

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC139356.2 + phase: 0
(180 letters)
Database: MTGI
36,976 sequences; 27,044,181 total letters
Searching..................................................done
Score E
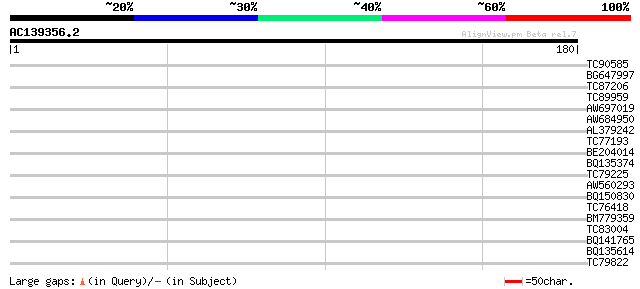
Sequences producing significant alignments: (bits) Value
TC90585 weakly similar to GP|13430834|gb|AAK26039.1 unknown prot... 30 0.68
BG647997 28 1.5
TC87206 similar to GP|10441354|gb|AAG17005.1 ARF GAP-like zinc f... 28 1.5
TC89959 similar to GP|15795137|dbj|BAB02515. transporter-like pr... 28 2.6
AW697019 weakly similar to GP|21554042|gb| AX110P-like protein {... 28 2.6
AW684950 similar to SP|Q39055|CNX2 Molybdopterin biosynthesis CN... 27 3.4
AL379242 27 4.4
TC77193 homologue to GP|21593495|gb|AAM65462.1 glycine rich prot... 27 4.4
BE204014 27 4.4
BQ135374 similar to PIR|S50766|S507 dehydrin-related protein - g... 27 5.7
TC79225 similar to PIR|T51820|T51820 F-actin capping protein alp... 27 5.7
AW560293 weakly similar to GP|21902022|db contains EST D24834(R2... 27 5.7
BQ150830 27 5.7
TC76418 similar to GP|16226228|gb|AAL16109.1 At1g20110/T20H2_10 ... 26 7.5
BM779359 similar to PIR|T05016|T050 hypothetical protein T19P19.... 26 9.8
TC83004 similar to PIR|T10985|T10985 regulator protein ROM2 - ki... 26 9.8
BQ141765 similar to GP|6721160|gb|A hypothetical protein {Arabid... 26 9.8
BQ135614 similar to GP|13486729|dbj hypothetical protein {Oryza ... 26 9.8
TC79822 similar to GP|21536770|gb|AAM61102.1 putative UDP-N-acet... 26 9.8
>TC90585 weakly similar to GP|13430834|gb|AAK26039.1 unknown protein
{Arabidopsis thaliana}, partial (12%)
Length = 803
Score = 29.6 bits (65), Expect = 0.68
Identities = 15/37 (40%), Positives = 20/37 (53%)
Frame = +1
Query: 109 NSVKPVIAEKKKNINWLFLLLGQMLGCCKLEQLKYFC 145
NS VIA+ +K ++ + CCK E LKYFC
Sbjct: 418 NSFLSVIAKMQKRRAENSTVMNPLQPCCKGEDLKYFC 528
>BG647997
Length = 790
Score = 28.5 bits (62), Expect = 1.5
Identities = 16/43 (37%), Positives = 24/43 (55%)
Frame = -1
Query: 35 IHNLNHLLQNKIFSITGDVQCKSCQTKFQMPFDLRTKFDVISQ 77
IH ++HL +F+ + + KS TKFQ F L KF ++ Q
Sbjct: 733 IHCVSHLFSTILFNPSCTISFKSKSTKFQHLFKL--KFSIVLQ 611
>TC87206 similar to GP|10441354|gb|AAG17005.1 ARF GAP-like zinc
finger-containing protein ZiGA4 {Arabidopsis thaliana},
partial (12%)
Length = 1984
Score = 28.5 bits (62), Expect = 1.5
Identities = 22/90 (24%), Positives = 35/90 (38%), Gaps = 8/90 (8%)
Frame = +1
Query: 44 NKIFSITGDVQCKSCQTKFQMPFDLRTKFDVISQYLVTNMNTMHDRAPKILM-------- 95
N F GD+Q K PFD DV + +MN++ P L+
Sbjct: 1396 NHEFPSNGDIQPNGINRKSTNPFDFPFDSDVEQNNMFLDMNSLQAALPDALLPATFGGIP 1575
Query: 96 NPRLPRCAHCNQENSVKPVIAEKKKNINWL 125
P LP +N+V P I+ + +++
Sbjct: 1576 EPWLP-------QNTVTPYISSAEGGFSFM 1644
>TC89959 similar to GP|15795137|dbj|BAB02515. transporter-like protein
{Arabidopsis thaliana}, partial (87%)
Length = 1681
Score = 27.7 bits (60), Expect = 2.6
Identities = 12/35 (34%), Positives = 22/35 (62%), Gaps = 1/35 (2%)
Frame = +3
Query: 24 PFIWATN-HRAKIHNLNHLLQNKIFSITGDVQCKS 57
PFIW+++ H + +HN +L I I+ + +C+S
Sbjct: 1293 PFIWSSHMHHSNLHNRVYLCPRDISHISKNHRCRS 1397
