

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
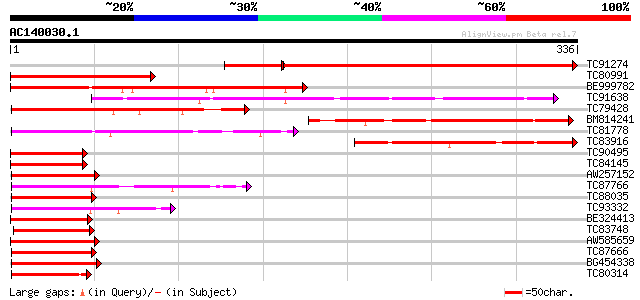
Query= AC140030.1 + phase: 0
(336 letters)
Database: MTGI
36,976 sequences; 27,044,181 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
TC91274 similar to PIR|T45710|T45710 H-protein promoter binding ... 373 e-120
TC80991 similar to GP|15983797|gb|AAL10495.1 AT5g39660/MIJ24_130... 189 1e-48
BE999782 similar to PIR|T09661|T096 ascorbate oxidase promoter-b... 171 3e-43
TC91638 similar to GP|15983797|gb|AAL10495.1 AT5g39660/MIJ24_130... 162 2e-40
TC79428 similar to PIR|T09661|T09661 ascorbate oxidase promoter-... 137 4e-33
BM814241 similar to GP|15983797|gb| AT5g39660/MIJ24_130 {Arabido... 126 1e-29
TC81778 similar to GP|7242908|dbj|BAA92506.1 ESTs C23582(S11122)... 110 6e-25
TC83916 similar to PIR|T09661|T09661 ascorbate oxidase promoter-... 110 6e-25
TC90495 similar to PIR|H84752|H84752 probable DOF zinc finger pr... 99 2e-21
TC84145 similar to GP|7242908|dbj|BAA92506.1 ESTs C23582(S11122)... 98 5e-21
AW257152 similar to PIR|D86409|D864 hypothetical protein AAF9842... 89 2e-18
TC87766 similar to GP|10177472|dbj|BAB10863. contains similarity... 88 4e-18
TC88035 similar to GP|6092016|dbj|BAA85655.1 elicitor-responsive... 87 1e-17
TC93332 similar to PIR|T48267|T48267 probable zinc finger protei... 87 1e-17
BE324413 homologue to GP|10176912|dbj zinc finger protein {Arabi... 87 1e-17
TC83748 homologue to PIR|T02375|T02375 finger protein BBF3 - com... 86 2e-17
AW585659 similar to GP|2262110|gb|A zinc finger protein isolog {... 86 2e-17
TC87666 homologue to GP|4996640|dbj|BAA78572.1 Dof zinc finger p... 86 2e-17
BG454338 homologue to PIR|T52044|T520 dof zinc finger protein [i... 84 6e-17
TC80314 similar to PIR|D86409|D86409 hypothetical protein AAF984... 82 3e-16
>TC91274 similar to PIR|T45710|T45710 H-protein promoter binding factor-2a
[imported] - Arabidopsis thaliana, partial (9%)
Length = 959
Score = 373 bits (958), Expect(2) = e-120
Identities = 174/174 (100%), Positives = 174/174 (100%)
Frame = +2
Query: 163 PFLPYTWNSAMPPPTFCPPNYPLAFYTPVTPPAYWGCMPPPWNIPCISPGSASVNESDSA 222
PFLPYTWNSAMPPPTFCPPNYPLAFYTPVTPPAYWGCMPPPWNIPCISPGSASVNESDSA
Sbjct: 140 PFLPYTWNSAMPPPTFCPPNYPLAFYTPVTPPAYWGCMPPPWNIPCISPGSASVNESDSA 319
