

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC139743.4 - phase: 0 /pseudo
(105 letters)
Database: MTGI
36,976 sequences; 27,044,181 total letters
Searching..................................................done
Score E
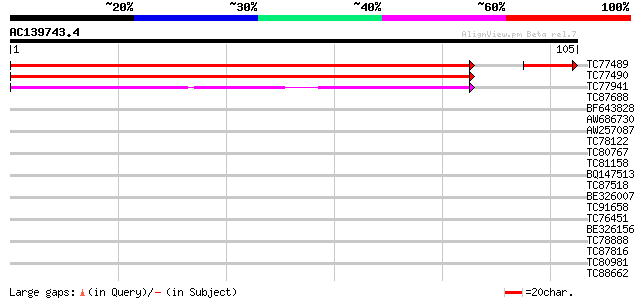
Sequences producing significant alignments: (bits) Value
TC77489 MTD1 174 2e-46
TC77490 GP|9294810|gb|AAF86687.1| MTD1 {Medicago truncatula}, pa... 174 6e-45
TC77941 weakly similar to GP|9294810|gb|AAF86687.1| MTD1 {Medica... 45 4e-06
TC87688 similar to GP|10177535|dbj|BAB10930. gene_id:K1F13.21~un... 34 0.008
BF643828 similar to GP|15810439|gb| unknown protein {Arabidopsis... 33 0.017
AW686730 similar to PIR|T10670|T10 hypothetical protein F6E21.80... 33 0.023
AW257087 homologue to GP|6683126|dbj KIAA0339 protein {Homo sapi... 32 0.029
TC78122 similar to GP|21593751|gb|AAM65718.1 unknown {Arabidopsi... 32 0.038
TC80767 weakly similar to GP|21593751|gb|AAM65718.1 unknown {Ara... 32 0.038
TC81158 similar to GP|9280664|gb|AAF86533.1| F21B7.15 {Arabidops... 32 0.038
BQ147513 similar to SP|Q09475|YP93 Putative helicase C28H8.3 (EC... 32 0.038
TC87518 weakly similar to PIR|E96714|E96714 probable DNA-binding... 31 0.086
BE326007 similar to GP|13676415|d hypothetical protein {Glycine ... 30 0.15
TC91658 similar to GP|14329812|emb|CAC40753. putative nucleosome... 29 0.25
TC76451 weakly similar to GP|10177810|dbj|BAB11176. serine carbo... 29 0.33
BE326156 similar to GP|9759609|dbj gene_id:MNA5.1~unknown protei... 28 0.56
TC78888 weakly similar to PIR|T43427|T43427 pob1 protein - fissi... 28 0.56
TC87816 similar to GP|23496590|gb|AAN36147.1 hypothetical protei... 28 0.56
TC80981 similar to GP|8698730|gb|AAF78488.1| ESTs gb|Z18526 gb|... 28 0.73
TC88662 similar to GP|17979181|gb|AAL49786.1 unknown protein {Ar... 28 0.73
>TC77489 MTD1
Length = 988
Score = 174 bits (440), Expect(2) = 2e-46
Identities = 86/86 (100%), Positives = 86/86 (100%)
Frame = +2
Query: 1 MELETVAFKRVGSRTSVLFDAPRDHDDDQGKVCSTTSSSSIGRNSDDDDDDEVSSERSMD 60
MELETVAFKRVGSRTSVLFDAPRDHDDDQGKVCSTTSSSSIGRNSDDDDDDEVSSERSMD
Sbjct: 89 MELETVAFKRVGSRTSVLFDAPRDHDDDQGKVCSTTSSSSIGRNSDDDDDDEVSSERSMD 268
Query: 61 ENEAESKYNGGALDCMEALEEVLPIR 86
ENEAESKYNGGALDCMEALEEVLPIR
Sbjct: 269 ENEAESKYNGGALDCMEALEEVLPIR 346
