

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC140031.2 + phase: 0 /pseudo
(484 letters)
Database: MTGI
36,976 sequences; 27,044,181 total letters
Searching..................................................done
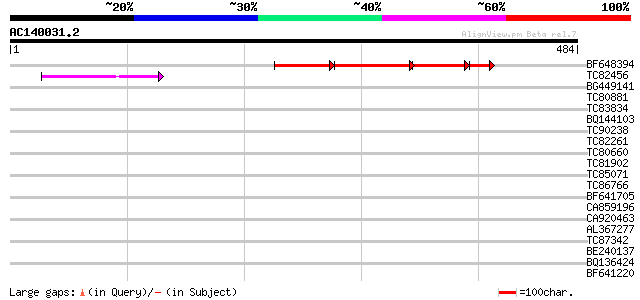
Score E
Sequences producing significant alignments: (bits) Value
BF648394 93 3e-43
TC82456 weakly similar to GP|18057159|gb|AAL58182.1 putative far... 51 1e-06
BG449141 similar to GP|15983442|gb At1g10240/F14N23_12 {Arabidop... 38 0.008
TC80881 similar to PIR|T11622|T11622 extensin class 1 precursor ... 35 0.064
TC83834 similar to PIR|G96565|G96565 F6D8.26 [imported] - Arabid... 35 0.084
BQ144103 homologue to GP|13436053|gb| Similar to aspartyl aminop... 33 0.24
TC90238 similar to PIR|T07126|T07126 magnesium chelatase (EC 4.9... 33 0.32
TC82261 similar to GP|10177249|dbj|BAB10717. contains similarity... 32 0.54
TC80660 similar to PIR|C86475|C86475 unknown protein 42479-4133... 32 0.54
TC81902 weakly similar to GP|18087521|gb|AAL58895.1 AT3g17120/K1... 31 0.93
TC85071 similar to GP|11994207|dbj|BAB01310. gene_id:MPN9.20~unk... 30 1.6
TC86766 weakly similar to PIR|T06291|T06291 extensin homolog T9E... 30 2.1
BF641705 similar to PIR|C86475|C864 unknown protein 42479-41336... 30 2.1
CA859196 weakly similar to PIR|C34715|R6BY 60s acidic ribosomal ... 30 2.7
CA920463 weakly similar to GP|2852684|gb|A resistance protein ca... 30 2.7
AL367277 homologue to PIR|S14974|S149 extensin class I (clone uG... 30 2.7
TC87342 PIR|S41431|S41431 GTP-binding protein ras-like - fava b... 29 3.5
BE240137 29 3.5
BQ136424 weakly similar to PIR|T04859|T048 extensin homolog F28A... 29 3.5
BF641220 weakly similar to PIR|G86203|G86 probable N-arginine di... 28 7.9
>BF648394
Length = 650
Score = 92.8 bits (229), Expect(4) = 3e-43
Identities = 44/68 (64%), Positives = 52/68 (75%)
Frame = -3
Query: 278 FKDVTFYSWLGGYMSRYALGNIVLEEERCRGTLCMDKEICGCV*RKSYGLPCACFVATKI 337
FK VT YS +GG++SRYAL NI LEE CR TLCM +I GCV R SYGLPC C +ATK+
Sbjct: 495 FKYVTLYSGIGGHVSRYALDNIALEETHCRKTLCMYNDI*GCVQRTSYGLPCPCEIATKL 316
Query: 338 RDDKPILL 345
++KPILL
Sbjct: 315 LEEKPILL 292
Score = 54.3 bits (129), Expect(4) = 3e-43
Identities = 30/51 (58%), Positives = 33/51 (63%)
Frame = -1
Query: 227 G*NRLMI**INIGAIVWVIWVLVGRKYLICW*CS*L*YKHHLVRVVPYWKI 277
G N+L I * NI IVWVIWVLVGRKY+ C S L YK LVR + W I