Query: 223 HSSVPTLGKHSRDGNIITSVNSPKEKPETEHNSTESSVLIPKTLRFDDPSEAAKSSLWSK 282
HSSVPTLGKHSRDGNIITSVNSPKEKPETEHNSTESSVLIPKTLRFDDPSEAAKSSLWSK
Sbjct: 320 HSSVPTLGKHSRDGNIITSVNSPKEKPETEHNSTESSVLIPKTLRFDDPSEAAKSSLWSK 499
Query: 283 LGINNDKANSLNGGGMFNGFQSKGKDMNHSVGTSPLMYANPAALSRSRTFHEET 336
LGINNDKANSLNGGGMFNGFQSKGKDMNHSVGTSPLMYANPAALSRSRTFHEET
Sbjct: 500 LGINNDKANSLNGGGMFNGFQSKGKDMNHSVGTSPLMYANPAALSRSRTFHEET 661
Score = 76.3 bits (186), Expect(2) = e-120
Identities = 35/37 (94%), Positives = 36/37 (96%)
Frame = +1
Query: 128 VTASNSSERKSHTSTNGLVDKGVEGFPPQLQHIPSPF 164
+ ASNSSERKSHTSTNGLVDKGVEGFPPQLQHIPSPF
Sbjct: 34 LXASNSSERKSHTSTNGLVDKGVEGFPPQLQHIPSPF 144
Score = 29.3 bits (64), Expect = 2.2
Identities = 15/23 (65%), Positives = 16/23 (69%), Gaps = 1/23 (4%)
Frame = +3
Query: 118 GENDHSIGVSVTASN-SSERKSH 139
GENDHSIGVSVT E+K H
Sbjct: 3 GENDHSIGVSVTGFKFIREKKPH 71
>TC80991 similar to GP|15983797|gb|AAL10495.1 AT5g39660/MIJ24_130
{Arabidopsis thaliana}, partial (19%)
Length = 782
Score = 189 bits (481), Expect = 1e-48
Identities = 86/86 (100%), Positives = 86/86 (100%)
Frame = +3
Query: 1 MDTKFCYFNNYNVNQPRHFCKKCQRYWTAGGTMRNVPVGAGRRKNKNLSTSHYRHLIIPE 60
MDTKFCYFNNYNVNQPRHFCKKCQRYWTAGGTMRNVPVGAGRRKNKNLSTSHYRHLIIPE
Sbjct: 522 MDTKFCYFNNYNVNQPRHFCKKCQRYWTAGGTMRNVPVGAGRRKNKNLSTSHYRHLIIPE 701
Query: 61 GAKLNSPNGIHSLGNGAAVLTFDSNP 86
GAKLNSPNGIHSLGNGAAVLTFDSNP
Sbjct: 702 GAKLNSPNGIHSLGNGAAVLTFDSNP 779
>BE999782 similar to PIR|T09661|T096 ascorbate oxidase promoter-binding
protein AOBP - winter squash, partial (17%)
Length = 630
Score = 171 bits (434), Expect = 3e-43
Identities = 97/197 (49%), Positives = 123/197 (62%), Gaps = 21/197 (10%)
Frame = +2
Query: 1 MDTKFCYFNNYNVNQPRHFCKKCQRYWTAGGTMRNVPVGAGRRKNKNLSTSHYRHLIIPE 60
MDTKFCY+NNYNVNQPRHFCKKCQRYWTAGG MRNVPVGAGRRKNKN S SH+R + +PE
Sbjct: 38 MDTKFCYYNNYNVNQPRHFCKKCQRYWTAGGAMRNVPVGAGRRKNKN-SASHFRQITVPE 214
Query: 61 GAKLN----SPNGIH--SLGNGAAVLTFDSNPPLCNTMASVLNIAERTQN-CVSNGFHHA 113
A N SPNG+H SL V TF ++ PLC +M S LN+A++ N NGF
Sbjct: 215 TAVQNSLSDSPNGVHHPSLNCNGTVFTFRTDTPLCESMESALNLADQGVNISQKNGFIRP 394
Query: 114 SS------YGGE---NDHSIGVSVTASNSSERKSHTSTNGLVDKGVEGFPPQLQHIPS-- 162
+ Y GE ++HSI S T++ +E + +S+ V + F PQ+ + PS