Score = 26.6 bits (57), Expect(2) = 2e-46
Identities = 10/10 (100%), Positives = 10/10 (100%)
Frame = +1
Query: 96 EKYLKFLQWE 105
EKYLKFLQWE
Sbjct: 346 EKYLKFLQWE 375
>TC77490 GP|9294810|gb|AAF86687.1| MTD1 {Medicago truncatula}, partial
(37%)
Length = 651
Score = 174 bits (440), Expect = 6e-45
Identities = 86/86 (100%), Positives = 86/86 (100%)
Frame = +2
Query: 1 MELETVAFKRVGSRTSVLFDAPRDHDDDQGKVCSTTSSSSIGRNSDDDDDDEVSSERSMD 60
MELETVAFKRVGSRTSVLFDAPRDHDDDQGKVCSTTSSSSIGRNSDDDDDDEVSSERSMD
Sbjct: 8 MELETVAFKRVGSRTSVLFDAPRDHDDDQGKVCSTTSSSSIGRNSDDDDDDEVSSERSMD 187
Query: 61 ENEAESKYNGGALDCMEALEEVLPIR 86
ENEAESKYNGGALDCMEALEEVLPIR
Sbjct: 188ENEAESKYNGGALDCMEALEEVLPIR 265
>TC77941 weakly similar to GP|9294810|gb|AAF86687.1| MTD1 {Medicago
truncatula}, partial (15%)
Length = 1059
Score = 45.1 bits (105), Expect = 4e-06
Identities = 30/86 (34%), Positives = 42/86 (47%)
Frame = +3
Query: 1 MELETVAFKRVGSRTSVLFDAPRDHDDDQGKVCSTTSSSSIGRNSDDDDDDEVSSERSMD 60
+ LET+ ++ + D DD C +SSSS+GRNSD +DD S D
Sbjct: 141 LSLETMERSSSFNQYDSICDQDFPEDDSDTDSC-VSSSSSLGRNSDSSEDD------SSD 299
Query: 61 ENEAESKYNGGALDCMEALEEVLPIR 86
E E G LD M+ LE+ LP++
Sbjct: 300 REEVEKNSFKGPLDTMKDLEKDLPVK 377
>TC87688 similar to GP|10177535|dbj|BAB10930. gene_id:K1F13.21~unknown
protein {Arabidopsis thaliana}, partial (46%)
Length = 2077
Score = 34.3 bits (77), Expect = 0.008
Identities = 19/60 (31%), Positives = 26/60 (42%), Gaps = 4/60 (6%)
Frame = +2
Query: 19 FDAPRDHDDDQGKVCSTTSSSSIGRNSDD----DDDDEVSSERSMDENEAESKYNGGALD 74
FD D DDD + + DD DD+DE SE DE + + K GG ++
Sbjct: 395 FDEELDDDDDDFEGVEKKKAKGGSEGEDDFEEEDDEDEEGSEDEDDEEDEKEKVKGGGIE 574
>BF643828 similar to GP|15810439|gb| unknown protein {Arabidopsis
thaliana}, partial (8%)
Length = 386
Score = 33.1 bits (74), Expect = 0.017
Identities = 17/42 (40%), Positives = 22/42 (51%)
Frame = +2
Query: 17 VLFDAPRDHDDDQGKVCSTTSSSSIGRNSDDDDDDEVSSERS 58
V+ +APR+ D G S++SSSS S D D SS S
Sbjct: 5 VIVEAPRERQADNGSKTSSSSSSSSDSGSSSSDSDSDSSSAS 130
>AW686730 similar to PIR|T10670|T10 hypothetical protein F6E21.80 -
Arabidopsis thaliana, partial (4%)
Length = 376
Score = 32.7 bits (73), Expect = 0.023
Identities = 16/43 (37%), Positives = 22/43 (50%)
Frame = +2
Query: 27 DDQGKVCSTTSSSSIGRNSDDDDDDEVSSERSMDENEAESKYN 69
DDQG + S+ S IGR DDD + S +E+E + N