Sbjct: 650 GSNQLXIY*RNIWTIVWVIWVLVGRKYMTCCCFSSLLYKQPLVRALACWNI 498
Score = 53.5 bits (127), Expect(4) = 3e-43
Identities = 26/48 (54%), Positives = 34/48 (70%)
Frame = -2
Query: 345 LDEIHLHWNRLCMGQESNEDCFSVEEEWNGIQERLKKIPNRMNLEMKE 392
+DEI+ HW +L MG+ESNE F VE E I E LKK+P +M +E+KE
Sbjct: 295 IDEIYHHWLKLRMGEESNEVGFCVEVELKAIVEHLKKLPFQMKVEVKE 152
Score = 33.9 bits (76), Expect(4) = 3e-43
Identities = 15/22 (68%), Positives = 19/22 (86%)
Frame = -1
Query: 393 TKSTSQGSQEKSGYCEVYRKGH 414
TKST+QGS+EKS +CEV RK +
Sbjct: 101 TKSTNQGSKEKSRHCEV*RKNY 36
>TC82456 weakly similar to GP|18057159|gb|AAL58182.1 putative far-red
impaired response protein {Oryza sativa}, partial (7%)
Length = 1244
Score = 50.8 bits (120), Expect = 1e-06
Identities = 24/104 (23%), Positives = 50/104 (48%)
Frame = +2
Query: 28 FNNFPTVLVMDSTYQTNMYKMPMFEIVGVTSTDLTYSVGFEFVTHEKEENFVWVLQMMCK 87
+ F V+ +D+TY T+ Y +P+ VGV T +G +++ + W+ +
Sbjct: 848 YEYFGEVITLDTTYLTSKYDLPLVPFVGVNHHGQTILLGCAILSNLDAKTLTWLFTRWLE 1027
Query: 88 LLTSRMNVPKVVVTDKDTTLMNAVSNVLPESFAMNCYFHVQKNV 131
+ + P ++T++D + NA+ P++ C +H+ K V
Sbjct: 1028CMHG--HAPNGIITEEDKAMKNAIEVAFPKARHRWCLWHIMKKV 1153
>BG449141 similar to GP|15983442|gb At1g10240/F14N23_12 {Arabidopsis
thaliana}, partial (33%)
Length = 690
Score = 38.1 bits (87), Expect = 0.008
Identities = 19/92 (20%), Positives = 44/92 (47%)
Frame = +3
Query: 25 LSFFNNFPTVLVMDSTYQTNMYKMPMFEIVGVTSTDLTYSVGFEFVTHEKEENFVWVLQM 84
+ ++ F +V D+T++ + MP+ VG+ + + G + E +F W ++
Sbjct: 354 IQLYDIFGDAVVFDTTHRLTAFDMPLGLWVGINNYGMPCFFGCVLLRDETVRSFSWAIKA 533
Query: 85 MCKLLTSRMNVPKVVVTDKDTTLMNAVSNVLP 116
+ + P+ ++TD++ L A+S +P
Sbjct: 534 FLGFMNGK--APQTILTDQNICLKEALSAEMP 623
>TC80881 similar to PIR|T11622|T11622 extensin class 1 precursor - cowpea,
partial (34%)
Length = 597
Score = 35.0 bits (79), Expect = 0.064
Identities = 23/77 (29%), Positives = 32/77 (40%), Gaps = 8/77 (10%)
Frame = +1
Query: 414 HQLVESHLFGSVLIPKILIVYRHRHQQRHHFLKGNVLVLVKHHVPHF--------HHLLG 465
H HL + +P + + H H HH L + + + HH PHF HHLL
Sbjct: 343 HTSTSHHLLHLLHLPHHMSIRVHHHPLHHHHLHMSTSLRLHHH-PHFPQTMYTSHHHLLP 519
Query: 466 FQISLPRFQSLFWIQLL 482
+ L F +L LL
Sbjct: 520 LHLLLHTFINLLLHHLL 570
>TC83834 similar to PIR|G96565|G96565 F6D8.26 [imported] - Arabidopsis
thaliana, partial (24%)
Length = 1032
Score = 34.7 bits (78), Expect = 0.084
Identities = 15/59 (25%), Positives = 30/59 (50%)
Frame = +3
