

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
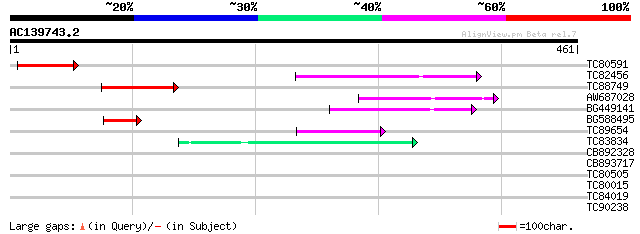
Query= AC139743.2 - phase: 0 /pseudo
(461 letters)
Database: MTGI
36,976 sequences; 27,044,181 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
TC80591 76 2e-14
TC82456 weakly similar to GP|18057159|gb|AAL58182.1 putative far... 68 7e-12
TC88749 68 8e-12
AW687028 weakly similar to GP|9909176|dbj| hypothetical protein~... 62 4e-10
BG449141 similar to GP|15983442|gb At1g10240/F14N23_12 {Arabidop... 55 6e-08
BG588495 47 2e-05
TC89654 weakly similar to PIR|T05644|T05644 hypothetical protein... 42 4e-04
TC83834 similar to PIR|G96565|G96565 F6D8.26 [imported] - Arabid... 41 9e-04
CB892328 GP|15075873|em PROBABLE GLYCOSYL HYDROLASE PROTEIN {Sin... 29 3.3
CB893717 similar to SP|O04933|SPS Sucrose-phosphate synthase 2 (... 29 3.3
TC80505 28 7.5
TC80015 similar to SP|Q9SGE9|SYFB_ARATH Probable phenylalanyl-tR... 28 7.5
TC84019 similar to GP|15451184|gb|AAK96863.1 putative protein {A... 28 9.7
TC90238 similar to PIR|T07126|T07126 magnesium chelatase (EC 4.9... 28 9.7
>TC80591
Length = 734
Score = 76.3 bits (186), Expect = 2e-14
Identities = 38/50 (76%), Positives = 42/50 (84%)
Frame = +1
Query: 7 VMVKNAKDVPGETKENFVENSGDVKPPNFEAKTNIDEASVHVEASKYVGG 56
VMV+NA+DV GETK+ FV SGDVKPP EAK+NID A VHVEASKYVGG
Sbjct: 580 VMVENAEDVLGETKQYFVRISGDVKPPKVEAKSNIDGAFVHVEASKYVGG 729
>TC82456 weakly similar to GP|18057159|gb|AAL58182.1 putative far-red
impaired response protein {Oryza sativa}, partial (7%)
Length = 1244
Score = 68.2 bits (165), Expect = 7e-12
Identities = 36/152 (23%), Positives = 72/152 (46%), Gaps = 1/152 (0%)
Frame = +2
Query: 233 RGDKKSLQFLISKLEEHNYTYYSRTQLESN-TIEDIFWAHPTSIKLFNNFPTVLVMDSTY 291
RGD ++ ++++ N +Y ++ I+++FWA + F V+ +D+TY
Sbjct: 710 RGDVDAIHSFFHRMQKQNSQFYCAMDMDDKRNIQNLFWADARCRAAYEYFGEVITLDTTY 889
Query: 292 KTNMYRMPMFEVVGVTSTDLTYSVGFGFMTHEKEENFVWVLTMLRKLLSSKMNMPKVIVT 351
T+ Y +P+ VGV T +G +++ + W+ T R L + P I+T
Sbjct: 890 LTSKYDLPLVPFVGVNHHGQTILLGCAILSNLDAKTLTWLFT--RWLECMHGHAPNGIIT 1063
Query: 352 DRDMSLMKAVAHVFPESYALNCFFHVQANVKQ 383
+ D ++ A+ FP++ C +H+ V +
Sbjct: 1064EEDKAMKNAIEVAFPKARHRWCLWHIMKKVPE 1159
>TC88749
Length = 1221
Score = 67.8 bits (164), Expect = 8e-12