Sbjct: 194 DDQGDMYSSARSYEIGRRRPTDDDSDPDDAESEEEDEDDDDDN 322
>AW257087 homologue to GP|6683126|dbj KIAA0339 protein {Homo sapiens},
partial (2%)
Length = 466
Score = 32.3 bits (72), Expect = 0.029
Identities = 18/59 (30%), Positives = 30/59 (50%)
Frame = -3
Query: 13 SRTSVLFDAPRDHDDDQGKVCSTTSSSSIGRNSDDDDDDEVSSERSMDENEAESKYNGG 71
S +S+L D D D+D+ + + D+D+DDE E S E+E++S + G
Sbjct: 398 SSSSLLDDEDEDEDEDEDE------------DEDEDEDDEELLEESESESESDSSFLAG 258
Score = 28.9 bits (63), Expect = 0.33
Identities = 12/36 (33%), Positives = 23/36 (63%)
Frame = -3
Query: 30 GKVCSTTSSSSIGRNSDDDDDDEVSSERSMDENEAE 65
G S++SSSS+ + D+D+D++ + DE++ E
Sbjct: 419 GFASSSSSSSSLLDDEDEDEDEDEDEDEDEDEDDEE 312
Score = 25.4 bits (54), Expect = 3.6
Identities = 10/34 (29%), Positives = 21/34 (61%)
Frame = -3
Query: 34 STTSSSSIGRNSDDDDDDEVSSERSMDENEAESK 67
+++SSSS D+D+D++ + DE+E + +
Sbjct: 413 ASSSSSSSSLLDDEDEDEDEDEDEDEDEDEDDEE 312
Score = 25.4 bits (54), Expect = 3.6
Identities = 10/19 (52%), Positives = 16/19 (83%)
Frame = +1
Query: 34 STTSSSSIGRNSDDDDDDE 52
S++SSSS NSDD++++E
Sbjct: 352 SSSSSSSSSSNSDDEEEEE 408
>TC78122 similar to GP|21593751|gb|AAM65718.1 unknown {Arabidopsis
thaliana}, partial (28%)
Length = 1394
Score = 32.0 bits (71), Expect = 0.038
Identities = 20/50 (40%), Positives = 30/50 (60%)
Frame = +3
Query: 37 SSSSIGRNSDDDDDDEVSSERSMDENEAESKYNGGALDCMEALEEVLPIR 86
SSSSIG D D+D+EV S+ ++ NG L +++LE+ LPI+
Sbjct: 375 SSSSIGTPDDSDNDEEVQSKLNLKGR------NG--LGSLDSLEDSLPIK 500
>TC80767 weakly similar to GP|21593751|gb|AAM65718.1 unknown {Arabidopsis
thaliana}, partial (35%)
Length = 750
Score = 32.0 bits (71), Expect = 0.038
Identities = 18/44 (40%), Positives = 27/44 (60%)
Frame = +3
Query: 43 RNSDDDDDDEVSSERSMDENEAESKYNGGALDCMEALEEVLPIR 86
RN D+DDDDEV +SK+ G L+ +++LE+ LPI+
Sbjct: 45 RNRDEDDDDEV-----------QSKFKG--LNSLDSLEDSLPIK 137
>TC81158 similar to GP|9280664|gb|AAF86533.1| F21B7.15 {Arabidopsis
thaliana}, partial (6%)
Length = 890
Score = 32.0 bits (71), Expect = 0.038
Identities = 15/41 (36%), Positives = 23/41 (55%)
Frame = +1
Query: 34 STTSSSSIGRNSDDDDDDEVSSERSMDENEAESKYNGGALD 74
ST+SSS+ G +SD+DD+D+ E ++ E A D
Sbjct: 469 STSSSSTSGSDSDEDDEDDAEDEEEGQISDDEKMVTWSAAD 591
>BQ147513 similar to SP|Q09475|YP93 Putative helicase C28H8.3 (EC 3.6.1.-).
