

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
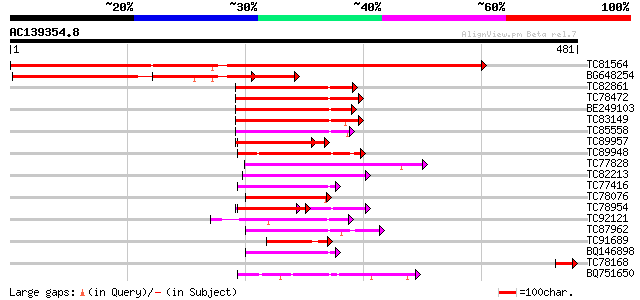
Query= AC139354.8 + phase: 0 /pseudo
(481 letters)
Database: MTGI
36,976 sequences; 27,044,181 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
TC81564 weakly similar to PIR|E85082|E85082 hypothetical protein... 741 0.0
BG648254 weakly similar to PIR|E85082|E85 hypothetical protein A... 244 7e-65
TC82861 similar to GP|6630450|gb|AAF19538.1| F23N19.10 {Arabidop... 82 5e-16
TC78472 similar to GP|16930441|gb|AAL31906.1 At2g42810/F7D19.19 ... 77 2e-14
BE249103 weakly similar to GP|6630450|gb| F23N19.10 {Arabidopsis... 76 3e-14
TC83149 similar to GP|13561982|gb|AAK30594.1 flagelliform silk p... 70 2e-12
TC85558 homologue to GP|7453538|gb|AAF62870.1| Toc64 {Pisum sati... 68 9e-12
TC89957 weakly similar to PIR|D96810|D96810 hypothetical protein... 65 8e-11
TC89948 similar to PIR|C96606|C96606 hypothetical protein F13N6.... 63 3e-10
TC77828 similar to GP|16648867|gb|AAL24285.1 HSP associated prot... 59 3e-09
TC82213 weakly similar to GP|7769858|gb|AAF69536.1| F12M16.20 {A... 57 1e-08
TC77416 similar to GP|18139887|gb|AAL60196.1 O-linked N-acetyl g... 53 2e-07
TC78076 similar to GP|17017308|gb|AAL33611.1 SGT1a {Arabidopsis ... 52 5e-07
TC78954 weakly similar to PIR|D96810|D96810 hypothetical protein... 44 3e-06
TC92121 weakly similar to GP|10177302|dbj|BAB10563. gene_id:MDC1... 49 4e-06
TC87962 similar to GP|14596039|gb|AAK68747.1 Unknown protein {Ar... 46 3e-05
TC91689 similar to PIR|T48150|T48150 stress-induced protein sti1... 45 6e-05
BQ146898 similar to GP|14596039|gb Unknown protein {Arabidopsis ... 45 8e-05
TC78168 similar to GP|18076028|emb|CAC80079. mercuric ion reduct... 45 8e-05
BQ751650 weakly similar to PIR|A32567|A32 stress-induced protein... 42 7e-04
>TC81564 weakly similar to PIR|E85082|E85082 hypothetical protein AT4g08320
[imported] - Arabidopsis thaliana, partial (46%)
Length = 1215
Score = 741 bits (1914), Expect = 0.0
Identities = 381/411 (92%), Positives = 384/411 (92%), Gaps = 7/411 (1%)
Frame = +3
Query: 1 MSHNRITTDSPLSHRIVRSFLHFLNQVEPSPGVDAEGIEVARECLVEAFKINNSASVTGE 60
MSHNRITTDSPLSH IVRSFLHFLNQVEPSPGVDAEGIEVARECLVEAFKINNSASVTGE
Sbjct: 6 MSHNRITTDSPLSHPIVRSFLHFLNQVEPSPGVDAEGIEVARECLVEAFKINNSASVTGE 185
Query: 61 PDSLIDIFKSFDANKQCERSRLDSMKASSSVSAQNAADAKTHPDESKPMDEDWTQGPHTS 120
PDSLIDIFKSFDANKQCERSRLDSMKASSSVSAQNAADAKTHPDESKPMDEDWTQGPHTS
Sbjct: 186 PDSLIDIFKSFDANKQCERSRLDSMKASSSVSAQNAADAKTHPDESKPMDEDWTQGPHTS 365
Query: 121 AVSKDELCGQFFAVLEKKHYFRTNIDGGDDIVQLEKASRLFDDGFTLYRR-------WKD 173
