

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC140025.10 - phase: 0
(344 letters)
Database: MTGI
36,976 sequences; 27,044,181 total letters
Searching..................................................done
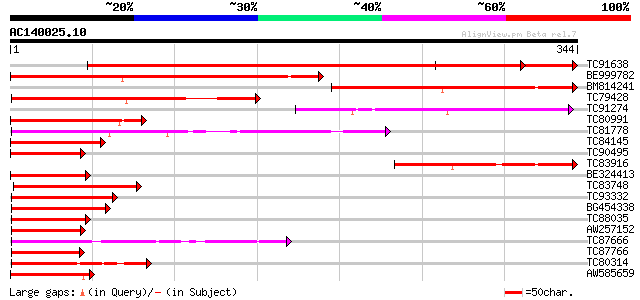
Score E
Sequences producing significant alignments: (bits) Value
TC91638 similar to GP|15983797|gb|AAL10495.1 AT5g39660/MIJ24_130... 566 e-162
BE999782 similar to PIR|T09661|T096 ascorbate oxidase promoter-b... 235 2e-62
BM814241 similar to GP|15983797|gb| AT5g39660/MIJ24_130 {Arabido... 182 2e-46
TC79428 similar to PIR|T09661|T09661 ascorbate oxidase promoter-... 147 8e-36
TC91274 similar to PIR|T45710|T45710 H-protein promoter binding ... 143 1e-34
TC80991 similar to GP|15983797|gb|AAL10495.1 AT5g39660/MIJ24_130... 122 2e-28
TC81778 similar to GP|7242908|dbj|BAA92506.1 ESTs C23582(S11122)... 118 3e-27
TC84145 similar to GP|7242908|dbj|BAA92506.1 ESTs C23582(S11122)... 102 3e-22
TC90495 similar to PIR|H84752|H84752 probable DOF zinc finger pr... 98 4e-21
TC83916 similar to PIR|T09661|T09661 ascorbate oxidase promoter-... 97 7e-21
BE324413 homologue to GP|10176912|dbj zinc finger protein {Arabi... 92 2e-19
TC83748 homologue to PIR|T02375|T02375 finger protein BBF3 - com... 92 2e-19
TC93332 similar to PIR|T48267|T48267 probable zinc finger protei... 92 4e-19
BG454338 homologue to PIR|T52044|T520 dof zinc finger protein [i... 92 4e-19
TC88035 similar to GP|6092016|dbj|BAA85655.1 elicitor-responsive... 89 2e-18
AW257152 similar to PIR|D86409|D864 hypothetical protein AAF9842... 88 4e-18
TC87666 homologue to GP|4996640|dbj|BAA78572.1 Dof zinc finger p... 87 1e-17
TC87766 similar to GP|10177472|dbj|BAB10863. contains similarity... 86 2e-17
TC80314 similar to PIR|D86409|D86409 hypothetical protein AAF984... 86 3e-17
AW585659 similar to GP|2262110|gb|A zinc finger protein isolog {... 86 3e-17
>TC91638 similar to GP|15983797|gb|AAL10495.1 AT5g39660/MIJ24_130
{Arabidopsis thaliana}, partial (16%)
Length = 914
Score = 566 bits (1458), Expect = e-162
Identities = 265/266 (99%), Positives = 265/266 (99%)
Frame = +2
Query: 48 SSSHYRQITVSEATLQNSRIHPSVKCNGTILTFGSNSPVCESMASVLKHADKTMQNYTRN 107
SSSHYRQITVSEATLQNSRIHPSVKCNGTILTFGSNSPVCESMASVLKHADKTMQNYTRN
