1 MRLKVWCLIYLRTWVLQRNNLTSLFFMLLLKKDGHPPHIPRILLPMPKICHSCLMPLYHM 60
MRLKVWCLIYLRTWVLQRNNLTSLFFMLLLKKDGHPPHIPRILLPMPKICHSCLMPLYHM
Sbjct: 600 MRLKVWCLIYLRTWVLQRNNLTSLFFMLLLKKDGHPPHIPRILLPMPKICHSCLMPLYHM 779
Query: 61 SPHQMQTLMHLSKC 74
SPHQMQTLMHLSKC
Sbjct: 780 SPHQMQTLMHLSKC 821
>TC77108 similar to GP|9758030|dbj|BAB08691.1 GTP-binding protein typA
(tyrosine phosphorylated protein A) {Arabidopsis
thaliana}, partial (98%)
Length = 2589
Score = 206 bits (523), Expect = 2e-53
Identities = 121/322 (37%), Positives = 177/322 (54%), Gaps = 32/322 (9%)
Frame = +2
Query: 131 VEIMSALPTVELDPPTISMTFGVNDSPLAGRDGTHLTGGKIGDRLMAESETNLAINVLPG 190
+++ LP+++++ PT+ M+F +N SP GR+G ++T + DRL E E NLA+ V G
Sbjct: 1256 IDLERHLPSIKVEEPTVKMSFSINTSPFVGREGKYVTSRNLRDRLYRELERNLAMKVEDG 1435
Query: 191 -MSETFEVQGRGELQLGILIENMRREGFELSVSPPKVMYKTDKGQKLEPIEE-------- 241
++TF V GRG L + ILIENMRREG+E V PPKV+ K + LEP E
Sbjct: 1436 ETADTFIVSGRGTLHITILIENMRREGYEFMVGPPKVINKKVDDKVLEPYENATCGATGR 1615
Query: 242 -----------------------VTIEVNDEHVGFVMEALSHRRAEITDMGPVAGTVGRT 278
T+EV +EH+G V+E L RR ++ DM V G+ G
Sbjct: 1616 KYSIIDMMNCLDRDQGKCSTCKIATVEVPEEHMGSVVELLGQRRGQMFDMQGV-GSEGTN 1792
Query: 279 RLCLTCPSRGLVGYRSVFSSETRGTGFMHRAFLKYEKFRGPLGNVRKGVLVSVGFGSITL 338
L P+RGL+G R+ + +RGT ++ F Y + G + G LV+ G++T
Sbjct: 1793 LLKYKIPTRGLLGLRNAILTASRGTAILNTIFDSYGPWAGDMSTRDLGSLVAFENGTVTS 1972
Query: 339 HALMSLEARGTLFVSPGMEAYDGMIVGEHSRDTDLDVNPVRAKQLTNVRSVNKDDTVKLI 398
+A+ S + RG +FV PG E Y G I+G H R DL +N + K TN+RS NK+ +V L
Sbjct: 1973 YAVASSQERGQMFVGPGAEVYKGQIIGIHQRPGDLSLNVCKKKAATNIRS-NKEQSVILD 2149
Query: 399 PPRLMTLEEAIGYVASDELIEV 420
P +L++ I Y+ DEL+EV
Sbjct: 2150 TPLDYSLDDCIEYIQEDELVEV 2215
>TC86809 homologue to PIR|S35701|S35701 translation elongation factor EF-G
chloroplast - soybean, partial (91%)
Length = 2674
Score = 40.4 bits (93), Expect = 0.001
Identities = 30/120 (25%), Positives = 57/120 (47%), Gaps = 1/120 (0%)
Frame = +2
Query: 111 GDIISIAGLSSPSIGHTVTTVEIMSALPTVELDPPTISMTFGVNDSPLAGRDGTHLTGGK 170
GDI+++AGL G T+ E L ++ P I + P D + G
Sbjct: 1406 GDIVALAGLKDTITGETLCDPESPVVLERMDFPDPVIKIAI----EPKTKADIDKMAAGL 1573
Query: 171 IGDRLMAESETNLAINVLPGMSETFEVQGRGELQLGILIENMRRE-GFELSVSPPKVMYK 229
+ +A+ + + + +++T ++G GEL L I+++ ++RE E +V P+V Y+
Sbjct: 1574 V---KLAQEDPSFHFSRDEEINQTV-IEGMGELHLEIIVDRLKREYKVEANVGAPQVNYR 1741
Score = 31.2 bits (69), Expect = 0.81
Identities = 15/38 (39%), Positives = 22/38 (57%)
Frame = +1
