

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
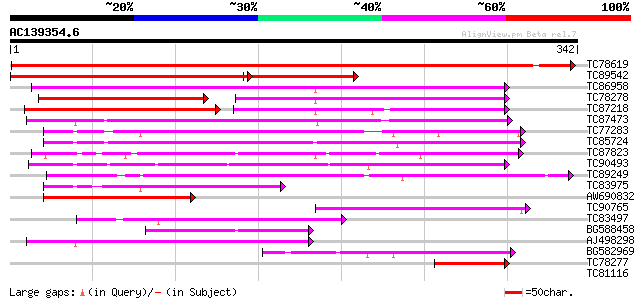
Query= AC139354.6 - phase: 0
(342 letters)
Database: MTGI
36,976 sequences; 27,044,181 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
TC78619 similar to GP|6552731|gb|AAF16530.1| T26F17.9 {Arabidops... 628 e-180
TC89542 homologue to PIR|F96805|F96805 unknown protein T5M16.20 ... 305 e-115
TC86958 similar to GP|12321869|gb|AAG50965.1 integral membrane p... 180 6e-46
TC78278 homologue to GP|12321869|gb|AAG50965.1 integral membrane... 99 4e-40
TC87218 similar to GP|8778643|gb|AAF79651.1| F5O11.25 {Arabidops... 91 1e-38
TC87473 similar to GP|15809998|gb|AAL06926.1 At2g25520/F13B15.18... 125 3e-29
TC77283 homologue to PIR|T06254|T06254 glucose-6-phosphate/phosp... 108 2e-24
TC85724 homologue to SP|P21727|CPTR_PEA Triose phosphate/phospha... 105 3e-23
TC87823 similar to GP|9295275|gb|AAF86907.1| phosphoenolpyruvate... 102 2e-22
TC90493 similar to GP|13937218|gb|AAK50101.1 AT5g17630/K10A8_110... 99 2e-21
TC89249 similar to GP|13486667|dbj|BAB39904. contains ESTs D4830... 95 3e-20
TC83975 similar to GP|14596173|gb|AAK68814.1 Similar to glucose-... 80 1e-15
AW690832 similar to GP|6056196|gb| unknown protein {Arabidopsis ... 75 4e-14
TC90765 similar to GP|6056196|gb|AAF02813.1| unknown protein {Ar... 71 7e-13
TC83497 70 2e-12
BG588458 similar to GP|9295275|gb| phosphoenolpyruvate/phosphate... 67 1e-11
AJ498298 similar to PIR|T05349|T05 hypothetical protein F8B4.90 ... 62 3e-10
BG582969 similar to PIR|T05401|T054 hypothetical protein F10M6.9... 52 3e-07
TC78277 homologue to GP|12321869|gb|AAG50965.1 integral membrane... 45 3e-05
TC81116 similar to PIR|T05401|T05401 hypothetical protein F10M6.... 39 0.004
>TC78619 similar to GP|6552731|gb|AAF16530.1| T26F17.9 {Arabidopsis
thaliana}, complete
Length = 1428
Score = 628 bits (1619), Expect = e-180
Identities = 297/340 (87%), Positives = 326/340 (95%)
Frame = +3
Query: 2 EEGLVCQWSVVRSLLAILQWWAFNVTVIIMNKWIFQKLDFKFPLSVSCIHFICSAIGAYV 61
E ++ QWSV+RSLL ILQWW FNVTVII+NKWIFQKLDFKFPLSVSC+HFICSAIGAY+
Sbjct: 120 ENSVLFQWSVIRSLLCILQWWTFNVTVIIVNKWIFQKLDFKFPLSVSCVHFICSAIGAYI 299
Query: 62 VIKVLKLKPLISVDPQDRWRRIFPMSFVFCINIVLGNVSLRYIPVSFMQTIKSFTPATTV 121
VIKVLKLKPLI+VDP+DRW+RIFPMSFVFCINIVLGNVSLRYIPVSFMQTIKSFTPATTV
Sbjct: 300 VIKVLKLKPLITVDPEDRWKRIFPMSFVFCINIVLGNVSLRYIPVSFMQTIKSFTPATTV 479
Query: 122 VLQWLVWRKYFDWRIWASLVPIVGGILLTSITELSFNMFGFCAALFGCLATSTKTILAEA 181
VLQWLVWRKYFDWRIWASL+PIVGGILLTS+TE+SFNMFGFCAAL GCLATSTKTILAE+