Sbjct: 2 SSSHYRQITVSEATLQNSRIHPSVKCNGTILTFGSNSPVCESMASVLKHADKTMQNYTRN 181
Query: 108 GYHKHEELRICVPHTSEEQGEDQSNKSSVTSTKSTEGATTNVSQEQAMWNDHSFPPQGGY 167
GYHKHEELRICVPHTSEEQGEDQSNKSSVTSTKSTEGATTNVSQEQAMWNDHSFPPQGGY
Sbjct: 182 GYHKHEELRICVPHTSEEQGEDQSNKSSVTSTKSTEGATTNVSQEQAMWNDHSFPPQGGY 361
Query: 168 FPHGTPWHLPWNPVQMSSPIPPPAFCPPGFSMPFYPATTYWGCTMPSAWNIPRQAQPSSP 227
FPHGTPWHLPWNPVQMSSPIPPPAFCPPGFSMPFYPATTYWGCTMPSAWNIPRQAQPSSP
Sbjct: 362 FPHGTPWHLPWNPVQMSSPIPPPAFCPPGFSMPFYPATTYWGCTMPSAWNIPRQAQPSSP 541
Query: 228 NGANHDSTPNSPTLGKHSREDNMLKSSEGDGKKEISEEKSLWFPKTLRIDDSEEAEKSSF 287
NGANHDSTPNSPTLGKHSREDNMLKSSEGDGKKEISEEKSLWFPKTLRIDDSEEAEKSSF
Sbjct: 542 NGANHDSTPNSPTLGKHSREDNMLKSSEGDGKKEISEEKSLWFPKTLRIDDSEEAEKSSF 721
Query: 288 WTTLGIKNNNADSVPPRRLFQAFPSK 313
WTTLGIKNNNADSVPPRRLFQAFP K
Sbjct: 722 WTTLGIKNNNADSVPPRRLFQAFPFK 799
Score = 74.7 bits (182), Expect = 5e-14
Identities = 41/86 (47%), Positives = 55/86 (63%)
Frame = +3
Query: 259 KKEISEEKSLWFPKTLRIDDSEEAEKSSFWTTLGIKNNNADSVPPRRLFQAFPSKCDEKN 318
+K+ +++++ F K + + +K+ LG++ F+ PSKCDEKN
Sbjct: 636 RKK*AKKRACGFLKL*GLMTLRKQKKALSGQHLGLRTTTPTQFLREDSFRHSPSKCDEKN 815
Query: 319 HLVQVSSVLQANPAALSRSLHFHETS 344
HLVQVSSVLQANPAALSRSLHFHETS
Sbjct: 816 HLVQVSSVLQANPAALSRSLHFHETS 893
>BE999782 similar to PIR|T09661|T096 ascorbate oxidase promoter-binding
protein AOBP - winter squash, partial (17%)
Length = 630
Score = 235 bits (600), Expect = 2e-62
Identities = 116/197 (58%), Positives = 139/197 (69%), Gaps = 7/197 (3%)
Frame = +2
Query: 1 MDTKFCYYNNYNVNQPRHFCKNCQRYWTAGGTMRNVPVGAGRRKNKSSSSHYRQITVSEA 60
MDTKFCYYNNYNVNQPRHFCK CQRYWTAGG MRNVPVGAGRRKNK+S+SH+RQITV E
Sbjct: 38 MDTKFCYYNNYNVNQPRHFCKKCQRYWTAGGAMRNVPVGAGRRKNKNSASHFRQITVPET 217
Query: 61 TLQNSRI-------HPSVKCNGTILTFGSNSPVCESMASVLKHADKTMQNYTRNGYHKHE 113
+QNS HPS+ CNGT+ TF +++P+CESM S L AD+ + +NG+ + E
Sbjct: 218 AVQNSLSDSPNGVHHPSLNCNGTVFTFRTDTPLCESMESALNLADQGVNISQKNGFIRPE 397
Query: 114 ELRICVPHTSEEQGEDQSNKSSVTSTKSTEGATTNVSQEQAMWNDHSFPPQGGYFPHGTP 173
LRI VP+ EE+ ++ S KSS TST TE A + S EQ M N SF PQ Y+P P
Sbjct: 398 ALRIHVPYVGEEKSDEHSIKSSDTSTTLTEDAAASSSVEQVMPNCQSFQPQVPYYP-SAP 574
Query: 174 WHLPWNPVQMSSPIPPP 190
W LPW+P Q SS + PP
Sbjct: 575 WLLPWSPSQWSSQVQPP 625
>BM814241 similar to GP|15983797|gb| AT5g39660/MIJ24_130 {Arabidopsis