Query: 236 LEPIEEVTIEVNDEHVGFVMEALSHRRAEITDMGPVAG 273
LEPI +V + +EH+G V+ L+ RR +I G G
Sbjct: 2095 LEPIMKVEVVTPEEHLGDVIGDLNSRRGQINSFGDKPG 2208
>TC77242 similar to GP|22655123|gb|AAM98152.1 putative protein {Arabidopsis
thaliana}, partial (43%)
Length = 1244
Score = 30.8 bits (68), Expect = 1.1
Identities = 14/32 (43%), Positives = 18/32 (55%)
Frame = -1
Query: 35 HPPHIPRILLPMPKICHSCLMPLYHMSPHQMQ 66
H + R+LL P ICHSC +S HQ+Q
Sbjct: 554 HLVSVRRLLLRYPSICHSCHNERRAVSSHQLQ 459
>BE202569
Length = 568
Score = 30.0 bits (66), Expect = 1.8
Identities = 18/62 (29%), Positives = 31/62 (49%)
Frame = +3
Query: 303 TGFMHRAFLKYEKFRGPLGNVRKGVLVSVGFGSITLHALMSLEARGTLFVSPGMEAYDGM 362
TGF H LKY +FRG +G++ L S S T M+ +R ++ G + + +
Sbjct: 51 TGFSHTN-LKYVEFRGCVGSINVVELASHLLRSATSLGKMTFSSRDKAYIGAGRDGPETL 227
Query: 363 IV 364
++
Sbjct: 228 MI 233
>AW686016 weakly similar to GP|3581899|emb hypothetical serine-rich protein
{Schizosaccharomyces pombe}, partial (4%)
Length = 593
Score = 29.6 bits (65), Expect = 2.4
Identities = 19/55 (34%), Positives = 29/55 (52%)
Frame = -1
Query: 16 LQRNNLTSLFFMLLLKKDGHPPHIPRILLPMPKICHSCLMPLYHMSPHQMQTLMH 70
LQ+N + S LLL+ H PH+ +LLP + + + L H+ HQ Q + H
Sbjct: 503 LQQNLIVSHLLQLLLQLPQHLPHLLPLLLPQ-NLVPTHFL*LLHL--HQNQVVTH 348
>BM813189 weakly similar to GP|17104669|gb| unknown protein {Arabidopsis
thaliana}, partial (19%)
Length = 454
Score = 29.6 bits (65), Expect = 2.4
Identities = 17/44 (38%), Positives = 22/44 (49%)
Frame = +2
Query: 8 LIYLRTWVLQRNNLTSLFFMLLLKKDGHPPHIPRILLPMPKICH 51
+ LR+ +RN +L F L L HPPH PR +P CH
Sbjct: 137 IFLLRSHHARRNQSHALPFQLHLHLPLHPPHQPR----LPSSCH 256
>AW776126 similar to GP|12643044|gb| putative AT-Hook DNA-binding protein
{Oryza sativa}, partial (19%)
Length = 742
Score = 29.3 bits (64), Expect = 3.1
Identities = 11/30 (36%), Positives = 17/30 (56%)
Frame = -3
Query: 45 PMPKICHSCLMPLYHMSPHQMQTLMHLSKC 74
P+ +CH C P + + QT++H SKC
Sbjct: 689 PLEFLCHYCRKPCKIQTKIESQTVLHKSKC 600
>TC78204 similar to GP|20197010|gb|AAM14872.1 Expressed protein {Arabidopsis
thaliana}, partial (45%)
Length = 1313
Score = 29.3 bits (64), Expect = 3.1
Identities = 20/54 (37%), Positives = 26/54 (48%), Gaps = 5/54 (9%)
Frame = -3
Query: 22 TSLFFMLLLKKDGHPPHIPRILLPMPKICHSCLM----PL-YHMSPHQMQTLMH 70
++LF LLL + PH PR L+ CHSC PL H+S Q +H
Sbjct: 873 STLFSSLLLSRPELLPHFPRQLM-----CHSCFF**HHPLMLHVSSFSFQKKLH 727
>TC85688 similar to SP|P50346|RLA0_SOYBN 60S acidic ribosomal protein P0.