Sbjct: 395 EALRIHVPYVGEEKSDEHSIKSSDTSTTLTEDAAASSSVEQVMPNCQSFQPQVPYYPSAP 574
Query: 163 ---PFLPYTWNSAMPPP 176
P+ P W+S + PP
Sbjct: 575 WLLPWSPSQWSSQVQPP 625
>TC91638 similar to GP|15983797|gb|AAL10495.1 AT5g39660/MIJ24_130
{Arabidopsis thaliana}, partial (16%)
Length = 914
Score = 162 bits (409), Expect = 2e-40
Identities = 115/297 (38%), Positives = 153/297 (50%), Gaps = 20/297 (6%)
Frame = +2
Query: 49 STSHYRHLIIPEGAKLNSPNGIH-SLGNGAAVLTFDSNPPLCNTMASVLNIAERT-QNCV 106
S+SHYR + + E NS IH S+ +LTF SN P+C +MASVL A++T QN
Sbjct: 2 SSSHYRQITVSEATLQNSR--IHPSVKCNGTILTFGSNSPVCESMASVLKHADKTMQNYT 175
Query: 107 SNGFH----------HASSYGGENDHSIGVSVTASNSSERKSHTSTNGLVDKGVEGFPPQ 156
NG+H H S GE D S SVT++ S+E + + FPPQ
Sbjct: 176 RNGYHKHEELRICVPHTSEEQGE-DQSNKSSVTSTKSTEGATTNVSQEQAMWNDHSFPPQ 352
Query: 157 LQHIPS------PFLPYTWNSAMPPPTFCPPNYPLAFYTPVTPPAYWGC-MPPPWNIPC- 208
+ P P+ P +S +PPP FCPP + + FY T YWGC MP WNIP
Sbjct: 353 GGYFPHGTPWHLPWNPVQMSSPIPPPAFCPPGFSMPFYPATT---YWGCTMPSAWNIPRQ 523
Query: 209 ISPGSASVNESDSAHSSVPTLGKHSRDGNIITSVNSPKEKPETEHNSTESSVLIPKTLRF 268
P S + DS +S PTLGKHSR+ N++ S +K +E E S+ PKTLR
Sbjct: 524 AQPSSPNGANHDSTPNS-PTLGKHSREDNMLKSSEGDGKKEISE----EKSLWFPKTLRI 688
Query: 269 DDPSEAAKSSLWSKLGINNDKANSLNGGGMFNGFQSKGKDMNHSVGTSPLMYANPAA 325
DD EA KSS W+ LGI N+ A+S+ +F F K S+GTS A+ ++
Sbjct: 689 DDSEEAEKSSFWTTLGIKNNNADSVPPRRLFQAFPFK-V*REKSLGTSVFSLASQSS 856
Score = 36.6 bits (83), Expect = 0.014
Identities = 19/36 (52%), Positives = 22/36 (60%)
Frame = +3
Query: 299 FNGFQSKGKDMNHSVGTSPLMYANPAALSRSRTFHE 334
F SK + NH V S ++ ANPAALSRS FHE
Sbjct: 780 FRHSPSKCDEKNHLVQVSSVLQANPAALSRSLHFHE 887
>TC79428 similar to PIR|T09661|T09661 ascorbate oxidase promoter-binding
protein AOBP - winter squash, partial (27%)
Length = 997
Score = 137 bits (346), Expect = 4e-33
Identities = 78/150 (52%), Positives = 95/150 (63%), Gaps = 9/150 (6%)
Frame = +1
Query: 2 DTKFCYFNNYNVNQPRHFCKKCQRYWTAGGTMRNVPVGAGRRKNKNLSTSHYRHLIIPE- 60
DTKFCY+NNYNVNQPR+FCK CQRYWTAGGTMRNVPVGAGRRKNKN S+SHYRH+ I E
Sbjct: 550 DTKFCYYNNYNVNQPRYFCKACQRYWTAGGTMRNVPVGAGRRKNKNNSSSHYRHITISEA 729
Query: 61 --GAKLNSPNGIHSLGN---GAAVLTFD-SNPPLCNTMASVLNIAER--TQNCVSNGFHH 112
A++ SPNG H L N VL F +P + ++M++ LN AE+ + +NG
Sbjct: 730 LDAARIISPNGTHHLQNLKTNGRVLNFGLDHPHIYDSMSNDLNPAEKKVLNDTRNNGDRF 909