VSKDELCGQFFAVLEKKHYFRTNIDGGDDIVQLEKASRLFDDGFT R K+
Sbjct: 366 -VSKDELCGQFFAVLEKKHYFRTNIDGGDDIVQLEKASRLFDDGFTEMERSGCQQFNLKN 542
Query: 174 LGVSSLI*RIWLNH*KH*AMQSKQYFDAIELYNCAIAIYEKSAVYYCNRAAAYTQINRYT 233
L S L + AMQSKQYFDAIELYNCAIAIYEKSAVYYCNRAAAYTQINRYT
Sbjct: 543 LAES-------LKTLGNKAMQSKQYFDAIELYNCAIAIYEKSAVYYCNRAAAYTQINRYT 701
Query: 234 EAIQDSLRSIEIDPNYSKAYSRLGLAYYAQGNYRDAIDKGFKKALQLDPNNESVKENIRV 293
EAIQDSLRSIEIDPNYSKAYSRLGLAYYAQGNYRDAIDKGFKKALQLDPNNESVKENIRV
Sbjct: 702 EAIQDSLRSIEIDPNYSKAYSRLGLAYYAQGNYRDAIDKGFKKALQLDPNNESVKENIRV 881
Query: 294 AEHKLMEERHRADHNQNSRSQEFQNHYARGSRSHAAPASFGSMPFNPSNLASMFMAAANG 353
AEHKLMEERHRADHNQNSRSQEFQNHYARGSRSHAAPASFGSMPFNPSNLASMFMAAANG
Sbjct: 882 AEHKLMEERHRADHNQNSRSQEFQNHYARGSRSHAAPASFGSMPFNPSNLASMFMAAANG 1061
Query: 354 GQGSHSQEGQEDANSSGANEPEIRFGGNVNVNQDQIPQELRGAFQSVMHMF 404
GQGSHSQEGQEDANSSGANEPEIRFGG+VNVNQDQIPQELRGAFQSVMHMF
Sbjct: 1062GQGSHSQEGQEDANSSGANEPEIRFGGHVNVNQDQIPQELRGAFQSVMHMF 1214
>BG648254 weakly similar to PIR|E85082|E85 hypothetical protein AT4g08320
[imported] - Arabidopsis thaliana, partial (32%)
Length = 696
Score = 244 bits (622), Expect = 7e-65
Identities = 138/210 (65%), Positives = 150/210 (70%), Gaps = 3/210 (1%)
Frame = +1
Query: 3 HNRITTDSPLSHRIVRSFLHFLNQVEPSPGVDAEGIEVARECLVEAFKINNSASVTGEPD 62
HNRITTDSPLSHRIVRSFLHFLNQVEPSPGVDAEGIEVARECLVEAFKINNSASVTGEPD
Sbjct: 1 HNRITTDSPLSHRIVRSFLHFLNQVEPSPGVDAEGIEVARECLVEAFKINNSASVTGEPD 180
Query: 63 SLIDIFKSFDANKQCERSRLDSMKASSSVSAQNAADAKTHPDESKPMDEDWTQGPHTSAV 122
SLIDIFKSFDANKQCERSRLDSMKASSSVSAQNAADAKTHPDESKPM
Sbjct: 181 SLIDIFKSFDANKQCERSRLDSMKASSSVSAQNAADAKTHPDESKPM------------- 321
Query: 123 SKDELCGQFFAVLEKKHYFRTNIDGGDDIVQLE---KASRLFDDGFTLYRRWKDLGVSSL 179
C + V ++R N G ++++ + T+ RRWKDLGVSSL
Sbjct: 322 -----CLKMNYVDNSLLFWRKNTISGPILMEVMT*CNWRKHLAYLMTVLRRWKDLGVSSL 486
Query: 180 I*RIWLNH*KH*AMQSKQYFDAIELYNCAI 209
I*RIWLNH*KH* + + +C I
Sbjct: 487 I*RIWLNH*KH*VTRQCNPSSILMQLSCII 576
Score = 191 bits (486), Expect = 4e-49
Identities = 105/132 (79%), Positives = 107/132 (80%), Gaps = 7/132 (5%)
Frame = +3
Query: 122 VSKDELCGQFFAVLEKKHYFRTNIDGGDDIVQLEKASRLFDDGFTLYRR-------WKDL 174
VSKDELCGQFFAVLEKKHYFRTNIDGGDDIVQLEKASRLFDDGFT R K+L
Sbjct: 321 VSKDELCGQFFAVLEKKHYFRTNIDGGDDIVQLEKASRLFDDGFTEMERSGCQQFNLKNL 500
Query: 175 GVSSLI*RIWLNH*KH*AMQSKQYFDAIELYNCAIAIYEKSAVYYCNRAAAYTQINRYTE 234
S L + AMQSKQYFDAIELYNCAIAIYEKSAVYYCNRAAAYTQINRYTE
Sbjct: 501 AES-------LKTLGNKAMQSKQYFDAIELYNCAIAIYEKSAVYYCNRAAAYTQINRYTE 659
Query: 235 AIQDSLRSIEID 246