Sbjct: 480 VLQWLVWRKYFDWRIWASLIPIVGGILLTSVTEMSFNMFGFCAALLGCLATSTKTILAES 659
Query: 182 LLHGYKFDSINTVYHMAPFATLIMVFPALLLEGNGILEWFSIHPYPWAAMIIIFSSGVLA 241
LLHGYKFDSINTVY+MAP+AT+I+V PA+LLEGNG+LEW + HPYPW+A+IIIFSSGVLA
Sbjct: 660 LLHGYKFDSINTVYYMAPYATMILVLPAMLLEGNGVLEWLNTHPYPWSALIIIFSSGVLA 839
Query: 242 FCLNFSIFYVIHSTTAVTFNVAGNLKVAVAVLISWLIFRNPISYMNAVGCAITLVGCTFY 301
FCLNFSIFYVIHSTTAVTFNVAGNLKVAVAVL+SWLIFRNPISY+NAVGCAITLVGCTFY
Sbjct: 840 FCLNFSIFYVIHSTTAVTFNVAGNLKVAVAVLVSWLIFRNPISYLNAVGCAITLVGCTFY 1019
Query: 302 GYVRNMISQQPAVPGTPRTPRTPRSKMELLPLVNDKLDDK 341
GYVR+++SQQP VPG TPRTPRSKME LPLVNDKL++K
Sbjct: 1020GYVRHLLSQQPPVPG---TPRTPRSKMESLPLVNDKLENK 1130
>TC89542 homologue to PIR|F96805|F96805 unknown protein T5M16.20 [imported]
- Arabidopsis thaliana, partial (58%)
Length = 854
Score = 305 bits (781), Expect(2) = e-115
Identities = 146/146 (100%), Positives = 146/146 (100%)
Frame = +2
Query: 1 MEEGLVCQWSVVRSLLAILQWWAFNVTVIIMNKWIFQKLDFKFPLSVSCIHFICSAIGAY 60
MEEGLVCQWSVVRSLLAILQWWAFNVTVIIMNKWIFQKLDFKFPLSVSCIHFICSAIGAY
Sbjct: 224 MEEGLVCQWSVVRSLLAILQWWAFNVTVIIMNKWIFQKLDFKFPLSVSCIHFICSAIGAY 403
Query: 61 VVIKVLKLKPLISVDPQDRWRRIFPMSFVFCINIVLGNVSLRYIPVSFMQTIKSFTPATT 120
VVIKVLKLKPLISVDPQDRWRRIFPMSFVFCINIVLGNVSLRYIPVSFMQTIKSFTPATT
Sbjct: 404 VVIKVLKLKPLISVDPQDRWRRIFPMSFVFCINIVLGNVSLRYIPVSFMQTIKSFTPATT 583
Query: 121 VVLQWLVWRKYFDWRIWASLVPIVGG 146
VVLQWLVWRKYFDWRIWASLVPIVGG
Sbjct: 584 VVLQWLVWRKYFDWRIWASLVPIVGG 661
Score = 129 bits (323), Expect(2) = e-115
Identities = 63/69 (91%), Positives = 65/69 (93%)
Frame = +3
Query: 142 PIVGGILLTSITELSFNMFGFCAALFGCLATSTKTILAEALLHGYKFDSINTVYHMAPFA 201
P++ GILLTSITEL FNMFGFCAALFGCLATSTKTILA ALLHGYKFDSINTVYHMAPFA
Sbjct: 648 PLLEGILLTSITELXFNMFGFCAALFGCLATSTKTILAXALLHGYKFDSINTVYHMAPFA 827
Query: 202 TLIMVFPAL 210
TLIMV PAL
Sbjct: 828 TLIMVXPAL 854
>TC86958 similar to GP|12321869|gb|AAG50965.1 integral membrane protein
putative; 85705-84183 {Arabidopsis thaliana}, partial
(86%)
Length = 1467
Score = 180 bits (457), Expect = 6e-46
Identities = 97/290 (33%), Positives = 168/290 (57%), Gaps = 2/290 (0%)
Frame = +2
Query: 14 SLLAILQWWAFNVTVIIMNKWIFQKLDFKFPLSVSCIHFICSAIGAYVVIKVLKLKPLIS 73
++ I W++ N+ V+++NK++ FK+P+ ++ H ++ +YV I K+ P+
Sbjct: 116 TITLISAWYSSNIGVLLLNKYLLSNYGFKYPIFLTMCHMTACSLFSYVAIAWFKMVPMQF 295
Query: 74 VDPQDRWRRIFPMSFVFCINIVLGNVSLRYIPVSFMQTIKSFTPATTVVLQWLVWRKYFD 133
+ + ++ +I +SF+FC+++V GNVSLRY+PVSF Q I + TP T V + + K
Sbjct: 296 MRSRLQFFKIATLSFIFCVSVVFGNVSLRYLPVSFNQAIGATTPFFTAVFAYAMTLKREA 475