Query: 113 ASSYGGENDHSIGVSVTASNSSERKSHTST 142
+S+ SVT S S E T
Sbjct: 910 SSA----------SSVTVSKSMEESGKNMT 969
>BM814241 similar to GP|15983797|gb| AT5g39660/MIJ24_130 {Arabidopsis
thaliana}, partial (10%)
Length = 527
Score = 126 bits (316), Expect = 1e-29
Identities = 75/162 (46%), Positives = 98/162 (60%), Gaps = 5/162 (3%)
Frame = +3
Query: 178 FCPPNYPLAFYTPVTPPAYWG-CMPPPWNIPCI----SPGSASVNESDSAHSSVPTLGKH 232
F P YP PPAYWG MP WN P + SP SA+VN ++ PTLGKH
Sbjct: 6 FAMPLYP--------PPAYWGFSMPGAWNNPWLAQPSSPNSATVNSGPNS----PTLGKH 149
Query: 233 SRDGNIITSVNSPKEKPETEHNSTESSVLIPKTLRFDDPSEAAKSSLWSKLGINNDKANS 292
SR+ +++ +S + +N E S+ +PKTLR DD EA KSS+ + LGI NDKA++
Sbjct: 150 SREESMLKPTDSTGS--DEGNNKEEKSLWVPKTLRIDDLGEAEKSSILTTLGIKNDKADA 323
Query: 293 LNGGGMFNGFQSKGKDMNHSVGTSPLMYANPAALSRSRTFHE 334
+ GGG+F F SK + + SV SP M ANPAA+SRS +FHE
Sbjct: 324 IRGGGLFKAFASKSNEKD-SVQNSPAMQANPAAMSRSISFHE 446
>TC81778 similar to GP|7242908|dbj|BAA92506.1 ESTs C23582(S11122)
AU056531(S20663) correspond to a region of the predicted
gene.~Similar to AOBP, partial (17%)
Length = 1081
Score = 110 bits (276), Expect = 6e-25
Identities = 73/195 (37%), Positives = 97/195 (49%), Gaps = 25/195 (12%)
Frame = +3
Query: 2 DTKFCYFNNYNVNQPRHFCKKCQRYWTAGGTMRNVPVGAGRRKNKNLSTSHYRHLII--- 58
DTKFCYFNNYNVNQPRHFC+ C RYWT GGTMRNV VGAGRRKNK+++ S YRH+I+
Sbjct: 330 DTKFCYFNNYNVNQPRHFCRSCHRYWTNGGTMRNVAVGAGRRKNKHIA-SQYRHMIVASD 506
Query: 59 ---------------PEGAKLNSPNGIHSLGNGAAVLTFD-SNPPLCNTMASVLNIAERT 102
G+ L S + VL F N L + S+LN+
Sbjct: 507 GIPTASLETNDSSRYQHGSNLESAAVFRCSNDNGIVLKFGRENASLDESKGSMLNLMNNR 686
Query: 103 QNCVSNGFHHASSYGGENDHSIGVSVTASNSSERKSHTSTNGLVD------KGVEGFPPQ 156
++ ++G + GE + S+ V SS HT N L + K ++ +P
Sbjct: 687 RHVDASG--NNCRENGEEETSLCV------SSVTNGHTRGNELFESEQNRSKPMQSYPAS 842
Query: 157 LQHIPSPFLPYTWNS 171
IP + WN+
Sbjct: 843 SWIIP---MNQRWNN 878
>TC83916 similar to PIR|T09661|T09661 ascorbate oxidase promoter-binding
protein AOBP - winter squash, partial (12%)
Length = 703
Score = 110 bits (276), Expect = 6e-25
Identities = 63/134 (47%), Positives = 83/134 (61%), Gaps = 2/134 (1%)
Frame = +3
Query: 205 NIPCISPGSASVNESDSAHSSVPTLGKHSRDGNIITSVNSPKEKPETEHNSTESS--VLI 262
N+P P + + S S PTLGKHSRD + N+ ++ +TE + + + VL+