{Caenorhabditis elegans}, partial (0%)
Length = 201
Score = 32.0 bits (71), Expect = 0.038
Identities = 20/56 (35%), Positives = 31/56 (54%)
Frame = +3
Query: 46 DDDDDDEVSSERSMDENEAESKYNGGALDCMEALEEVLPIRLFQSFMVMQEKYLKF 101
D+DDDDEV SK+ G L+ +++LE+ LPI+L SF + ++ F
Sbjct: 60 DEDDDDEVX-----------SKFKG--LNSLDSLEDSLPIKLGSSFS*FLD*FVNF 188
>TC87518 weakly similar to PIR|E96714|E96714 probable DNA-binding protein
T6L1.19 [imported] - Arabidopsis thaliana, partial (41%)
Length = 1337
Score = 30.8 bits (68), Expect = 0.086
Identities = 22/86 (25%), Positives = 42/86 (48%), Gaps = 8/86 (9%)
Frame = +2
Query: 26 DDDQGKVCSTTSSSSIGRNSDDDDDDEVSSERS----MDENEAESKYNGGALDCMEALEE 81
DDD + + +SSSI N++ D+ + +R+ +E E + + +AL +
Sbjct: 92 DDDDQDIFTPNTSSSINNNNNVKVDEPIRGKRANPHRSKHSETEQRRRSKINERFQALRD 271
Query: 82 VLP----IRLFQSFMVMQEKYLKFLQ 103
++P R SF++ +Y+ FLQ
Sbjct: 272 LIPENDSKRDKASFLLEVIEYIHFLQ 349
>BE326007 similar to GP|13676415|d hypothetical protein {Glycine max},
partial (8%)
Length = 544
Score = 30.0 bits (66), Expect = 0.15
Identities = 20/68 (29%), Positives = 35/68 (51%), Gaps = 1/68 (1%)
Frame = +1
Query: 19 FDAPRDHDDDQGKVCSTTSS-SSIGRNSDDDDDDEVSSERSMDENEAESKYNGGALDCME 77
FD+ ++ D D CS +SS + N+D+DD ++ S+R+ S + D M
Sbjct: 106 FDSFKEGDAD----CSPSSSWKRVDNNTDEDDSEDSGSDRA-----ESSSPDASMADIMP 258
Query: 78 ALEEVLPI 85
L+E+ P+
Sbjct: 259 MLDELHPL 282
>TC91658 similar to GP|14329812|emb|CAC40753. putative nucleosome assembly
protein 1 {Atropa belladonna}, partial (46%)
Length = 583
Score = 29.3 bits (64), Expect = 0.25
Identities = 16/53 (30%), Positives = 23/53 (43%)
Frame = +2
Query: 20 DAPRDHDDDQGKVCSTTSSSSIGRNSDDDDDDEVSSERSMDENEAESKYNGGA 72
D D DD+ G + DDDDD+E + DE + +SK G+
Sbjct: 74 DIEVDEDDEDG------DEEDDDDDDDDDDDEEDEDDEEEDEGKGKSKSKRGS 214
>TC76451 weakly similar to GP|10177810|dbj|BAB11176. serine carboxypeptidase
II-like protein {Arabidopsis thaliana}, partial (6%)
Length = 774
Score = 28.9 bits (63), Expect = 0.33
Identities = 17/57 (29%), Positives = 29/57 (50%), Gaps = 8/57 (14%)
Frame = -3
Query: 24 DHDDDQGKVCSTTSS--SSIGRNSDDDDDDEVSSERSMDENE------AESKYNGGA 72
DHD + + S S+ S+ +NSDD+D + + ++NE + + NGGA
Sbjct: 256 DHDGEGSDIDSPISNVAGSLQQNSDDNDSGLIEGSETEEDNEFSHSDQEDIRTNGGA 86
>BE326156 similar to GP|9759609|dbj gene_id:MNA5.1~unknown protein
{Arabidopsis thaliana}, partial (22%)
Length = 522
Score = 28.1 bits (61), Expect = 0.56
Identities = 21/90 (23%), Positives = 37/90 (40%), Gaps = 14/90 (15%)
Frame = -2
Query: 10 RVGSRTSVLFDAPRDHDDDQGKVCS---------TTSSSSIGRN-----SDDDDDDEVSS 55
R TS+ + HD + C+ +S +GR S D+DDD +
Sbjct: 320 RYSKVTSIFKTLNKGHDSPSKQKCTPI*PAPEERNSSIVQLGRYCHPDISGDEDDDGEGN 141