thaliana}, partial (10%)
Length = 527
Score = 182 bits (461), Expect = 2e-46
Identities = 95/151 (62%), Positives = 111/151 (72%), Gaps = 2/151 (1%)
Frame = +3
Query: 196 GFSMPFYPATTYWGCTMPSAWNIPRQAQPSSPNGANHDSTPNSPTLGKHSREDNMLKSSE 255
GF+MP YP YWG +MP AWN P AQPSSPN A +S PNSPTLGKHSRE++MLK ++
Sbjct: 3 GFAMPLYPPPAYWGFSMPGAWNNPWLAQPSSPNSATVNSGPNSPTLGKHSREESMLKPTD 182
Query: 256 GDGKKE--ISEEKSLWFPKTLRIDDSEEAEKSSFWTTLGIKNNNADSVPPRRLFQAFPSK 313
G E EEKSLW PKTLRIDD EAEKSS TTLGIKN+ AD++ LF+AF SK
Sbjct: 183 STGSDEGNNKEEKSLWVPKTLRIDDLGEAEKSSILTTLGIKNDKADAIRGGGLFKAFASK 362
Query: 314 CDEKNHLVQVSSVLQANPAALSRSLHFHETS 344
+EK+ VQ S +QANPAA+SRS+ FHETS
Sbjct: 363 SNEKDS-VQNSPAMQANPAAMSRSISFHETS 452
>TC79428 similar to PIR|T09661|T09661 ascorbate oxidase promoter-binding
protein AOBP - winter squash, partial (27%)
Length = 997
Score = 147 bits (370), Expect = 8e-36
Identities = 84/160 (52%), Positives = 103/160 (63%), Gaps = 9/160 (5%)
Frame = +1
Query: 2 DTKFCYYNNYNVNQPRHFCKNCQRYWTAGGTMRNVPVGAGRRKNK-SSSSHYRQITVSEA 60
DTKFCYYNNYNVNQPR+FCK CQRYWTAGGTMRNVPVGAGRRKNK +SSSHYR IT+SEA
Sbjct: 550 DTKFCYYNNYNVNQPRYFCKACQRYWTAGGTMRNVPVGAGRRKNKNNSSSHYRHITISEA 729
Query: 61 TLQNSRIHP-------SVKCNGTILTFGSNSP-VCESMASVLKHADKTMQNYTRNGYHKH 112
I P ++K NG +L FG + P + +SM++ L A+K + N TRN
Sbjct: 730 LDAARIISPNGTHHLQNLKTNGRVLNFGLDHPHIYDSMSNDLNPAEKKVLNDTRN----- 894
Query: 113 EELRICVPHTSEEQGEDQSNKSSVTSTKSTEGATTNVSQE 152
G+ S+ SSVT +KS E + N++QE
Sbjct: 895 -------------NGDRFSSASSVTVSKSMEESGKNMTQE 975
>TC91274 similar to PIR|T45710|T45710 H-protein promoter binding factor-2a
[imported] - Arabidopsis thaliana, partial (9%)
Length = 959
Score = 143 bits (360), Expect = 1e-34
Identities = 81/177 (45%), Positives = 104/177 (57%), Gaps = 8/177 (4%)
Frame = +2
Query: 174 WHLPWNPVQMSSPIPPPAFCPPGFSMPFYPATT---YWGCTMPSAWNIPRQAQPSSPNGA 230
+ +P+ P +S +PPP FCPP + + FY T YWGC MP WNIP P S +
Sbjct: 131 FQVPFLPYTWNSAMPPPTFCPPNYPLAFYTPVTPPAYWGC-MPPPWNIPC-ISPGSASVN 304
Query: 231 NHDSTPNS-PTLGKHSREDNMLKSSEGDGKKEISE----EKSLWFPKTLRIDDSEEAEKS 285
DS +S PTLGKHSR+ N++ S +K +E E S+ PKTLR DD EA KS
Sbjct: 305 ESDSAHSSVPTLGKHSRDGNIITSVNSPKEKPETEHNSTESSVLIPKTLRFDDPSEAAKS 484
Query: 286 SFWTTLGIKNNNADSVPPRRLFQAFPSKCDEKNHLVQVSSVLQANPAALSRSLHFHE 342
S W+ LGI N+ A+S+ +F F SK + NH V S ++ ANPAALSRS FHE