[Soybean] {Glycine max}, partial (96%)
Length = 1289
Score = 28.9 bits (63), Expect = 4.0
Identities = 23/90 (25%), Positives = 44/90 (48%), Gaps = 7/90 (7%)
Frame = +3
Query: 321 GNVRKGVLVSVGFGSITLHALMSLEARGTLFVSPGM----EAYDG---MIVGEHSRDTDL 373
GNVR + +G +L + L+ + T+ V P + YD ++ E+++ +
Sbjct: 72 GNVRLD*TICLGPLISSLSLFLLLKTQSTMAVKPTKAEKKQNYDAKLCQLIDEYTQILVV 251
Query: 374 DVNPVRAKQLTNVRSVNKDDTVKLIPPRLM 403
+ + V +KQL N+R + D+V L+ M
Sbjct: 252 NADNVGSKQLQNIRQGLRGDSVVLMGKNTM 341
>BQ137193
Length = 936
Score = 28.5 bits (62), Expect = 5.3
Identities = 9/17 (52%), Positives = 13/17 (75%)
Frame = -3
Query: 42 ILLPMPKICHSCLMPLY 58
+L P+ +CH CL+PLY
Sbjct: 508 LLYPVVMLCHECLLPLY 458
>BG586576 similar to GP|10129885|db mitochondrial elongation factor G {Oryza
sativa (japonica cultivar-group)}, partial (30%)
Length = 745
Score = 28.5 bits (62), Expect = 5.3
Identities = 19/74 (25%), Positives = 35/74 (46%), Gaps = 3/74 (4%)
Frame = +1
Query: 167 TGGKIGDRL--MAESETNLAINVLPGMSETFEVQGRGELQLGILIENMRRE-GFELSVSP 223
+GG+ L + + + P +T + G GEL L I +E +RRE + +V
Sbjct: 175 SGGQFSKALNRFQREDPTFRVGLDPESGQTI-ISGMGELHLDIYVERIRREYKVDATVGK 351
Query: 224 PKVMYKTDKGQKLE 237
P+V ++ Q+ +
Sbjct: 352 PRVNFRETVTQRAD 393
>TC91300 similar to PIR|T33564|T33564 hypothetical protein R160.4 -
Caenorhabditis elegans, partial (10%)
Length = 734
Score = 28.1 bits (61), Expect = 6.9
Identities = 22/62 (35%), Positives = 32/62 (51%), Gaps = 8/62 (12%)
Frame = -1
Query: 368 SRDTDLDVNPVRAKQLTNVRSVNKDDTVKLIPPRLMTLE--------EAIGYVASDELIE 419
S + DL VN V+ K+ VRS KDD VK+ P ++E E IG+ ++ I+
Sbjct: 197 SDEKDLKVNAVKKKKKKPVRSKKKDD-VKISPAVTDSVEVVG*SRSFEKIGFQREED*IQ 21
Query: 420 VR 421
R
Sbjct: 20 RR 15
>TC89284 similar to GP|1890154|emb|CAA72432.1 alpha-mannosidase precursor
{Arabidopsis thaliana}, partial (36%)
Length = 1246
Score = 27.7 bits (60), Expect = 9.0
Identities = 20/90 (22%), Positives = 40/90 (44%), Gaps = 4/90 (4%)
Frame = +1
Query: 49 ICHSCLMPLYHMSPHQMQTLMHLSKC*LRNKDSGAEKIE----DGKVVKVMKKKGTTMVV 104
+C +CL+ LYH S + + K ++D G K++ K++ + MVV
Sbjct: 247 VCRTCLILLYHHSWRIQIASLSMLKWHF-SRDGGGSKVKLRNLKSKILSTRVSSNS*MVV 423
Query: 105 TDCAGAGDIISIAGLSSPSIGHTVTTVEIM 134
C I++ L P +G ++ + ++
Sbjct: 424 CACMTRPPRITLT*LIRPLLGINLSKMSLV 513
>BG585889 weakly similar to SP|Q40287|UFO5_ Flavonol 3-O-glucosyltransferase
5 (EC 2.4.1.91) (UDP-glucose flavonoid
3-O-glucosyltransferase 5)., partial (8%)
Length = 304
Score = 27.7 bits (60), Expect = 9.0
Identities = 14/40 (35%), Positives = 20/40 (50%)
Frame = +3
Query: 100 TTMVVTDCAGAGDIISIAGLSSPSIGHTVTTVEIMSALPT 139
+T +T C I L+SP IGH + T+E+ L T
Sbjct: 3 STREITACNLVSQKIHSVLLASPGIGHLIPTIELGKRLTT 122
Database: MTGI
Posted date: Oct 22, 2004 3:39 PM
Number of letters in database: 27,044,181
Number of sequences in database: 36,976
Lambda K H
0.321 0.138 0.405
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 11,908,612
Number of Sequences: 36976
Number of extensions: 168434
Number of successful extensions: 967
Number of sequences better than 10.0: 30
Number of HSP's better than 10.0 without gapping: 959
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 964
length of query: 433
length of database: 9,014,727
effective HSP length: 99
effective length of query: 334
effective length of database: 5,354,103
effective search space: 1788270402
effective search space used: 1788270402
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.9 bits)
S2: 60 (27.7 bits)
Medicago: description of AC140034.13