Query: 31 FPTVLVMDSTYQTNMYKMPMFEIVGVTSTDLTYSVGFEFVTHEKEENFVWVLQMMCKLL 89
F V+ D+TY N Y++P+ +VG+ + +G + E E++ W+ + K +
Sbjct: 831 FSDVIYFDNTYLVNKYEIPLVALVGINHHGQSVLLGCGLLAGEIIESYKWLFRTWIKCI 1007
>BQ144103 homologue to GP|13436053|gb| Similar to aspartyl aminopeptidase
{Homo sapiens}, partial (3%)
Length = 971
Score = 33.1 bits (74), Expect = 0.24
Identities = 13/42 (30%), Positives = 21/42 (49%)
Frame = +1
Query: 430 ILIVYRHRHQQRHHFLKGNVLVLVKHHVPHFHHLLGFQISLP 471
+++ Y H + R+ F+ LVL H +PH H F +P
Sbjct: 490 LILFYHHDFRNRYRFIPPRYLVLPTHLIPHSFHPFHFSYPIP 615
>TC90238 similar to PIR|T07126|T07126 magnesium chelatase (EC 4.99.1.-)
chain chlH - soybean, partial (15%)
Length = 844
Score = 32.7 bits (73), Expect = 0.32
Identities = 14/46 (30%), Positives = 25/46 (53%)
Frame = -1
Query: 25 LSFFNNFPTVLVMDSTYQTNMYKMPMFEIVGVTSTDLTYSVGFEFV 70
LSF+N +++M+ Y N+Y P+F + +T +L +S V
Sbjct: 160 LSFYNKEKVLVMMERWYLRNLYTFPLFFQICLTKVELCFSTQLNLV 23
>TC82261 similar to GP|10177249|dbj|BAB10717. contains similarity to
phytocyanin/early nodulin-like protein~gene_id:K19P17.3,
partial (7%)
Length = 740
Score = 32.0 bits (71), Expect = 0.54
Identities = 19/60 (31%), Positives = 27/60 (44%), Gaps = 4/60 (6%)
Frame = +2
Query: 407 CEVYRKGHQLVESHLFGSVLIPKILIVYR----HRHQQRHHFLKGNVLVLVKHHVPHFHH 462
C + KG + + ++P++LI R H HQ H FL V + HH FHH
Sbjct: 308 CTLANKGIYHTIENFYKHQILPRVLITKRILHLHHHQHLHQFLHQKVPQI--HHHFFFHH 481
>TC80660 similar to PIR|C86475|C86475 unknown protein 42479-41336
[imported] - Arabidopsis thaliana, partial (19%)
Length = 614
Score = 32.0 bits (71), Expect = 0.54
Identities = 17/52 (32%), Positives = 28/52 (53%), Gaps = 2/52 (3%)
Frame = -2
Query: 435 RHRHQQRHHFLKGNV--LVLVKHHVPHFHHLLGFQISLPRFQSLFWIQLLFQ 484
+H H RH K + L+ + HH+ HF H + + I L SL +++L F+
Sbjct: 325 KHEHYHRHPLTKAPLH*LIHMHHHLSHFPHSVNYMILL----SLQYLELNFE 182
>TC81902 weakly similar to GP|18087521|gb|AAL58895.1 AT3g17120/K14A17_24
{Arabidopsis thaliana}, partial (21%)
Length = 788
Score = 31.2 bits (69), Expect = 0.93
Identities = 15/38 (39%), Positives = 19/38 (49%)
Frame = -1
Query: 426 LIPKILIVYRHRHQQRHHFLKGNVLVLVKHHVPHFHHL 463
L P ++ Y H HQQ H LK ++L H P HL
Sbjct: 389 LYPPTILNYIHLHQQHHKNLKNCYMMLQFHFYPFLIHL 276
>TC85071 similar to GP|11994207|dbj|BAB01310. gene_id:MPN9.20~unknown
protein {Arabidopsis thaliana}, partial (28%)
Length = 958
Score = 30.4 bits (67), Expect = 1.6