AIQDSLRSIEID
Sbjct: 660 AIQDSLRSIEID 695
>TC82861 similar to GP|6630450|gb|AAF19538.1| F23N19.10 {Arabidopsis
thaliana}, partial (29%)
Length = 750
Score = 82.0 bits (201), Expect = 5e-16
Identities = 40/104 (38%), Positives = 64/104 (61%)
Frame = +2
Query: 192 AMQSKQYFDAIELYNCAIAIYEKSAVYYCNRAAAYTQINRYTEAIQDSLRSIEIDPNYSK 251
A S + AI ++ AI + + V Y NR+AAY + YT+A+ D+ +++E+ P++SK
Sbjct: 116 AFSSGDFSTAIRHFSEAIDLSPTNHVLYSNRSAAYASLQNYTDALTDAKKTVELKPDWSK 295
Query: 252 AYSRLGLAYYAQGNYRDAIDKGFKKALQLDPNNESVKENIRVAE 295
YSRLG A+ Y DA+ +KK L++DPNNE +K + A+
Sbjct: 296 GYSRLGAAHLGLSQYDDAV-SAYKKGLEIDPNNEPLKSGLADAQ 424
>TC78472 similar to GP|16930441|gb|AAL31906.1 At2g42810/F7D19.19
{Arabidopsis thaliana}, partial (56%)
Length = 1050
Score = 76.6 bits (187), Expect = 2e-14
Identities = 37/109 (33%), Positives = 67/109 (60%)
Frame = +1
Query: 192 AMQSKQYFDAIELYNCAIAIYEKSAVYYCNRAAAYTQINRYTEAIQDSLRSIEIDPNYSK 251
A + +++ AI+LY AI + ++AVY NRA A+ ++ Y AI D+ ++IE+DP YSK
Sbjct: 256 AFKERKFAHAIDLYTQAIELNGQNAVYLANRAFAHLRLEEYGSAILDATKAIEVDPKYSK 435
Query: 252 AYSRLGLAYYAQGNYRDAIDKGFKKALQLDPNNESVKENIRVAEHKLME 300
Y R G A+ G +++A+ K F++ ++ PN+ + ++ E +M+
Sbjct: 436 GYYRRGAAHLGLGKFKEAL-KDFQQVKKMCPNDPDATKKLKECEKAVMK 579
>BE249103 weakly similar to GP|6630450|gb| F23N19.10 {Arabidopsis thaliana},
partial (20%)
Length = 613
Score = 75.9 bits (185), Expect = 3e-14
Identities = 39/103 (37%), Positives = 65/103 (62%)
Frame = +1
Query: 192 AMQSKQYFDAIELYNCAIAIYEKSAVYYCNRAAAYTQINRYTEAIQDSLRSIEIDPNYSK 251
A + + AI+ + IA+ + V Y NR+AA + RY EA++D+ + +E+ P++ K
Sbjct: 73 AFAAGDFETAIQHFTDGIAVDATNHVLYSNRSAAQASLKRYKEALEDAQKVVELKPDWPK 252
Query: 252 AYSRLGLAYYAQGNYRDAIDKGFKKALQLDPNNESVKENIRVA 294
+SRLG A G++ DAID +K+ALQL+P NE+ K+ ++ A
Sbjct: 253 GHSRLGTAKQGLGDWDDAID-AYKRALQLEPTNEAAKKALQDA 378
>TC83149 similar to GP|13561982|gb|AAK30594.1 flagelliform silk protein
{Argiope trifasciata}, partial (3%)
Length = 1177
Score = 69.7 bits (169), Expect = 2e-12
Identities = 40/111 (36%), Positives = 70/111 (63%), Gaps = 2/111 (1%)
Frame = +2
Query: 192 AMQSKQYFDAIELYNCAIAIYEKSAVYYCNRAAAYTQINRYTEAIQDSLRSIEIDPNYSK 251
AM SK Y AI LY A+++ +AVY NRAAA++ ++ A D+ ++ IDP Y+K
Sbjct: 272 AMASKDYPSAIALYTEALSLNPGNAVYLSNRAAAHSAAKDHSSARADAEAAVAIDPAYTK 451
Query: 252 AYSRLGLAYYAQGNYRDAIDKGFKKALQLDPN--NESVKENIRVAEHKLME 300
A+SRLGLA +A G+ + A++ + K ++ + + +E++K+ A+ ++ E
Sbjct: 452 AWSRLGLARFALGDAKGAME-AYGKGIEYEGSGGSEAMKKGYETAKRRVEE 601
>TC85558 homologue to GP|7453538|gb|AAF62870.1| Toc64 {Pisum sativum},
partial (40%)
Length = 1387
