

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
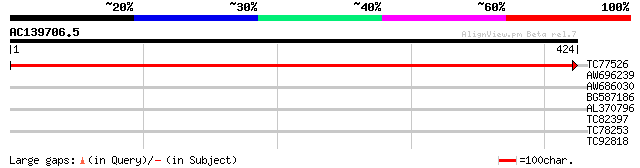
Query= AC139706.5 - phase: 0
(424 letters)
Database: MTGI
36,976 sequences; 27,044,181 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
TC77526 similar to GP|6177796|dbj|BAA86060.1 senescencs-related ... 849 0.0
AW696239 homologue to PIR|D71441|D714 hypothetical protein - Ara... 29 3.0
AW686030 weakly similar to PIR|T47508|T47 probable transporter p... 29 3.9
BG587186 similar to GP|18844936|dbj putative IAA-Ala hydrolase {... 28 5.1
AL370796 weakly similar to PIR|T47508|T475 probable transporter ... 28 6.7
TC82397 similar to GP|18377877|gb|AAL67124.1 AT3g53630/F4P12_330... 28 8.7
TC78253 weakly similar to GP|22654983|gb|AAM98084.1 AT5g45110/K1... 28 8.7
TC92818 homologue to GP|21592920|gb|AAM64870.1 putative microtub... 28 8.7
>TC77526 similar to GP|6177796|dbj|BAA86060.1 senescencs-related protein
{Pyrus pyrifolia}, partial (92%)
Length = 1725
Score = 849 bits (2194), Expect = 0.0
Identities = 424/424 (100%), Positives = 424/424 (100%)
Frame = +1
Query: 1 MANLSMTQLKTRNSASRFSFNFSKKVYPRPSRVDFKVFASEGQVSEPDLSVKVNGLHMPN 60
MANLSMTQLKTRNSASRFSFNFSKKVYPRPSRVDFKVFASEGQVSEPDLSVKVNGLHMPN
Sbjct: 187 MANLSMTQLKTRNSASRFSFNFSKKVYPRPSRVDFKVFASEGQVSEPDLSVKVNGLHMPN 366
Query: 61 PFVIGSGPPGTNYTVMKRAFDEGWGGVIAKTVSLDAAKVINVTPRYARLRANANGSAKGE 120
PFVIGSGPPGTNYTVMKRAFDEGWGGVIAKTVSLDAAKVINVTPRYARLRANANGSAKGE
Sbjct: 367 PFVIGSGPPGTNYTVMKRAFDEGWGGVIAKTVSLDAAKVINVTPRYARLRANANGSAKGE 546
Query: 121 IIGWQNIELISDRPLETMLKEFKQLKEEYPDRILIASIMEEYNKAAWEELIDRVEQTGID 180
IIGWQNIELISDRPLETMLKEFKQLKEEYPDRILIASIMEEYNKAAWEELIDRVEQTGID
Sbjct: 547 IIGWQNIELISDRPLETMLKEFKQLKEEYPDRILIASIMEEYNKAAWEELIDRVEQTGID 726
Query: 181 AIEINFSCPHGMPERKMGAAVGQDCALLEEVCGWINAKATVPVWAKMTPNITDISQPARV 240
AIEINFSCPHGMPERKMGAAVGQDCALLEEVCGWINAKATVPVWAKMTPNITDISQPARV
Sbjct: 727 AIEINFSCPHGMPERKMGAAVGQDCALLEEVCGWINAKATVPVWAKMTPNITDISQPARV 906
Query: 241 ALSSGCEGVAAINTIMSVMGINLNTLRPEPCVEGYSTPGGYSAKAVHPIALGKVMSIAKM 300
ALSSGCEGVAAINTIMSVMGINLNTLRPEPCVEGYSTPGGYSAKAVHPIALGKVMSIAKM
Sbjct: 907 ALSSGCEGVAAINTIMSVMGINLNTLRPEPCVEGYSTPGGYSAKAVHPIALGKVMSIAKM 1086
Query: 301 MKSEFDSENYSLSAIGGVETGGDAAEFILLGANTVQVCTGVMMHGYGLVKKLSLELKDFM 360
MKSEFDSENYSLSAIGGVETGGDAAEFILLGANTVQVCTGVMMHGYGLVKKLSLELKDFM
Sbjct: 1087MKSEFDSENYSLSAIGGVETGGDAAEFILLGANTVQVCTGVMMHGYGLVKKLSLELKDFM 1266
Query: 361 KKHNFTSIEDFRGASLQYFTTHTDLVHRQQEAIKQRKAIKKGLQSDKDWTGDGFVKESES 420
KKHNFTSIEDFRGASLQYFTTHTDLVHRQQEAIKQRKAIKKGLQSDKDWTGDGFVKESES
Sbjct: 1267KKHNFTSIEDFRGASLQYFTTHTDLVHRQQEAIKQRKAIKKGLQSDKDWTGDGFVKESES 1446
Query: 421 MVSN 424
MVSN
Sbjct: 1447MVSN 1458
>AW696239 homologue to PIR|D71441|D714 hypothetical protein - Arabidopsis
thaliana, partial (9%)
Length = 621
Score = 29.3 bits (64), Expect = 3.0
Identities = 13/45 (28%), Positives = 25/45 (54%)
Frame = +3
Query: 17 RFSFNFSKKVYPRPSRVDFKVFASEGQVSEPDLSVKVNGLHMPNP 61
+ +F+FSKK R S D + + S+P S ++N +++ +P
Sbjct: 3 QMAFSFSKKPSSRYSSYDSRSSTTSSHFSDPSSSYELNNMNLKHP 137
>AW686030 weakly similar to PIR|T47508|T47 probable transporter protein -