Identities = 18/42 (42%), Positives = 23/42 (53%)
Frame = +1
Query: 436 HRHQQRHHFLKGNVLVLVKHHVPHFHHLLGFQISLPRFQSLF 477
HRHQ R +L V+ H+PH HHL + LP+ Q LF
Sbjct: 64 HRHQLRRP--PSLLLPPVRTHLPHLHHL--HRSHLPQLQLLF 177
>TC86766 weakly similar to PIR|T06291|T06291 extensin homolog T9E8.80 -
Arabidopsis thaliana, partial (18%)
Length = 1260
Score = 30.0 bits (66), Expect = 2.1
Identities = 15/50 (30%), Positives = 24/50 (48%), Gaps = 1/50 (2%)
Frame = +3
Query: 414 HQLVESHLFGSVLIPKILIVYRHRH-QQRHHFLKGNVLVLVKHHVPHFHH 462
HQ + H L+ + ++RH HH + ++ L+ HH PH HH
Sbjct: 252 HQSIHHHRHLHHLV*NLHHLHRHLV*NLHHHHRQRHIKHLITHHHPHHHH 401
>BF641705 similar to PIR|C86475|C864 unknown protein 42479-41336 [imported]
- Arabidopsis thaliana, partial (19%)
Length = 526
Score = 30.0 bits (66), Expect = 2.1
Identities = 13/38 (34%), Positives = 20/38 (52%), Gaps = 2/38 (5%)
Frame = -2
Query: 435 RHRHQQRHHFLKGNV--LVLVKHHVPHFHHLLGFQISL 470
+H H RH K + L+ + HH+ HF H + + I L
Sbjct: 249 KHEHYHRHPLTKAPLH*LIHMHHHLSHFPHSVNYMILL 136
>CA859196 weakly similar to PIR|C34715|R6BY 60s acidic ribosomal protein
p1-alpha - fission yeast (Schizosaccharomyces pombe),
partial (88%)
Length = 512
Score = 29.6 bits (65), Expect = 2.7
Identities = 19/64 (29%), Positives = 31/64 (47%), Gaps = 6/64 (9%)
Frame = -1
Query: 414 HQLVESHLFGSVLIPKILIVYRHRHQQRHHFLKGNVLVLVKHH----VPHFHHLLG--FQ 467
H + L V + +L + H H++ HH + LVL++ H + H H L +Q
Sbjct: 371 HHQILLFLLQQVFLLLLLFLPLHHHRRHHHQMLHRPLVLLRRHQYSSINH*HFCLQVLWQ 192
Query: 468 ISLP 471
I+LP
Sbjct: 191 INLP 180
>CA920463 weakly similar to GP|2852684|gb|A resistance protein candidate
{Lactuca sativa}, partial (6%)
Length = 810
Score = 29.6 bits (65), Expect = 2.7
Identities = 12/34 (35%), Positives = 20/34 (58%), Gaps = 5/34 (14%)
Frame = +3
Query: 436 HRHQQRHHFLKGNVLVLVK-----HHVPHFHHLL 464
H+H QRH L+G +L++ + H+P F+ L
Sbjct: 105 HKHYQRHRNLEGKILIVARAKIDYRHLPRFYPCL 206
>AL367277 homologue to PIR|S14974|S149 extensin class I (clone uG-18) -
tomato (fragment), partial (46%)
Length = 254
Score = 29.6 bits (65), Expect = 2.7
Identities = 19/62 (30%), Positives = 26/62 (41%), Gaps = 11/62 (17%)
Frame = +2
Query: 414 HQLVESHLFGSVLIPKILIVYRHRHQQR---HHFLKGNVLVLVKHHV--------PHFHH 462
H ++ HL L ++I+ H H R HH L L K+H+ PH HH
Sbjct: 47 HTIISLHLHHHHLPHHLIIISHHPHHPRPLLHHTTTSLHLHLKKYHILHITISHHPHHHH 226
Query: 463 LL 464
L
Sbjct: 227 PL 232