Score = 67.8 bits (164), Expect = 9e-12
Identities = 42/104 (40%), Positives = 57/104 (54%), Gaps = 3/104 (2%)
Frame = +3
Query: 192 AMQSKQYFDAIELYNCAIAIYEKSAVYYCNRAAAYTQINRYTEAIQDSLRSIEIDPNYSK 251
A + KQ+ AI Y AI + +A YY NRA AY ++ Y +A D ++I +D K
Sbjct: 606 AYKDKQWQKAIGFYTEAIKLCGNNATYYSNRAQAYLELGSYLQAEADCTKAISLDKKSVK 785
Query: 252 AYSRLGLAYYAQGNYRDAIDKGFKKALQLDPNNE---SVKENIR 292
AY R G A G Y++AID FK AL L+P N+ S E +R
Sbjct: 786 AYFRRGTAREMLGYYKEAID-DFKYALVLEPTNKRAASAAERLR 914
>TC89957 weakly similar to PIR|D96810|D96810 hypothetical protein T11I11.6
[imported] - Arabidopsis thaliana, partial (51%)
Length = 1843
Score = 64.7 bits (156), Expect = 8e-11
Identities = 30/80 (37%), Positives = 51/80 (63%)
Frame = +3
Query: 192 AMQSKQYFDAIELYNCAIAIYEKSAVYYCNRAAAYTQINRYTEAIQDSLRSIEIDPNYSK 251
A + ++ +A+ LY+ AIAI A Y+CN++AA + R+ EAI + SI +DP+Y++
Sbjct: 636 AYKKGKFEEALALYDKAIAIDSNKATYHCNKSAALIGLGRFQEAIIECEESIRLDPSYNR 815
Query: 252 AYSRLGLAYYAQGNYRDAID 271
A++RL Y+ G+ A+D
Sbjct: 816 AHNRLATIYFRLGDVEKALD 875
Score = 41.2 bits (95), Expect = 9e-04
Identities = 22/67 (32%), Positives = 41/67 (60%)
Frame = +3
Query: 194 QSKQYFDAIELYNCAIAIYEKSAVYYCNRAAAYTQINRYTEAIQDSLRSIEIDPNYSKAY 253
++ ++ +A +YN + ++V CNRAA +++ +Y +AI+D ++ ++P YSKA
Sbjct: 1341 KASKFMEACAVYNEGLDHDPHNSVLLCNRAACRSKLGQYEKAIEDCDAALMLNPCYSKA- 1517
Query: 254 SRLGLAY 260
RL AY
Sbjct: 1518 -RLRRAY 1535
>TC89948 similar to PIR|C96606|C96606 hypothetical protein F13N6.2
[imported] - Arabidopsis thaliana, partial (59%)
Length = 1691
Score = 62.8 bits (151), Expect = 3e-10
Identities = 36/109 (33%), Positives = 67/109 (61%)
Frame = +2
Query: 194 QSKQYFDAIELYNCAIAIYEKSAVYYCNRAAAYTQINRYTEAIQDSLRSIEIDPNYSKAY 253
+ K++ +AI+ Y+ +IA + +AV Y NRA A+ ++ R+ EA D ++ +D Y KAY
Sbjct: 350 KQKKFKEAIDCYSRSIA-FSPTAVAYANRAMAHIKLRRFQEAEDDCTEALNLDDRYIKAY 526
Query: 254 SRLGLAYYAQGNYRDAIDKGFKKALQLDPNNESVKENIRVAEHKLMEER 302
SR A G +++++ + A++L+PNN+ VK+ + A+ K + E+
Sbjct: 527 SRRATARKELGKNKESMEDA-EFAMRLEPNNQEVKK--QYADAKSLYEK 664
>TC77828 similar to GP|16648867|gb|AAL24285.1 HSP associated protein like
{Arabidopsis thaliana}, partial (83%)
Length = 1587
Score = 59.3 bits (142), Expect = 3e-09
Identities = 43/163 (26%), Positives = 71/163 (43%), Gaps = 8/163 (4%)
Frame = +3
Query: 200 DAIELYNCAIAIYEKSAVYYCNRAAAYTQINRYTEAIQDSLRSIEIDPNYSKAYSRLGLA 259
+AIE AI + SA+ Y NRA+ Y ++ + AI+D+ ++EI+P+ +K Y G+A
Sbjct: 504 EAIENLTEAIILNPTSAIMYANRASVYIKMKKPNAAIRDANAALEINPDSAKGYKSRGIA 683
Query: 260 YYAQGNYRDAI-DKGFKKALQLDPNNESVKENIRVAEHKLMEERHRADHNQNSRSQEFQN 318