Query: 134 WRIWASLVPIVGGILLTSITELSFNMFGFCAALFGCLATSTKTILAEALL--HGYKFDSI 191
W + +LVP+V G+++ S E SF++FGF + A + KT+L LL G K +S+
Sbjct: 476 WLTYLALVPVVTGVIIASGGEPSFHLFGFIICVAATAARALKTVLQGILLSSEGEKLNSM 655
Query: 192 NTVYHMAPFATLIMVFPALLLEGNGILEWFSIHPYPWAAMIIIFSSGVLAFCLNFSIFYV 251
N + +MAP A + ++ L +E N + ++ + + + LA+ +N + F V
Sbjct: 656 NLLLYMAPMAVVFLLPATLYMEENVVGITLALARDDMKIIWYLLFNSALAYFVNLTNFLV 835
Query: 252 IHSTTAVTFNVAGNLKVAVAVLISWLIFRNPISYMNAVGCAITLVGCTFY 301
T+A+T V GN K AVAV++S LIFRNP+S +G ++T++G Y
Sbjct: 836 TKHTSALTLQVLGNAKGAVAVVVSILIFRNPVSVTGMMGYSLTVLGVVLY 985
>TC78278 homologue to GP|12321869|gb|AAG50965.1 integral membrane protein
putative; 85705-84183 {Arabidopsis thaliana}, partial
(87%)
Length = 1415
Score = 98.6 bits (244), Expect(2) = 4e-40
Identities = 57/167 (34%), Positives = 95/167 (56%), Gaps = 2/167 (1%)
Frame = +3
Query: 137 WASLVPIVGGILLTSITELSFNMFGFCAALFGCLATSTKTILAEALL--HGYKFDSINTV 194
+ +LVP+V G+++ S E SF++FGF + A + K++L LL G K +S+N +
Sbjct: 714 YLTLVPVVTGVVIASGGEPSFHLFGFIVCVAATAARALKSVLQGILLSSEGEKLNSMNLL 893
Query: 195 YHMAPFATLIMVFPALLLEGNGILEWFSIHPYPWAAMIIIFSSGVLAFCLNFSIFYVIHS 254
+MAP A + ++ L++E N + F++ + + + LA+ +N + F V
Sbjct: 894 LYMAPMAVVFLLPATLIMEENVVGITFALARDDTKIIWYLLFNSALAYFVNLTNFLVTKH 1073
Query: 255 TTAVTFNVAGNLKVAVAVLISWLIFRNPISYMNAVGCAITLVGCTFY 301
T+A+T V GN K AVAV++S LIFRNP+S +G +T+ G Y
Sbjct: 1074TSALTLQVLGNAKGAVAVVVSILIFRNPVSVTGMMGYGLTVFGVILY 1214
Score = 84.0 bits (206), Expect(2) = 4e-40
Identities = 34/103 (33%), Positives = 67/103 (65%)
Frame = +2
Query: 18 ILQWWAFNVTVIIMNKWIFQKLDFKFPLSVSCIHFICSAIGAYVVIKVLKLKPLISVDPQ 77
+ W++ N+ V+++NK++ FK+P+ ++ H ++ +YV I +K+ P+ ++ +
Sbjct: 356 VAAWYSSNIGVLLLNKYLLSNYGFKYPIFLTMCHMTACSLFSYVAIAWMKIVPMQTIRSR 535
Query: 78 DRWRRIFPMSFVFCINIVLGNVSLRYIPVSFMQTIKSFTPATT 120
++ +I +S +FC+++V GN+SLRY+PVSF Q I + TP T
Sbjct: 536 VQFFKISALSLIFCVSVVFGNISLRYLPVSFNQAIGATTPFFT 664
>TC87218 similar to GP|8778643|gb|AAF79651.1| F5O11.25 {Arabidopsis
thaliana}, partial (85%)
Length = 1557
Score = 90.9 bits (224), Expect(2) = 1e-38
Identities = 52/171 (30%), Positives = 95/171 (55%), Gaps = 5/171 (2%)
Frame = +2
Query: 136 IWASLVPIVGGILLTSITELSFNMFGFCAALFGCLATSTKTILAEALL--HGYKFDSINT 193
++ +L+P+V GI++++ +E F++ GF + + K+++ +L K S+N
Sbjct: 626 VYLALLPVVLGIVVSTNSEPLFHLLGFLVCVGSTAGRALKSVVQGIILTSEAEKLHSMNL 805
Query: 194 VYHMAPFATLIMVFPALLLEGNGI---LEWFSIHPYPWAAMIIIFSSGVLAFCLNFSIFY 250
+ +MAP A +I++ L +EGN +E P+ + ++ + +A+ +N + F