Query: 56 ERSMDENEAESKYNGGALDCMEALEEVLPI 85
E S+ E ++ G + + +A+ +PI
Sbjct: 140 EPSVSNGEENVEWYGRSSEVPKAVNRTIPI 51
>TC78888 weakly similar to PIR|T43427|T43427 pob1 protein - fission yeast
(Schizosaccharomyces pombe), partial (4%)
Length = 956
Score = 28.1 bits (61), Expect = 0.56
Identities = 17/71 (23%), Positives = 33/71 (45%), Gaps = 7/71 (9%)
Frame = +3
Query: 23 RDHDDDQ-------GKVCSTTSSSSIGRNSDDDDDDEVSSERSMDENEAESKYNGGALDC 75
+D DDD G + + +S +S + DDD+ +S + + + S ++ G L
Sbjct: 216 QDVDDDDVTDSISNGSITEDSMNSMCSSSSSELDDDQEASSSTSSLSSSSSSHSNGPLYE 395
Query: 76 MEALEEVLPIR 86
+ L LP++
Sbjct: 396 LSELMNHLPLK 428
>TC87816 similar to GP|23496590|gb|AAN36147.1 hypothetical protein
{Plasmodium falciparum 3D7}, partial (0%)
Length = 960
Score = 28.1 bits (61), Expect = 0.56
Identities = 11/20 (55%), Positives = 12/20 (60%)
Frame = -2
Query: 33 CSTTSSSSIGRNSDDDDDDE 52
C S + G N DDDDDDE
Sbjct: 218 CQNISLADDGNNDDDDDDDE 159
>TC80981 similar to GP|8698730|gb|AAF78488.1| ESTs gb|Z18526 gb|AI994480
gb|T44186 gb|Z30806 come from this gene. {Arabidopsis
thaliana}, partial (37%)
Length = 699
Score = 27.7 bits (60), Expect = 0.73
Identities = 16/51 (31%), Positives = 26/51 (50%), Gaps = 7/51 (13%)
Frame = +3
Query: 27 DDQGKVCSTTSSSSIG-------RNSDDDDDDEVSSERSMDENEAESKYNG 70
D + ++ +T +SSS N+DDDD+D+ E S +E + K G
Sbjct: 174 DQETELPTTVNSSSDAVPNNEPENNTDDDDEDDEDEENSDEEPVVDRKGKG 326
>TC88662 similar to GP|17979181|gb|AAL49786.1 unknown protein {Arabidopsis
thaliana}, partial (7%)
Length = 674
Score = 27.7 bits (60), Expect = 0.73
Identities = 14/36 (38%), Positives = 21/36 (57%)
Frame = +1
Query: 49 DDDEVSSERSMDENEAESKYNGGALDCMEALEEVLP 84
DDD S ERS +++A+SK N ++ E L + P
Sbjct: 376 DDDRRSRERSGSQSKAKSKSNSSSVLSSEELSPLRP 483
Database: MTGI
Posted date: Oct 22, 2004 3:39 PM
Number of letters in database: 27,044,181
Number of sequences in database: 36,976
Lambda K H
0.312 0.129 0.360
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 2,689,206
Number of Sequences: 36976
Number of extensions: 30466
Number of successful extensions: 557
Number of sequences better than 10.0: 115
Number of HSP's better than 10.0 without gapping: 388
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 475
length of query: 105
length of database: 9,014,727
effective HSP length: 81
effective length of query: 24
effective length of database: 6,019,671
effective search space: 144472104
effective search space used: 144472104
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.2 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 42 (21.9 bits)
S2: 50 (23.9 bits)
Medicago: description of AC139743.4