G + +A D + D +V + + HKL E R + D + R ++
Sbjct: 684 RAMLGQWAEAAKDLHVASNIDYDEEINAVLKKVEPNAHKLEEHRRKYDRLRKEREEKKAE 863
Query: 319 HYARGSRSHAAPA-------SFGSMPFNPSNLASMFMAAANGG 354
+ R+ A A S NP + F GG
Sbjct: 864 RERQRRRAEAQAAYEKAKKQEQSSSSRNPGGMPGGFPGGMPGG 992
>TC82213 weakly similar to GP|7769858|gb|AAF69536.1| F12M16.20 {Arabidopsis
thaliana}, partial (25%)
Length = 604
Score = 57.4 bits (137), Expect = 1e-08
Identities = 34/109 (31%), Positives = 54/109 (49%)
Frame = +1
Query: 198 YFDAIELYNCAIAIYEKSAVYYCNRAAAYTQINRYTEAIQDSLRSIEIDPNYSKAYSRLG 257
+ DA+ LY+ AI++ SA Y NRAAA T + R EA+++ ++ +DPNYS+A+ RL
Sbjct: 97 FSDALSLYDRAISMSPGSASYRSNRAAALTGLGRLAEAVKECEEAVRLDPNYSRAHYRLA 276
Query: 258 LAYYAQGNYRDAIDKGFKKALQLDPNNESVKENIRVAEHKLMEERHRAD 306
+ G +A L DP + + +K + R D
Sbjct: 277 SLFIRLGQVENARKHLCHPGLTPDPTEMQKLKMVEKHINKCADVRRVGD 423
>TC77416 similar to GP|18139887|gb|AAL60196.1 O-linked N-acetyl glucosamine
transferase {Arabidopsis thaliana}, partial (94%)
Length = 3465
Score = 53.1 bits (126), Expect = 2e-07
Identities = 31/87 (35%), Positives = 48/87 (54%)
Frame = +2
Query: 194 QSKQYFDAIELYNCAIAIYEKSAVYYCNRAAAYTQINRYTEAIQDSLRSIEIDPNYSKAY 253
Q Y DAI YN + I +A NR Y +I R ++AIQD +R+I + P ++A+
Sbjct: 1445 QQGNYADAISCYNEVLRIDPLAADGLVNRGNTYKEIGRVSDAIQDYIRAITVRPTMAEAH 1624
Query: 254 SRLGLAYYAQGNYRDAIDKGFKKALQL 280
+ L AY G+ A+ K +++AL L
Sbjct: 1625 ANLASAYKDSGHVEAAV-KSYRQALIL 1702
Score = 40.8 bits (94), Expect = 0.001
Identities = 35/138 (25%), Positives = 57/138 (40%)
Frame = +2
Query: 145 IDGGDDIVQLEKASRLFDDGFTLYRRWKDLGVSSLI*RIWLNH*KH*AMQSKQYFDAIEL 204
+D ++ L KA L + ++ Y + L + W N M+S + A++
Sbjct: 797 VDAHSNLGNLMKAQGLVQEAYSCYL--EALRIQPTFAIAWSNL-AGLFMESGDFNRALQY 967
Query: 205 YNCAIAIYEKSAVYYCNRAAAYTQINRYTEAIQDSLRSIEIDPNYSKAYSRLGLAYYAQG 264
Y A+ + Y N Y + EAI +++ PNY AY L +Y QG
Sbjct: 968 YKEAVKLKPSFPDAYLNLGNVYKALGMPQEAIACYQHALQTRPNYGMAYGNLASIHYEQG 1147
Query: 265 NYRDAIDKGFKKALQLDP 282
AI +K+A+ DP
Sbjct: 1148QLDMAI-LHYKQAIACDP 1198
Score = 39.3 bits (90), Expect = 0.003
Identities = 28/82 (34%), Positives = 43/82 (52%)
Frame = +2
Query: 201 AIELYNCAIAIYEKSAVYYCNRAAAYTQINRYTEAIQDSLRSIEIDPNYSKAYSRLGLAY 260
AI Y AI + A + N A+AY + R TEA Q +++ I+P A+S LG
Sbjct: 650 AIRYYLIAIELRPNFADAWSNLASAYMRKGRLTEAAQCCRQALAINPLMVDAHSNLGNLM 829
Query: 261 YAQGNYRDAIDKGFKKALQLDP 282
AQG ++A + +AL++ P
Sbjct: 830 KAQGLVQEAY-SCYLEALRIQP 892
Score = 38.5 bits (88), Expect = 0.006
Identities = 22/83 (26%), Positives = 37/83 (44%)
Frame = +2
Query: 200 DAIELYNCAIAIYEKSAVYYCNRAAAYTQINRYTEAIQDSLRSIEIDPNYSKAYSRLGLA 259