Sbjct: 485 SLWSKLGINNDKANSLNGGGMFNGFQSKGKDMNHSVGTSPLMYANPAALSRSRTFHE 655
>TC80991 similar to GP|15983797|gb|AAL10495.1 AT5g39660/MIJ24_130
{Arabidopsis thaliana}, partial (19%)
Length = 782
Score = 122 bits (306), Expect = 2e-28
Identities = 60/86 (69%), Positives = 67/86 (77%), Gaps = 3/86 (3%)
Frame = +3
Query: 1 MDTKFCYYNNYNVNQPRHFCKNCQRYWTAGGTMRNVPVGAGRRKNKS-SSSHYRQITVSE 59
MDTKFCY+NNYNVNQPRHFCK CQRYWTAGGTMRNVPVGAGRRKNK+ S+SHYR + + E
Sbjct: 522 MDTKFCYFNNYNVNQPRHFCKKCQRYWTAGGTMRNVPVGAGRRKNKNLSTSHYRHLIIPE 701
Query: 60 ATLQNS--RIHPSVKCNGTILTFGSN 83
NS IH S+ +LTF SN
Sbjct: 702 GAKLNSPNGIH-SLGNGAAVLTFDSN 776
>TC81778 similar to GP|7242908|dbj|BAA92506.1 ESTs C23582(S11122)
AU056531(S20663) correspond to a region of the predicted
gene.~Similar to AOBP, partial (17%)
Length = 1081
Score = 118 bits (296), Expect = 3e-27
Identities = 87/256 (33%), Positives = 114/256 (43%), Gaps = 26/256 (10%)
Frame = +3
Query: 2 DTKFCYYNNYNVNQPRHFCKNCQRYWTAGGTMRNVPVGAGRRKNKSSSSHYRQITVSE-- 59
DTKFCY+NNYNVNQPRHFC++C RYWT GGTMRNV VGAGRRKNK +S YR + V+
Sbjct: 330 DTKFCYFNNYNVNQPRHFCRSCHRYWTNGGTMRNVAVGAGRRKNKHIASQYRHMIVASDG 509
Query: 60 -----------------ATLQNSRIHPSVKCNGTILTFG-SNSPVCESMASVL------K 95
+ L+++ + NG +L FG N+ + ES S+L +
Sbjct: 510 IPTASLETNDSSRYQHGSNLESAAVFRCSNDNGIVLKFGRENASLDESKGSMLNLMNNRR 689
Query: 96 HADKTMQNYTRNGYHKHEELRICVPHTSEEQGEDQSNKSSVTSTKSTEGATTNVSQEQAM 155
H D + N NG EE +CV SSVT+ T G S++
Sbjct: 690 HVDASGNNCRENG---EEETSLCV--------------SSVTN-GHTRGNELFESEQNRS 815
Query: 156 WNDHSFPPQGGYFPHGTPWHLPWNPVQMSSPIPPPAFCPPGFSMPFYPATTYWGCTMPSA 215
S+P P W+ + VQ S + P P A + M +
Sbjct: 816 KPMQSYPASSWIIPMNQRWNNVTSMVQSSMQMCNPYGIDP-------TAMQWCHAPMVAV 974
Query: 216 WNIPRQAQPSSPNGAN 231
NI Q P S N
Sbjct: 975 TNIGLQFGPGSNRNGN 1022
>TC84145 similar to GP|7242908|dbj|BAA92506.1 ESTs C23582(S11122)
AU056531(S20663) correspond to a region of the predicted
gene.~Similar to AOBP, partial (16%)
Length = 673
Score = 102 bits (253), Expect = 3e-22
Identities = 41/58 (70%), Positives = 50/58 (85%)
Frame = +3
Query: 1 MDTKFCYYNNYNVNQPRHFCKNCQRYWTAGGTMRNVPVGAGRRKNKSSSSHYRQITVS 58
++TKFCY+NNYNVNQPRHFCKNCQRYWTAGG +RNVP+GAG+R+NK S Q+ V+
Sbjct: 267 LETKFCYFNNYNVNQPRHFCKNCQRYWTAGGVIRNVPIGAGKRRNKQSPLQNCQVPVT 440