Identities = 32/63 (50%), Positives = 46/63 (72%)
Frame = +2
Query: 75 DDVKSKNRDELLEWVRRQANKAGFTIVTQRSSLINPMFRLVCERSGSHIVPEKKPKHANT 134
+ V K+++++L +RRQAN+AGFT+ +RSS+ NPM L+CERSG H V +K+ KH T
Sbjct: 539 NQVTWKDKEDMLSSIRRQANRAGFTVGIKRSSIQNPMLELMCERSGDHKVSKKRLKHEAT 718
Query: 135 GSR 137
SR
Sbjct: 719 *SR 727
>AW687028 weakly similar to GP|9909176|dbj| hypothetical protein~similar to
Arabidopsis thaliana F20D10.300, partial (11%)
Length = 662
Score = 62.4 bits (150), Expect = 4e-10
Identities = 35/114 (30%), Positives = 56/114 (48%)
Frame = +1
Query: 284 VLVMDSTYKTNMYRMPMFEVVGVTSTDLTYSVGFGFMTHEKEENFVWVLTMLRKLLSSKM 343
VL D+TYK N Y P+ G T G + E E++ WVL + + +K
Sbjct: 4 VLAFDTTYKKNKYNYPLCIFSGCNHHSQTIIFGVALLEDETIESYKWVLNRFLECMENKF 183
Query: 344 NMPKVIVTDRDMSLMKAVAHVFPESYALNCFFHVQANVKQRCVLNCKYPLGFKK 397
PK +VTD D S+ +A+ VFP++ C +H+ N ++ + + GF+K
Sbjct: 184 --PKAVVTDGDGSMREAIKQVFPDASHRLCAWHLHKNAQEN-IKKTPFLEGFRK 336
>BG449141 similar to GP|15983442|gb At1g10240/F14N23_12 {Arabidopsis
thaliana}, partial (33%)
Length = 690
Score = 55.1 bits (131), Expect = 6e-08
Identities = 29/119 (24%), Positives = 63/119 (52%)
Frame = +3
Query: 261 SNTIEDIFWAHPTSIKLFNNFPTVLVMDSTYKTNMYRMPMFEVVGVTSTDLTYSVGFGFM 320
+N +E+I W++ +SI+L++ F +V D+T++ + MP+ VG+ + + G +
Sbjct: 312 NNRLENIAWSYASSIQLYDIFGDAVVFDTTHRLTAFDMPLGLWVGINNYGMPCFFGCVLL 491
Query: 321 THEKEENFVWVLTMLRKLLSSKMNMPKVIVTDRDMSLMKAVAHVFPESYALNCFFHVQA 379
E +F W + ++ K P+ I+TD+++ L +A++ P + C + + A
Sbjct: 492 RDETVRSFSWAIKAFLGFMNGK--APQTILTDQNICLKEALSAEMPMTKHAFCIWMIVA 662
>BG588495
Length = 540
Score = 47.0 bits (110), Expect = 2e-05
Identities = 22/31 (70%), Positives = 25/31 (79%)
Frame = +3
Query: 77 VKSKNRDELLEWVRRQANKAGFTIVTQRSSL 107
VKSK+RDELLEW R QANKAGFTI ++ L
Sbjct: 12 VKSKDRDELLEWTRHQANKAGFTIDDLKAQL 104
>TC89654 weakly similar to PIR|T05644|T05644 hypothetical protein F20D10.290
- Arabidopsis thaliana, partial (65%)
Length = 1358
Score = 42.4 bits (98), Expect = 4e-04
Identities = 22/72 (30%), Positives = 35/72 (48%)
Frame = +2
Query: 234 GDKKSLQFLISKLEEHNYTYYSRTQLESNTIEDIFWAHPTSIKLFNNFPTVLVMDSTYKT 293
G + L +L E++ +Y+ N +IFWA T ++ F ++ D+TYKT
Sbjct: 119 GGHQVLDYLKRMRAENHAFFYAVQSDVDNAGGNIFWADETCRTNYSYFGDTVIFDTTYKT 298
Query: 294 NMYRMPMFEVVG 305
N YR+P G