Sbjct: 3 NVPWFPPHTPATTPRSSPKS--PTLGKHSRDDDNTNDENAKQDSLQTEESPKQRNGCVLV 176
Query: 263 PKTLRFDDPSEAAKSSLWSKLGINNDKANSLNGGGMFNGFQSKGKDMNHSVGTSPLMYAN 322
PKTLR DDP+EAAKSS+W LGI N+ L+ GGM FQSK NH V TSP++ AN
Sbjct: 177 PKTLRIDDPTEAAKSSIWETLGIKNE---GLSRGGMTKAFQSKKDGKNH-VQTSPMLMAN 344
Query: 323 PAALSRSRTFHEET 336
PAAL+RS FHE +
Sbjct: 345 PAALARSLNFHENS 386
>TC90495 similar to PIR|H84752|H84752 probable DOF zinc finger protein
[imported] - Arabidopsis thaliana, partial (41%)
Length = 620
Score = 99.0 bits (245), Expect = 2e-21
Identities = 41/46 (89%), Positives = 43/46 (93%)
Frame = +3
Query: 1 MDTKFCYFNNYNVNQPRHFCKKCQRYWTAGGTMRNVPVGAGRRKNK 46
M+TKFCYFNNYNVNQPRHFCK CQRYWTAGG +RNVPVGAGRRK K
Sbjct: 276 METKFCYFNNYNVNQPRHFCKSCQRYWTAGGALRNVPVGAGRRKAK 413
>TC84145 similar to GP|7242908|dbj|BAA92506.1 ESTs C23582(S11122)
AU056531(S20663) correspond to a region of the predicted
gene.~Similar to AOBP, partial (16%)
Length = 673
Score = 97.8 bits (242), Expect = 5e-21
Identities = 38/46 (82%), Positives = 44/46 (95%)
Frame = +3
Query: 1 MDTKFCYFNNYNVNQPRHFCKKCQRYWTAGGTMRNVPVGAGRRKNK 46
++TKFCYFNNYNVNQPRHFCK CQRYWTAGG +RNVP+GAG+R+NK
Sbjct: 267 LETKFCYFNNYNVNQPRHFCKNCQRYWTAGGVIRNVPIGAGKRRNK 404
>AW257152 similar to PIR|D86409|D864 hypothetical protein AAF98424.1
[imported] - Arabidopsis thaliana, partial (20%)
Length = 587
Score = 89.0 bits (219), Expect = 2e-18
Identities = 35/52 (67%), Positives = 44/52 (84%)
Frame = +1
Query: 2 DTKFCYFNNYNVNQPRHFCKKCQRYWTAGGTMRNVPVGAGRRKNKNLSTSHY 53
+TKFCY+NNY+++QPRHFCK C+RYWT GGT+RNVPVG G RKNK L ++
Sbjct: 148 NTKFCYYNNYSLSQPRHFCKACKRYWTRGGTLRNVPVGGGCRKNKRLKRPNH 303
>TC87766 similar to GP|10177472|dbj|BAB10863. contains similarity to dof
zinc finger protein~gene_id:MQB2.26 {Arabidopsis
thaliana}, partial (21%)
Length = 1516
Score = 88.2 bits (217), Expect = 4e-18
Identities = 59/155 (38%), Positives = 78/155 (50%), Gaps = 13/155 (8%)
Frame = +2
Query: 2 DTKFCYFNNYNVNQPRHFCKKCQRYWTAGGTMRNVPVGAGRRKNKN-----------LST 50
+TKFCY+NNYN++QPRHFCK C+RYWT GG +RNVPVG G R+N L
Sbjct: 353 NTKFCYYNNYNLSQPRHFCKTCRRYWTKGGALRNVPVGGGCRRNNKRRKGNSVSKSPLKP 532
Query: 51 SHYRHLIIPEGAKLNSPNGIHSLGNGAAVLTFDSNPPLCNTMASV--LNIAERTQNCVSN 108
+H HL I GA G+ +A SN N+ ++ N + S
Sbjct: 533 NHNDHLQIGTGAAAG--------GSSSASANSSSNNGCTNSNVNIGMPNFPTQFPFVTSL 688