>TC90495 similar to PIR|H84752|H84752 probable DOF zinc finger protein
[imported] - Arabidopsis thaliana, partial (41%)
Length = 620
Score = 98.2 bits (243), Expect = 4e-21
Identities = 40/46 (86%), Positives = 44/46 (94%)
Frame = +3
Query: 1 MDTKFCYYNNYNVNQPRHFCKNCQRYWTAGGTMRNVPVGAGRRKNK 46
M+TKFCY+NNYNVNQPRHFCK+CQRYWTAGG +RNVPVGAGRRK K
Sbjct: 276 METKFCYFNNYNVNQPRHFCKSCQRYWTAGGALRNVPVGAGRRKAK 413
>TC83916 similar to PIR|T09661|T09661 ascorbate oxidase promoter-binding
protein AOBP - winter squash, partial (12%)
Length = 703
Score = 97.4 bits (241), Expect = 7e-21
Identities = 59/117 (50%), Positives = 71/117 (60%), Gaps = 6/117 (5%)
Frame = +3
Query: 234 STPNSPTLGKHSREDNMLKSSEGDGKKEISEEKS------LWFPKTLRIDDSEEAEKSSF 287
S+P SPTLGKHSR+D+ +EE + PKTLRIDD EA KSS
Sbjct: 48 SSPKSPTLGKHSRDDDNTNDENAKQDSLQTEESPKQRNGCVLVPKTLRIDDPTEAAKSSI 227
Query: 288 WTTLGIKNNNADSVPPRRLFQAFPSKCDEKNHLVQVSSVLQANPAALSRSLHFHETS 344
W TLGIKN + + + +AF SK D KNH VQ S +L ANPAAL+RSL+FHE S
Sbjct: 228 WETLGIKN---EGLSRGGMTKAFQSKKDGKNH-VQTSPMLMANPAALARSLNFHENS 386
>BE324413 homologue to GP|10176912|dbj zinc finger protein {Arabidopsis
thaliana}, partial (21%)
Length = 339
Score = 92.4 bits (228), Expect = 2e-19
Identities = 37/49 (75%), Positives = 44/49 (89%)
Frame = +2
Query: 1 MDTKFCYYNNYNVNQPRHFCKNCQRYWTAGGTMRNVPVGAGRRKNKSSS 49
++TKFCYYNNYN++QPRHFCKNC+RYWT GG +RNVPVG G RK+K SS
Sbjct: 47 INTKFCYYNNYNLSQPRHFCKNCRRYWTKGGVLRNVPVGGGCRKSKRSS 193
>TC83748 homologue to PIR|T02375|T02375 finger protein BBF3 - common tobacco
(fragment), partial (60%)
Length = 658
Score = 92.4 bits (228), Expect = 2e-19
Identities = 43/79 (54%), Positives = 53/79 (66%), Gaps = 1/79 (1%)
Frame = +3
Query: 3 TKFCYYNNYNVNQPRHFCKNCQRYWTAGGTMRNVPVGAGRRKNKSSSSHYRQITVSEATL 62
TKFCYYNNY+++QPR+FCK C+RYWT GGT+RN+PVG G RKNK SS ++ +
Sbjct: 405 TKFCYYNNYSLSQPRYFCKTCRRYWTKGGTLRNIPVGGGCRKNKKVSSKKSNDHLANTNI 584
Query: 63 QNSRIH-PSVKCNGTILTF 80
QN H PS N L F
Sbjct: 585 QNQPHHVPSYHQNPKDLHF 641
>TC93332 similar to PIR|T48267|T48267 probable zinc finger protein -
Arabidopsis thaliana, partial (20%)
Length = 666
Score = 91.7 bits (226), Expect = 4e-19
Identities = 37/64 (57%), Positives = 48/64 (74%)
Frame = +1
Query: 2 DTKFCYYNNYNVNQPRHFCKNCQRYWTAGGTMRNVPVGAGRRKNKSSSSHYRQITVSEAT 61