+AI+ YN +++ N Y + N A ++ + S Y+ L +
Sbjct: 1259 EAIQCYNQCLSLQPNHPQALTNLGNIYMEWNMVAAAASYYKATLNVTTGLSAPYNNLAII 1438
Query: 260 YYAQGNYRDAIDKGFKKALQLDP 282
Y QGNY DAI + + L++DP
Sbjct: 1439 YKQQGNYADAI-SCYNEVLRIDP 1504
Score = 33.5 bits (75), Expect = 0.19
Identities = 26/108 (24%), Positives = 52/108 (48%), Gaps = 3/108 (2%)
Frame = +2
Query: 190 H*AMQSKQYFDAIELYNCAIAIYEKSAVYYCNR---AAAYTQINRYTEAIQDSLRSIEID 246
H +S Y A+E N +YE++ + N A Y Q++ + + + ++ I+
Sbjct: 413 HQMYKSGSYKKALEHSN---TVYERNPLRTDNLLLLGAIYYQLHDFDMCVAKNEEALRIE 583
Query: 247 PNYSKAYSRLGLAYYAQGNYRDAIDKGFKKALQLDPNNESVKENIRVA 294
P++++ Y + A+ +GN AI + + A++L PN N+ A
Sbjct: 584 PHFAECYGNMANAWKEKGNIDLAI-RYYLIAIELRPNFADAWSNLASA 724
Score = 29.3 bits (64), Expect = 3.5
Identities = 23/94 (24%), Positives = 35/94 (36%)
Frame = +2
Query: 201 AIELYNCAIAIYEKSAVYYCNRAAAYTQINRYTEAIQDSLRSIEIDPNYSKAYSRLGLAY 260
A Y + + + Y N A Y Q Y +AI + IDP + G Y
Sbjct: 1364 AASYYKATLNVTTGLSAPYNNLAIIYKQQGNYADAISCYNEVLRIDPLAADGLVNRGNTY 1543
Query: 261 YAQGNYRDAIDKGFKKALQLDPNNESVKENIRVA 294
G DAI + + +A+ + P N+ A
Sbjct: 1544 KEIGRVSDAI-QDYIRAITVRPTMAEAHANLASA 1642
>TC78076 similar to GP|17017308|gb|AAL33611.1 SGT1a {Arabidopsis thaliana},
partial (87%)
Length = 1493
Score = 52.0 bits (123), Expect = 5e-07
Identities = 27/76 (35%), Positives = 48/76 (62%), Gaps = 3/76 (3%)
Frame = +1
Query: 201 AIELYNCAIAIYEKSAVYYCNRAAAYTQINRYTEAIQDSLRSIEIDPNYSKAYSRLGLAY 260
A++ Y+ AI I +A + +RA ++ ++N +TEA+ D+ ++I+++PN SKAY R G A
Sbjct: 145 AVDFYSQAIEIDPTNANLFADRAQSHIKLNAFTEAVSDANKAIQLNPNLSKAYLRKGTAC 324
Query: 261 YAQGNY---RDAIDKG 273
Y + A++KG
Sbjct: 325 INLEEYHTAKVALEKG 372
>TC78954 weakly similar to PIR|D96810|D96810 hypothetical protein T11I11.6
[imported] - Arabidopsis thaliana, partial (45%)
Length = 2329
Score = 43.9 bits (102), Expect(2) = 3e-06
Identities = 21/56 (37%), Positives = 33/56 (58%)
Frame = +3
Query: 192 AMQSKQYFDAIELYNCAIAIYEKSAVYYCNRAAAYTQINRYTEAIQDSLRSIEIDP 247
A + ++ +A+ LY AIAI A Y+CN++AA + R+ AI + +I IDP
Sbjct: 834 AYKQGRFSEALALYERAIAIDSNKATYHCNKSAALIGLGRFQHAIVECEEAIRIDP 1001
Score = 42.0 bits (97), Expect = 5e-04
Identities = 18/62 (29%), Positives = 38/62 (61%)
Frame = +1
Query: 194 QSKQYFDAIELYNCAIAIYEKSAVYYCNRAAAYTQINRYTEAIQDSLRSIEIDPNYSKAY 253
++ ++ +A +YN + +++ CNRAA +++ ++ +AI+D ++ + PNYSKA
Sbjct: 1540 KASKFTEACAVYNEGLEYDPFNSILLCNRAACRSKLGQFEKAIEDCNVALTVQPNYSKAM 1719
Query: 254 SR 255
R
Sbjct: 1720 LR 1725
Score = 25.0 bits (53), Expect(2) = 3e-06
Identities = 16/62 (25%), Positives = 29/62 (45%)
Frame = +1