>AW697019 weakly similar to GP|21554042|gb| AX110P-like protein {Arabidopsis
thaliana}, partial (22%)
Length = 759
Score = 27.7 bits (60), Expect = 2.6
Identities = 16/46 (34%), Positives = 23/46 (49%), Gaps = 1/46 (2%)
Frame = -2
Query: 3 ANHTHVPPRRRSAPRSGPIPIPFI-WATNHRAKIHNLNHLLQNKIF 47
A H PPR+RS PIP+ + + IH+ H+LQ + F
Sbjct: 524 ATHCTPPPRQRSPSTPPPIPLHHVPPSVPAHIPIHSSPHILQARWF 387
>AW684950 similar to SP|Q39055|CNX2 Molybdopterin biosynthesis CNX2 protein
(Molybdenum cofactor biosynthesis enzyme CNX2)., partial
(30%)
Length = 603
Score = 27.3 bits (59), Expect = 3.4
Identities = 24/106 (22%), Positives = 39/106 (36%)
Frame = +1
Query: 7 HVPPRRRSAPRSGPIPIPFIWATNHRAKIHNLNHLLQNKIFSITGDVQCKSCQTKFQMPF 66
H P R + +S IP+ F + +I NL L + K + C S +
Sbjct: 64 HSPISRFNPSKSNSIPVNFEMGSYCATRICNLQTLFKGKSSGYSTVTSCDSFSEE----- 228
Query: 67 DLRTKFDVISQYLVTNMNTMHDRAPKILMNPRLPRCAHCNQENSVK 112
+ K + +S LV + +H L RC +C V+
Sbjct: 229 --KPKDNSVSDMLVDSFGRLHTYLRISLTERCNLRCQYCMPAEGVE 360
>AL379242
Length = 506
Score = 26.9 bits (58), Expect = 4.4
Identities = 22/86 (25%), Positives = 33/86 (37%), Gaps = 4/86 (4%)
Frame = +3
Query: 27 WATNHRAKIHNLNHLLQNKIFSITGDVQCKSCQTKFQMPFDLRTKFDVISQYLVTNMNTM 86
W HR + H+L H S G + +K++V++ YLV + T+
Sbjct: 33 WKIMHR*ETHSLVH-------SFNGTIS--------------ESKYEVLATYLVKTLETL 149
Query: 87 HDRAPKILMNPRL----PRCAHCNQE 108
H P + RL P C QE
Sbjct: 150 HPERPAEMTIWRLDW*GPTCLCIYQE 227
>TC77193 homologue to GP|21593495|gb|AAM65462.1 glycine rich protein-like
{Arabidopsis thaliana}, partial (85%)
Length = 989
Score = 26.9 bits (58), Expect = 4.4
Identities = 9/19 (47%), Positives = 12/19 (62%)
Frame = -3
Query: 9 PPRRRSAPRSGPIPIPFIW 27
PP + P S P P+PF+W
Sbjct: 492 PPAPLAPPSSKPRPVPFVW 436
>BE204014
Length = 658
Score = 26.9 bits (58), Expect = 4.4
Identities = 12/25 (48%), Positives = 16/25 (64%)
Frame = -1
Query: 31 HRAKIHNLNHLLQNKIFSITGDVQC 55
HR K+ NLN LL+N +F + QC
Sbjct: 148 HRIKVGNLNPLLENLVFHPSFAGQC 74
>BQ135374 similar to PIR|S50766|S507 dehydrin-related protein - garden pea,
partial (28%)
Length = 687
Score = 26.6 bits (57), Expect = 5.7
Identities = 15/49 (30%), Positives = 22/49 (44%)
Frame = -1
Query: 125 LFLLLGQMLGCCKLEQLKYFCKYNNHHRTGAKNRVLYLTYFELCQQLDP 173
LFL L ++ KL C YN K+ V++L Y+ + Q P
Sbjct: 285 LFLFLQPLVS*WKLHPKVTQCDYNQPQ*LPIKDDVIHLFYYHIANQKHP 139
>TC79225 similar to PIR|T51820|T51820 F-actin capping protein alpha chain
[imported] - Arabidopsis thaliana, partial (97%)
Length = 1223
Score = 26.6 bits (57), Expect = 5.7
Identities = 10/18 (55%), Positives = 12/18 (66%)
Frame = -1
Query: 135 CCKLEQLKYFCKYNNHHR 152
CCKLE L+ + NHHR
Sbjct: 239 CCKLEMLRKQLR*TNHHR 186
>AW560293 weakly similar to GP|21902022|db contains EST D24834(R2633)~similar
to Arabidopsis thaliana chromosome 5 At5g12010~unknown
protein, partial (25%)
Length = 623
Score = 26.6 bits (57), Expect = 5.7
Identities = 9/20 (45%), Positives = 14/20 (70%)
Frame = +2
Query: 80 VTNMNTMHDRAPKILMNPRL 99
+ N N++H R P+I NPR+
Sbjct: 491 IRNQNSIHQREPRISNNPRI 550
>BQ150830
Length = 544
Score = 26.6 bits (57), Expect = 5.7
Identities = 15/51 (29%), Positives = 28/51 (54%)
Frame = -2
Query: 74 VISQYLVTNMNTMHDRAPKILMNPRLPRCAHCNQENSVKPVIAEKKKNINW 124