+TKFCYYNNYN++QPRHFCK C+RYWT GG +RNVPVG G R++K + T S++
Sbjct: 154 NTKFCYYNNYNLSQPRHFCKTCRRYWTRGGALRNVPVGGGCRRSKKTKKSNTSSTPSKSP 333
Query: 62 LQNS 65
+ NS
Sbjct: 334 ISNS 345
>BG454338 homologue to PIR|T52044|T520 dof zinc finger protein [imported] -
Arabidopsis thaliana, partial (37%)
Length = 678
Score = 91.7 bits (226), Expect = 4e-19
Identities = 39/61 (63%), Positives = 48/61 (77%), Gaps = 1/61 (1%)
Frame = +1
Query: 2 DTKFCYYNNYNVNQPRHFCKNCQRYWTAGGTMRNVPVGAGRRK-NKSSSSHYRQITVSEA 60
+TKFCYYNNYN++QPRHFCKNC+RYWT GG +RN+PVG G RK K SS+ R T S +
Sbjct: 238 NTKFCYYNNYNLSQPRHFCKNCKRYWTKGGALRNIPVGGGTRKVTKRSSNSKRSTTPSSS 417
Query: 61 T 61
+
Sbjct: 418 S 420
>TC88035 similar to GP|6092016|dbj|BAA85655.1 elicitor-responsive Dof
protein ERDP {Pisum sativum}, partial (77%)
Length = 1397
Score = 89.4 bits (220), Expect = 2e-18
Identities = 36/48 (75%), Positives = 43/48 (89%)
Frame = +3
Query: 2 DTKFCYYNNYNVNQPRHFCKNCQRYWTAGGTMRNVPVGAGRRKNKSSS 49
+TKFCYYNNY++ QPR+FCK+C+RYWT GGT+RNVPVG G RKNK SS
Sbjct: 264 NTKFCYYNNYSLTQPRYFCKSCRRYWTKGGTLRNVPVGGGCRKNKRSS 407
>AW257152 similar to PIR|D86409|D864 hypothetical protein AAF98424.1
[imported] - Arabidopsis thaliana, partial (20%)
Length = 587
Score = 88.2 bits (217), Expect = 4e-18
Identities = 35/45 (77%), Positives = 41/45 (90%)
Frame = +1
Query: 2 DTKFCYYNNYNVNQPRHFCKNCQRYWTAGGTMRNVPVGAGRRKNK 46
+TKFCYYNNY+++QPRHFCK C+RYWT GGT+RNVPVG G RKNK
Sbjct: 148 NTKFCYYNNYSLSQPRHFCKACKRYWTRGGTLRNVPVGGGCRKNK 282
>TC87666 homologue to GP|4996640|dbj|BAA78572.1 Dof zinc finger protein
{Oryza sativa}, partial (20%)
Length = 1828
Score = 86.7 bits (213), Expect = 1e-17
Identities = 55/170 (32%), Positives = 83/170 (48%)
Frame = +2
Query: 2 DTKFCYYNNYNVNQPRHFCKNCQRYWTAGGTMRNVPVGAGRRKNKSSSSHYRQITVSEAT 61
+TKFCYYNNY++ QPR+FCK C+RYWT GG++RNVPVG G RKNK S +++
Sbjct: 731 NTKFCYYNNYSLTQPRYFCKTCRRYWTEGGSLRNVPVGGGSRKNKKIPS-----SLANPN 895
Query: 62 LQNSRIHPSVKCNGTILTFGSNSPVCESMASVLKHADKTMQNYTRNGYHKHEELRICVPH 121
+S+I + S +P + L A +M+NY +H H +P
Sbjct: 896 NSSSKIPDLNPPTLQVSHLSSQNPKIQG-GQDLNLAFPSMENY----HHNHG-----MPS 1045
Query: 122 TSEEQGEDQSNKSSVTSTKSTEGATTNVSQEQAMWNDHSFPPQGGYFPHG 171
+ E + + SS + A+ ++ A N+ P +P G
Sbjct: 1046SYVEMHNNNESSSSALDLLRSSMASRGMN-PYANNNNLGLPNSNAIYPSG 1192