Arabidopsis thaliana, partial (20%)
Length = 630
Score = 28.9 bits (63), Expect = 3.9
Identities = 13/32 (40%), Positives = 20/32 (61%)
Frame = +3
Query: 240 VALSSGCEGVAAINTIMSVMGINLNTLRPEPC 271
VA +S C VA + TI+ + +N+L+P PC
Sbjct: 297 VAFASSC--VALLGTILLFLTATINSLKPHPC 386
>BG587186 similar to GP|18844936|dbj putative IAA-Ala hydrolase {Oryza sativa
(japonica cultivar-group)}, partial (11%)
Length = 598
Score = 28.5 bits (62), Expect = 5.1
Identities = 11/39 (28%), Positives = 19/39 (48%)
Frame = +2
Query: 150 PDRILIASIMEEYNKAAWEELIDRVEQTGIDAIEINFSC 188
PD + I ++K ++ +L R+E+ I I F C
Sbjct: 89 PDHVTIGGTFRAFSKESFNQLRQRIEEVSIGNKSIQFEC 205
>AL370796 weakly similar to PIR|T47508|T475 probable transporter protein -
Arabidopsis thaliana, partial (14%)
Length = 493
Score = 28.1 bits (61), Expect = 6.7
Identities = 12/32 (37%), Positives = 20/32 (62%)
Frame = +3
Query: 240 VALSSGCEGVAAINTIMSVMGINLNTLRPEPC 271
VA ++ C VA + TI+ + +N+L+P PC
Sbjct: 294 VAFTTSC--VALLGTILLTLTATINSLKPHPC 383
>TC82397 similar to GP|18377877|gb|AAL67124.1 AT3g53630/F4P12_330
{Arabidopsis thaliana}, partial (9%)
Length = 716
Score = 27.7 bits (60), Expect = 8.7
Identities = 13/38 (34%), Positives = 20/38 (52%)
Frame = -1
Query: 348 LVKKLSLELKDFMKKHNFTSIEDFRGASLQYFTTHTDL 385
L++KL L L F HN ++ SLQ+F T+ +
Sbjct: 572 LLQKLPLPLPLFSLHHNLPQFQNLNSQSLQFFFTNISI 459
>TC78253 weakly similar to GP|22654983|gb|AAM98084.1 AT5g45110/K17O22_11
{Arabidopsis thaliana}, partial (32%)
Length = 1669
Score = 27.7 bits (60), Expect = 8.7
Identities = 15/43 (34%), Positives = 25/43 (57%)
Frame = +1
Query: 153 ILIASIMEEYNKAAWEELIDRVEQTGIDAIEINFSCPHGMPER 195
IL+AS + N+ A + +DRV ++ +D I I PH + E+
Sbjct: 1483 ILMASFHCQLNQLA-AQCVDRVARSDLDQISIEKELPHELSEK 1608
>TC92818 homologue to GP|21592920|gb|AAM64870.1 putative
microtubule-associated protein {Arabidopsis thaliana},
partial (97%)
Length = 656
Score = 27.7 bits (60), Expect = 8.7
Identities = 14/53 (26%), Positives = 27/53 (50%)
Frame = +2
Query: 131 SDRPLETMLKEFKQLKEEYPDRILIASIMEEYNKAAWEELIDRVEQTGIDAIE 183
++ PLE E +++E+YPDRI + +++R E+T + I+
Sbjct: 155 TEHPLERRQAEAARIREKYPDRIPV--------------IVERAEKTDVPEID 271
Database: MTGI
Posted date: Oct 22, 2004 3:39 PM
Number of letters in database: 27,044,181
Number of sequences in database: 36,976
Lambda K H
0.316 0.133 0.386
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 10,923,662
Number of Sequences: 36976
Number of extensions: 131870
Number of successful extensions: 608
Number of sequences better than 10.0: 16
Number of HSP's better than 10.0 without gapping: 604
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 608
length of query: 424
length of database: 9,014,727
effective HSP length: 99
effective length of query: 325
effective length of database: 5,354,103
effective search space: 1740083475
effective search space used: 1740083475
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.3 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.6 bits)
S2: 60 (27.7 bits)
Medicago: description of AC139706.5