Sbjct: 299 NQYRVPFASFTG 334
>TC83834 similar to PIR|G96565|G96565 F6D8.26 [imported] - Arabidopsis
thaliana, partial (24%)
Length = 1032
Score = 41.2 bits (95), Expect = 9e-04
Identities = 34/196 (17%), Positives = 79/196 (39%), Gaps = 2/196 (1%)
Frame = +3
Query: 138 KCGCLFMISGYQSKQTKEWGLNILNGVHNHPMEPALEGHILAGRLKEDDKKIVRDLTKSK 197
+ GC MI + +++ W + + HNH + + I L D + K+
Sbjct: 414 RTGCPAMIR-MKLVESQRWRICEVTLEHNHVLGAKIHKSIKKNSLPSSDAE-----GKTI 575
Query: 198 MLPRNILIHLK-NQRPHCMTNVKQVYNERQQIWKANRGDKKSLQFLISKLEEHNYTYYSR 256
+ ++I + N + + +++ +GD +++ + +++ N ++
Sbjct: 576 KVYHALVIDTEGNDNLNSNARDDRAFSKYSNKLNLRKGDTQAIYNFLCRMQLTNPNFFYL 755
Query: 257 TQL-ESNTIEDIFWAHPTSIKLFNNFPTVLVMDSTYKTNMYRMPMFEVVGVTSTDLTYSV 315
+ + + W S F V+ D+TY N Y +P+ +VG+ + +
Sbjct: 756 MDFNDEGHLRNALWVDAKSRAACGYFSDVIYFDNTYLVNKYEIPLVALVGINHHGQSVLL 935
Query: 316 GFGFMTHEKEENFVWV 331
G G + E E++ W+
Sbjct: 936 GCGLLAGEIIESYKWL 983
>CB892328 GP|15075873|em PROBABLE GLYCOSYL HYDROLASE PROTEIN {Sinorhizobium
meliloti}, partial (1%)
Length = 828
Score = 29.3 bits (64), Expect = 3.3
Identities = 50/233 (21%), Positives = 88/233 (37%), Gaps = 20/233 (8%)
Frame = +1
Query: 63 QPHEVDTAKFFSDDVKSKNRDELLEWVRRQANKAGFTIVTQRSSLINPMFR------LVC 116
QP E D + ++DE + + A K GF++ + + + C
Sbjct: 124 QPIETDEPRDIYLGQVVNSKDEAYDLYQEHAFKTGFSVRKGKELYYDNEKKKTRLKDFYC 303
Query: 117 ERSG-SHIVPEKKPKHANTGSRKCGCLFMISGYQSKQTKE--WGLNILNGVHNHPMEPAL 173
+ G + P+ + + SR CL M+ + TKE W + L HNH P
Sbjct: 304 SKQGFKNNEPDGEVAYKRADSRT-NCLAMV---RFNVTKEGVWKVTKLILDHNHEFVPLQ 471
Query: 174 EGHILAG-----RLKEDDKKIVRDLTKSKMLPRNILIHLKNQ-----RPHCMTNVKQVYN 223
+ ++L LKED ++ L + N+ +L + + M Y
Sbjct: 472 QRYLLRSMRNMSNLKEDR---IKSLVNDAIRVTNVGGYLGKEVGGVDKRGVMLKDMHNYV 642
Query: 224 ERQQIWKANRGDKKSL-QFLISKLEEHNYTYYSRTQLESNTIEDIFWAHPTSI 275
++ GD +SL L SK + + YYS + + ++FW S+
Sbjct: 643 STGKLKFIEAGDAQSLLNHLQSKQAQDSMFYYSVQLDHESRLNNVFWRDGKSV 801
>CB893717 similar to SP|O04933|SPS Sucrose-phosphate synthase 2 (EC 2.4.1.14)
(UDP-glucose-fructose- phosphate glucosyltransferase
2)., partial (22%)
Length = 843
Score = 29.3 bits (64), Expect = 3.3
Identities = 18/41 (43%), Positives = 27/41 (64%), Gaps = 1/41 (2%)
Frame = -3
Query: 171 PALEGHILAGRLKEDDK-KIVRDLTKSKMLPRNILIHLKNQ 210
P LEGH+L +LK D K ++V+DL K + NI I L+++