Query: 109 GFHHASSYGGENDHSIGVSVTASNSSERKSHTSTN 143
H+ +SY E IG S+ A N S + T+TN
Sbjct: 689 HRHNDNSYASE---GIG-SLLAKNMS---NSTTTN 772
>TC88035 similar to GP|6092016|dbj|BAA85655.1 elicitor-responsive Dof
protein ERDP {Pisum sativum}, partial (77%)
Length = 1397
Score = 86.7 bits (213), Expect = 1e-17
Identities = 35/50 (70%), Positives = 42/50 (84%)
Frame = +3
Query: 2 DTKFCYFNNYNVNQPRHFCKKCQRYWTAGGTMRNVPVGAGRRKNKNLSTS 51
+TKFCY+NNY++ QPR+FCK C+RYWT GGT+RNVPVG G RKNK S S
Sbjct: 264 NTKFCYYNNYSLTQPRYFCKSCRRYWTKGGTLRNVPVGGGCRKNKRSSPS 413
>TC93332 similar to PIR|T48267|T48267 probable zinc finger protein -
Arabidopsis thaliana, partial (20%)
Length = 666
Score = 86.7 bits (213), Expect = 1e-17
Identities = 46/104 (44%), Positives = 61/104 (58%), Gaps = 7/104 (6%)
Frame = +1
Query: 2 DTKFCYFNNYNVNQPRHFCKKCQRYWTAGGTMRNVPVGAGRRKNK-----NLSTSHYRHL 56
+TKFCY+NNYN++QPRHFCK C+RYWT GG +RNVPVG G R++K N S++ +
Sbjct: 154 NTKFCYYNNYNLSQPRHFCKTCRRYWTRGGALRNVPVGGGCRRSKKTKKSNTSSTPSKSP 333
Query: 57 IIPEGAK--LNSPNGIHSLGNGAAVLTFDSNPPLCNTMASVLNI 98
I K S + IHS +N P N M S+ N+
Sbjct: 334 ISNSNDKEYSTSASAIHSTSTPLGEFLHPTNRP--NYMTSLQNL 459
>BE324413 homologue to GP|10176912|dbj zinc finger protein {Arabidopsis
thaliana}, partial (21%)
Length = 339
Score = 86.7 bits (213), Expect = 1e-17
Identities = 34/49 (69%), Positives = 42/49 (85%)
Frame = +2
Query: 1 MDTKFCYFNNYNVNQPRHFCKKCQRYWTAGGTMRNVPVGAGRRKNKNLS 49
++TKFCY+NNYN++QPRHFCK C+RYWT GG +RNVPVG G RK+K S
Sbjct: 47 INTKFCYYNNYNLSQPRHFCKNCRRYWTKGGVLRNVPVGGGCRKSKRSS 193
>TC83748 homologue to PIR|T02375|T02375 finger protein BBF3 - common tobacco
(fragment), partial (60%)
Length = 658
Score = 86.3 bits (212), Expect = 2e-17
Identities = 33/48 (68%), Positives = 43/48 (88%)
Frame = +3
Query: 3 TKFCYFNNYNVNQPRHFCKKCQRYWTAGGTMRNVPVGAGRRKNKNLST 50
TKFCY+NNY+++QPR+FCK C+RYWT GGT+RN+PVG G RKNK +S+
Sbjct: 405 TKFCYYNNYSLSQPRYFCKTCRRYWTKGGTLRNIPVGGGCRKNKKVSS 548
>AW585659 similar to GP|2262110|gb|A zinc finger protein isolog {Arabidopsis
thaliana}, partial (27%)
Length = 483
Score = 85.9 bits (211), Expect = 2e-17
Identities = 35/54 (64%), Positives = 46/54 (84%), Gaps = 1/54 (1%)
Frame = +2
Query: 1 MDTKFCYFNNYNVNQPRHFCKKCQRYWTAGGTMRNVPVGAGRRK-NKNLSTSHY 53