>TC87342 PIR|S41431|S41431 GTP-binding protein ras-like - fava bean,
complete
Length = 995
Score = 29.3 bits (64), Expect = 3.5
Identities = 15/33 (45%), Positives = 19/33 (57%), Gaps = 4/33 (12%)
Frame = +2
Query: 352 WNRLCMGQE----SNEDCFSVEEEWNGIQERLK 380
W R+C GQ S E CFS EE W ++ RL+
Sbjct: 638 WKRVCAGQRRENRS*E*CFSFEESWLLLKLRLE 736
>BE240137
Length = 269
Score = 29.3 bits (64), Expect = 3.5
Identities = 22/81 (27%), Positives = 36/81 (44%), Gaps = 5/81 (6%)
Frame = -2
Query: 389 EMKETKSTSQGSQEKSGYCEVYRKGHQLVESHLFGSVLIPKILIVYRH--RHQQRHHFLK 446
E KE + QG Q+K + +ESH GS P+ Y+H +Q+ +
Sbjct: 223 EQKEQQLRIQGFQQK-------KMKQNSIESHSIGSAFQPREQDRYKHMLERKQKRVLQE 65
Query: 447 GNVLV---LVKHHVPHFHHLL 464
N+L+ + H + HH+L
Sbjct: 64 YNILLHFDIYLHRLQVLHHIL 2
>BQ136424 weakly similar to PIR|T04859|T048 extensin homolog F28A21.80 -
Arabidopsis thaliana, partial (5%)
Length = 779
Score = 29.3 bits (64), Expect = 3.5
Identities = 11/32 (34%), Positives = 21/32 (65%), Gaps = 1/32 (3%)
Frame = +2
Query: 432 IVYRHRHQQRHH-FLKGNVLVLVKHHVPHFHH 462
+ +R+ H + HH ++KG +L++ H+ H HH
Sbjct: 80 LYHRYIHPRHHHLYMKGLFHLLLESHMHHHHH 175
>BF641220 weakly similar to PIR|G86203|G86 probable N-arginine dibasic
convertase [imported] - Arabidopsis thaliana, partial
(5%)
Length = 634
Score = 28.1 bits (61), Expect = 7.9
Identities = 13/34 (38%), Positives = 15/34 (43%)
Frame = -3
Query: 431 LIVYRHRHQQRHHFLKGNVLVLVKHHVPHFHHLL 464
L ++ H HH L HH PH HHLL
Sbjct: 449 LXLHPHHLHPHHHLL---------HHPPHLHHLL 375
Database: MTGI
Posted date: Oct 22, 2004 3:39 PM
Number of letters in database: 27,044,181
Number of sequences in database: 36,976
Lambda K H
0.342 0.150 0.517
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 18,991,420
Number of Sequences: 36976
Number of extensions: 324884
Number of successful extensions: 3244
Number of sequences better than 10.0: 48
Number of HSP's better than 10.0 without gapping: 3101
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 3216
length of query: 484
length of database: 9,014,727
effective HSP length: 100
effective length of query: 384
effective length of database: 5,317,127
effective search space: 2041776768
effective search space used: 2041776768
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 15 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 39 (22.0 bits)
S2: 60 (27.7 bits)
Medicago: description of AC140031.2