Query: 245 IDPNYSKAYSRLGLAYYAQGNYRDAIDKGFKKALQLDPNNESVKENIRVAEHKLMEERHR 304
+ P+Y +A+SRL Y+ G A++ K+ D + + ++V +K E R
Sbjct: 994 LTPSYPRAHSRLATIYFRLGEAEKALNCN-KETPYFDSELDFQAQALQVHFNKCSEARKV 1170
Query: 305 AD 306
D
Sbjct: 1171 KD 1176
>TC92121 weakly similar to GP|10177302|dbj|BAB10563.
gene_id:MDC12.18~pir||F69210~similar to unknown protein
{Arabidopsis thaliana}, partial (76%)
Length = 845
Score = 48.9 bits (115), Expect = 4e-06
Identities = 35/123 (28%), Positives = 58/123 (46%), Gaps = 2/123 (1%)
Frame = +3
Query: 171 WKDLGVSSLI*RIWLNH*KH*AMQSKQYFDAIELYNCAIAIYEKSAVY--YCNRAAAYTQ 228
W +LGVS + S++ A E+Y A+++ K + N Y Q
Sbjct: 261 WSNLGVSLQL--------------SEEQSQAEEVYKWALSLATKQEAHAILSNMGILYRQ 398
Query: 229 INRYTEAIQDSLRSIEIDPNYSKAYSRLGLAYYAQGNYRDAIDKGFKKALQLDPNNESVK 288
+Y A +S+E+ P Y+ A++ LGL + A+G +A F+KALQ D ++ K
Sbjct: 399 QKKYELAKAMFTKSLELQPGYAPAFNNLGLVFIAEGLLEEA-KHCFEKALQSDSMLDAAK 575
Query: 289 ENI 291
N+
Sbjct: 576 SNL 584
>TC87962 similar to GP|14596039|gb|AAK68747.1 Unknown protein {Arabidopsis
thaliana}, partial (90%)
Length = 1268
Score = 46.2 bits (108), Expect = 3e-05
Identities = 35/122 (28%), Positives = 59/122 (47%), Gaps = 4/122 (3%)
Frame = +3
Query: 201 AIELYNCAIAIYEKSAVYYCNRAAAYTQINRYTEAIQDSLRSIEIDPNYSKAYSRLGLAY 260
AI+ Y AI + VY+ NRA + + N + +DS R+I++D + KA+ LGLA
Sbjct: 249 AIDAYTEAITLCPNVPVYFTNRALCHLKRNEWERVEEDSRRAIQLDSHSVKAHYMLGLAL 428
Query: 261 YAQGNYRDAIDKGFKKALQL----DPNNESVKENIRVAEHKLMEERHRADHNQNSRSQEF 316
+ + I + +KAL L +P V+E + + + + + + RS E
Sbjct: 429 LQKQEHTKGI-RELQKALDLGRGANPKGYMVEE---IWQEFAKAKYKEWERSSSQRSWEL 596
Query: 317 QN 318
QN
Sbjct: 597 QN 602
>TC91689 similar to PIR|T48150|T48150 stress-induced protein sti1-like
protein - Arabidopsis thaliana, partial (26%)
Length = 638
Score = 45.1 bits (105), Expect = 6e-05
Identities = 22/56 (39%), Positives = 34/56 (60%)
Frame = +2
Query: 219 YCNRAAAYTQINRYTEAIQDSLRSIEIDPNYSKAYSRLGLAYYAQGNYRDAIDKGF 274
Y NRAA YT++ E ++D+ + IE+DP ++K Y+R G A + + I KGF
Sbjct: 26 YSNRAACYTKLGAMPEGLKDAEKCIELDPTFTKGYTRKG----AGSVFHERI*KGF 181
>BQ146898 similar to GP|14596039|gb Unknown protein {Arabidopsis thaliana},
partial (44%)
Length = 669
Score = 44.7 bits (104), Expect = 8e-05
Identities = 27/80 (33%), Positives = 43/80 (53%)
Frame = +1
Query: 201 AIELYNCAIAIYEKSAVYYCNRAAAYTQINRYTEAIQDSLRSIEIDPNYSKAYSRLGLAY 260
AI+ Y AI + VY+ NRA + + N + +DS R+I++D + KA+ LGLA
Sbjct: 262 AIDAYTEAITLCPNVPVYFTNRALCHLKRNEWERVEEDSRRAIQLDSHSVKAHYMLGLAL 441
Query: 261 YAQGNYRDAIDKGFKKALQL 280
+ + I + +KAL L
Sbjct: 442 LQKQEHTKGI-RELQKALDL 498