++TKFCY+NNY++ QPR+FCK C+RYWT GG++RN+PVG G RK NKN S+S Y
Sbjct: 227 INTKFCYYNNYSLTQPRYFCKTCRRYWTQGGSIRNIPVGGGSRKNNKNRSSSSY 388
>TC87666 homologue to GP|4996640|dbj|BAA78572.1 Dof zinc finger protein
{Oryza sativa}, partial (20%)
Length = 1828
Score = 85.9 bits (211), Expect = 2e-17
Identities = 33/50 (66%), Positives = 43/50 (86%)
Frame = +2
Query: 2 DTKFCYFNNYNVNQPRHFCKKCQRYWTAGGTMRNVPVGAGRRKNKNLSTS 51
+TKFCY+NNY++ QPR+FCK C+RYWT GG++RNVPVG G RKNK + +S
Sbjct: 731 NTKFCYYNNYSLTQPRYFCKTCRRYWTEGGSLRNVPVGGGSRKNKKIPSS 880
>BG454338 homologue to PIR|T52044|T520 dof zinc finger protein [imported] -
Arabidopsis thaliana, partial (37%)
Length = 678
Score = 84.3 bits (207), Expect = 6e-17
Identities = 33/53 (62%), Positives = 42/53 (78%)
Frame = +1
Query: 2 DTKFCYFNNYNVNQPRHFCKKCQRYWTAGGTMRNVPVGAGRRKNKNLSTSHYR 54
+TKFCY+NNYN++QPRHFCK C+RYWT GG +RN+PVG G RK S++ R
Sbjct: 238 NTKFCYYNNYNLSQPRHFCKNCKRYWTKGGALRNIPVGGGTRKVTKRSSNSKR 396
>TC80314 similar to PIR|D86409|D86409 hypothetical protein AAF98424.1
[imported] - Arabidopsis thaliana, partial (19%)
Length = 986
Score = 82.0 bits (201), Expect = 3e-16
Identities = 33/47 (70%), Positives = 41/47 (87%)
Frame = +1
Query: 2 DTKFCYFNNYNVNQPRHFCKKCQRYWTAGGTMRNVPVGAGRRKNKNL 48
+TKFCY+NNYN +QPRHFC+ C+R+WT GGT+RNVPVG G RKNK +
Sbjct: 475 NTKFCYYNNYNKSQPRHFCRACKRHWTKGGTLRNVPVGGG-RKNKRI 612
Database: MTGI
Posted date: Oct 22, 2004 3:39 PM
Number of letters in database: 27,044,181
Number of sequences in database: 36,976
Lambda K H
0.313 0.130 0.407
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 13,892,790
Number of Sequences: 36976
Number of extensions: 269491
Number of successful extensions: 2257
Number of sequences better than 10.0: 188
Number of HSP's better than 10.0 without gapping: 1522
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 1970
length of query: 336
length of database: 9,014,727
effective HSP length: 97
effective length of query: 239
effective length of database: 5,428,055
effective search space: 1297305145
effective search space used: 1297305145
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.2 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 42 (21.9 bits)
S2: 59 (27.3 bits)
Medicago: description of AC140030.1