Sbjct: 127 PPLEGHVLQPQLK*DQKAQLVQDL*KHPLSLPNIGICLEHR 5
>TC80505
Length = 675
Score = 28.1 bits (61), Expect = 7.5
Identities = 20/63 (31%), Positives = 27/63 (42%)
Frame = +1
Query: 260 ESNTIEDIFWAHPTSIKLFNNFPTVLVMDSTYKTNMYRMPMFEVVGVTSTDLTYSVGFGF 319
ESN F H ++ + NFPT ++S K N+Y F V + VGFG
Sbjct: 316 ESNNFSFSFLFHK*KLQKYTNFPTPFSINSPPKPNIYFFLSFLV*DL-------RVGFGT 474
Query: 320 MTH 322
H
Sbjct: 475 CHH 483
>TC80015 similar to SP|Q9SGE9|SYFB_ARATH Probable phenylalanyl-tRNA synthetase
beta chain (EC 6.1.1.20), partial (27%)
Length = 1401
Score = 28.1 bits (61), Expect = 7.5
Identities = 10/20 (50%), Positives = 17/20 (85%)
Frame = +3
Query: 59 VLPLQPHEVDTAKFFSDDVK 78
+L L+P ++DTA+FF++ VK
Sbjct: 1311 ILSLKPRQIDTAEFFANTVK 1370
>TC84019 similar to GP|15451184|gb|AAK96863.1 putative protein {Arabidopsis
thaliana}, partial (13%)
Length = 539
Score = 27.7 bits (60), Expect = 9.7
Identities = 11/21 (52%), Positives = 14/21 (66%)
Frame = +2
Query: 113 RLVCERSGSHIVPEKKPKHAN 133
R+VC GSH+ PE +HAN
Sbjct: 167 RIVCACHGSHMTPEDFVRHAN 229
>TC90238 similar to PIR|T07126|T07126 magnesium chelatase (EC 4.99.1.-)
chain chlH - soybean, partial (15%)
Length = 844
Score = 27.7 bits (60), Expect = 9.7
Identities = 10/40 (25%), Positives = 22/40 (55%)
Frame = -1
Query: 275 IKLFNNFPTVLVMDSTYKTNMYRMPMFEVVGVTSTDLTYS 314
+ +N +++M+ Y N+Y P+F + +T +L +S
Sbjct: 160 LSFYNKEKVLVMMERWYLRNLYTFPLFFQICLTKVELCFS 41
Database: MTGI
Posted date: Oct 22, 2004 3:39 PM
Number of letters in database: 27,044,181
Number of sequences in database: 36,976
Lambda K H
0.319 0.134 0.398
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 14,460,395
Number of Sequences: 36976
Number of extensions: 196115
Number of successful extensions: 1015
Number of sequences better than 10.0: 28
Number of HSP's better than 10.0 without gapping: 1011
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 1014
length of query: 461
length of database: 9,014,727
effective HSP length: 99
effective length of query: 362
effective length of database: 5,354,103
effective search space: 1938185286
effective search space used: 1938185286
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.8 bits)
S2: 60 (27.7 bits)
Medicago: description of AC139743.2