++ YL++N + IL+ R+ R A C++ NS+ ++ K +NI W
Sbjct: 528 ILRLYLISNDILIRILYLSILL*LRV-RAARCSRSNSLLSIL*IKHRNIQW 379
>TC76418 similar to GP|16226228|gb|AAL16109.1 At1g20110/T20H2_10
{Arabidopsis thaliana}, partial (54%)
Length = 2020
Score = 26.2 bits (56), Expect = 7.5
Identities = 10/35 (28%), Positives = 17/35 (48%)
Frame = -2
Query: 16 PRSGPIPIPFIWATNHRAKIHNLNHLLQNKIFSIT 50
P P +P H ++HNL+H +I ++T
Sbjct: 654 PYHAPHKVPLFLLHKHTPRLHNLHHHHHQRIHTLT 550
>BM779359 similar to PIR|T05016|T050 hypothetical protein T19P19.180 -
Arabidopsis thaliana, partial (5%)
Length = 563
Score = 25.8 bits (55), Expect = 9.8
Identities = 18/60 (30%), Positives = 27/60 (45%)
Frame = -2
Query: 91 PKILMNPRLPRCAHCNQENSVKPVIAEKKKNINWLFLLLGQMLGCCKLEQLKYFCKYNNH 150
P L+NP L CN + S++ I K + F L+G+++ K F KY H
Sbjct: 280 PTELINPDLEIKLRCNLQTSMENKIKLKSLLMKIKFKLMGRLM------VSKNFWKYEEH 119
>TC83004 similar to PIR|T10985|T10985 regulator protein ROM2 - kidney bean,
partial (78%)
Length = 1250
Score = 25.8 bits (55), Expect = 9.8
Identities = 9/22 (40%), Positives = 13/22 (58%)
Frame = +1
Query: 6 THVPPRRRSAPRSGPIPIPFIW 27
+H+PP SA G P P++W
Sbjct: 79 SHIPPYFNSAAAPGHAPHPYMW 144
>BQ141765 similar to GP|6721160|gb|A hypothetical protein {Arabidopsis
thaliana}, partial (2%)
Length = 802
Score = 25.8 bits (55), Expect = 9.8
Identities = 12/30 (40%), Positives = 20/30 (66%), Gaps = 3/30 (10%)
Frame = +1
Query: 6 THVPPR-RRSAPRSGPIPIPFIWA--TNHR 32
T++P + R PR+GP+P P I++ NH+
Sbjct: 103 TYIPFKLNRLTPRTGPVPAPAIFSREANHK 192
>BQ135614 similar to GP|13486729|dbj hypothetical protein {Oryza sativa
(japonica cultivar-group)}, partial (11%)
Length = 1384
Score = 25.8 bits (55), Expect = 9.8
Identities = 10/22 (45%), Positives = 14/22 (63%)
Frame = -3
Query: 159 VLYLTYFELCQQLDPSGPFNVR 180
VLYL Y +CQ + P+ F+ R
Sbjct: 887 VLYLAYLRICQHICPARRFSSR 822
>TC79822 similar to GP|21536770|gb|AAM61102.1 putative
UDP-N-acetylglucosamine-dolichyl-phosphate
N-acetylglucosaminephosphotransferase, partial (85%)
Length = 1617
Score = 25.8 bits (55), Expect = 9.8
Identities = 11/33 (33%), Positives = 18/33 (54%)
Frame = -2
Query: 135 CCKLEQLKYFCKYNNHHRTGAKNRVLYLTYFEL 167
CC++E LKY+C N+ + N L +F +
Sbjct: 449 CCEVEILKYYC---NNQKDNTNNYTQ*LRHFNI 360
Database: MTGI
Posted date: Oct 22, 2004 3:39 PM
Number of letters in database: 27,044,181
Number of sequences in database: 36,976
Lambda K H
0.325 0.138 0.449
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 7,997,555
Number of Sequences: 36976
Number of extensions: 135827
Number of successful extensions: 1138
Number of sequences better than 10.0: 38
Number of HSP's better than 10.0 without gapping: 1133
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 1138
length of query: 180
length of database: 9,014,727
effective HSP length: 90
effective length of query: 90
effective length of database: 5,686,887
effective search space: 511819830
effective search space used: 511819830
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 15 ( 7.0 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 40 (21.6 bits)
S2: 55 (25.8 bits)
Medicago: description of AC139356.2