>TC78168 similar to GP|18076028|emb|CAC80079. mercuric ion reductase
{Pseudomonas sp.}, partial (2%)
Length = 629
Score = 44.7 bits (104), Expect = 8e-05
Identities = 18/18 (100%), Positives = 18/18 (100%)
Frame = -2
Query: 464 GNAPPGQPHDQTNERTAP 481
GNAPPGQPHDQTNERTAP
Sbjct: 445 GNAPPGQPHDQTNERTAP 392
Score = 32.7 bits (73), Expect = 0.32
Identities = 16/41 (39%), Positives = 21/41 (51%)
Frame = -2
Query: 378 FGGNVNVNQDQIPQELRGAFQSVMHMFSGNAPPGQPHDQMN 418
F + + ++ +P RG GNAPPGQPHDQ N
Sbjct: 499 FSSDDDDDETLVPNSARG----------GNAPPGQPHDQTN 407
>BQ751650 weakly similar to PIR|A32567|A32 stress-induced protein STI1 -
yeast (Saccharomyces cerevisiae), partial (32%)
Length = 723
Score = 41.6 bits (96), Expect = 7e-04
Identities = 47/172 (27%), Positives = 79/172 (45%), Gaps = 17/172 (9%)
Frame = +3
Query: 194 QSKQYFDAIELYNCAIAIYEKSAVYYCNRAAAYTQ-------INRYTEAIQDSLRSIEID 246
+ + + +AI Y A ++ K Y N AAY + I T+A+++ R+I D
Sbjct: 96 KKRNFDEAIVHYGKAWELH-KDITYLNNLGAAYFEKGDFEKCIEACTQAVEEG-RAIYAD 269
Query: 247 -PNYSKAYSRLGLAYYAQGNYRDAIDKGFKKALQLDPNNESVKENIRVAEHKLMEERHRA 305
+K+Y+R+G AY QGN A++ +KK+L + V +R AE +E+ A
Sbjct: 270 FKMIAKSYARIGSAYEKQGNLEAAVE-NYKKSL-TEHRTPDVNAKLRAAERNKVEQARLA 443
Query: 306 --DHNQNSRSQEFQNHYARGSRSHAAPASFGSM-------PFNPSNLASMFM 348
D + ++E N + S A A++ M P SN A+ F+
Sbjct: 444 YIDPAKAEEAREEGNKKFKESDWPGAVAAYSEMIKRAPEDPRGYSNRAAAFV 599
Score = 38.5 bits (88), Expect = 0.006
Identities = 16/48 (33%), Positives = 28/48 (58%)
Frame = +3
Query: 219 YCNRAAAYTQINRYTEAIQDSLRSIEIDPNYSKAYSRLGLAYYAQGNY 266
Y NRAAA+ ++ + A++D +I+ DP + +AY R A++ Y
Sbjct: 573 YSNRAAAFVKLLEFPSAVEDCNTAIKKDPTFIRAYIRKSQAFFGMREY 716
Database: MTGI
Posted date: Oct 22, 2004 3:39 PM
Number of letters in database: 27,044,181
Number of sequences in database: 36,976
Lambda K H
0.317 0.132 0.387
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 13,114,561
Number of Sequences: 36976
Number of extensions: 164019
Number of successful extensions: 831
Number of sequences better than 10.0: 56
Number of HSP's better than 10.0 without gapping: 802
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 824
length of query: 481
length of database: 9,014,727
effective HSP length: 100
effective length of query: 381
effective length of database: 5,317,127
effective search space: 2025825387
effective search space used: 2025825387
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.3 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.7 bits)
S2: 60 (27.7 bits)
Medicago: description of AC139354.8