>TC87766 similar to GP|10177472|dbj|BAB10863. contains similarity to dof
zinc finger protein~gene_id:MQB2.26 {Arabidopsis
thaliana}, partial (21%)
Length = 1516
Score = 85.9 bits (211), Expect = 2e-17
Identities = 33/44 (75%), Positives = 39/44 (88%)
Frame = +2
Query: 2 DTKFCYYNNYNVNQPRHFCKNCQRYWTAGGTMRNVPVGAGRRKN 45
+TKFCYYNNYN++QPRHFCK C+RYWT GG +RNVPVG G R+N
Sbjct: 353 NTKFCYYNNYNLSQPRHFCKTCRRYWTKGGALRNVPVGGGCRRN 484
>TC80314 similar to PIR|D86409|D86409 hypothetical protein AAF98424.1
[imported] - Arabidopsis thaliana, partial (19%)
Length = 986
Score = 85.5 bits (210), Expect = 3e-17
Identities = 42/85 (49%), Positives = 53/85 (61%)
Frame = +1
Query: 2 DTKFCYYNNYNVNQPRHFCKNCQRYWTAGGTMRNVPVGAGRRKNKSSSSHYRQITVSEAT 61
+TKFCYYNNYN +QPRHFC+ C+R+WT GGT+RNVPVG G RKNK +T S T
Sbjct: 475 NTKFCYYNNYNKSQPRHFCRACKRHWTKGGTLRNVPVGGG-RKNKRIKKPTTPVT-SSTT 648
Query: 62 LQNSRIHPSVKCNGTILTFGSNSPV 86
+ + S C +I SN +
Sbjct: 649 ITTT----STTCTTSITNMNSNMAI 711
>AW585659 similar to GP|2262110|gb|A zinc finger protein isolog {Arabidopsis
thaliana}, partial (27%)
Length = 483
Score = 85.5 bits (210), Expect = 3e-17
Identities = 35/54 (64%), Positives = 46/54 (84%), Gaps = 3/54 (5%)
Frame = +2
Query: 1 MDTKFCYYNNYNVNQPRHFCKNCQRYWTAGGTMRNVPVGAGRR---KNKSSSSH 51
++TKFCYYNNY++ QPR+FCK C+RYWT GG++RN+PVG G R KN+SSSS+
Sbjct: 227 INTKFCYYNNYSLTQPRYFCKTCRRYWTQGGSIRNIPVGGGSRKNNKNRSSSSY 388
Database: MTGI
Posted date: Oct 22, 2004 3:39 PM
Number of letters in database: 27,044,181
Number of sequences in database: 36,976
Lambda K H
0.312 0.127 0.401
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 14,071,046
Number of Sequences: 36976
Number of extensions: 245644
Number of successful extensions: 1638
Number of sequences better than 10.0: 107
Number of HSP's better than 10.0 without gapping: 1486
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 1605
length of query: 344
length of database: 9,014,727
effective HSP length: 97
effective length of query: 247
effective length of database: 5,428,055
effective search space: 1340729585
effective search space used: 1340729585
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.2 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 42 (21.9 bits)
S2: 59 (27.3 bits)
Medicago: description of AC140025.10