Sbjct: 806 LLYMAPLAAMILLPVTLYIEGNVFAITIEKARSDPF---IVFLLIGNATVAYLVNLTNFL 976
Query: 251 VIHSTTAVTFNVAGNLKVAVAVLISWLIFRNPISYMNAVGCAITLVGCTFY 301
V T+A+T V GN K AVA ++S LIFRNP++ M G IT++G Y
Sbjct: 977 VTKHTSALTLQVLGNAKAAVAAVVSVLIFRNPVTVMGMTGFGITIMGVVLY 1129
Score = 87.0 bits (214), Expect(2) = 1e-38
Identities = 37/118 (31%), Positives = 72/118 (60%)
Frame = +1
Query: 10 SVVRSLLAILQWWAFNVTVIIMNKWIFQKLDFKFPLSVSCIHFICSAIGAYVVIKVLKLK 69
++V + L I W+ N+ V+++NK++ +++P+ ++ +H + A +Y I V++
Sbjct: 247 NLVTTSLIIASWYFSNIGVLLLNKYLLSFYGYRYPIFLTMLHMLSCAAYSYAAINVVQFV 426
Query: 70 PLISVDPQDRWRRIFPMSFVFCINIVLGNVSLRYIPVSFMQTIKSFTPATTVVLQWLV 127
P + + ++ +IF +S +FC ++V GN SLRY+PVSF Q I + TP T + +L+
Sbjct: 427 PYQQIHSKKQFLKIFALSAIFCFSVVCGNTSLRYLPVSFNQAIGATTPFFTAIFAFLI 600
>TC87473 similar to GP|15809998|gb|AAL06926.1 At2g25520/F13B15.18
{Arabidopsis thaliana}, partial (95%)
Length = 1313
Score = 125 bits (313), Expect = 3e-29
Identities = 74/298 (24%), Positives = 156/298 (51%), Gaps = 5/298 (1%)
Frame = +2
Query: 11 VVRSLLAILQWWAFNVTVIIMNKWIFQK--LDFKFPLSVSCIHF-ICSAIGAYVVIKVLK 67
V+ S + W + TVI+ NK+I + ++ +P+S++ IH CS++ AYV+++V K
Sbjct: 173 VILSYTYVAIWIFLSFTVIVYNKYILDRKMYNWPYPISLTMIHMAFCSSL-AYVLVRVFK 349
Query: 68 LKPLISVDPQDRWRRIFPMSFVFCINIVLGNVSLRYIPVSFMQTIKSFTPATTVVLQWLV 127
L +S+ + + P+ ++ +++ N + Y+ VSF+Q +K+ P + L
Sbjct: 350 LVEPVSMSRDLYLKSVVPIGALYSLSLWFSNSAYIYLSVSFIQMLKALMPVAVYSIGVLF 529
Query: 128 WRKYFDWRIWASLVPIVGGILLTSITELSFNMFGFCAALFGCLATSTKTILAEALLH--G 185
++ F A+++ I G+ + + E F+ +G L +T+ +L + LL+ G
Sbjct: 530 KKEGFKNETMANMISISLGVAVAAYGEAKFDTWGVTLQLMAVAFEATRLVLIQILLNSKG 709
Query: 186 YKFDSINTVYHMAPFATLIMVFPALLLEGNGILEWFSIHPYPWAAMIIIFSSGVLAFCLN 245
+ I ++Y++AP + + P L++E + + S H I ++ + AF LN
Sbjct: 710 ISLNPITSLYYIAPCCLVFLSVPWLIVEYPSLRDNSSFH----LDFAIFGTNSLCAFALN 877
Query: 246 FSIFYVIHSTTAVTFNVAGNLKVAVAVLISWLIFRNPISYMNAVGCAITLVGCTFYGY 303
++F ++ T+A+T NVAG +K + + SW + ++ ++ +N +G + +G +Y +
Sbjct: 878 LAVFLLVGKTSALTMNVAGVVKDWLLIAFSWSVIKDTVTPINLIGYGLAFLGVAYYNH 1051
>TC77283 homologue to PIR|T06254|T06254
glucose-6-phosphate/phosphate-translocator precursor
plastid - garden pea, partial (98%)
Length = 1706
Score = 108 bits (271), Expect = 2e-24
Identities = 80/309 (25%), Positives = 144/309 (45%), Gaps = 18/309 (5%)
Frame = +1
Query: 21 WWAFNVTVIIMNKWIFQKLDFKFPLSVSCIHFICSAIGAYVVIKVLKLKPLISVDPQ--- 77
WWA NV I NK + + +P S + C ++ + ++ I+ P+
Sbjct: 538 WWALNVVFNIYNKKVLNA--YPYPWLTSTLSLACGSL-----MMLISWATRIAEAPKTDL 696
Query: 78 DRWRRIFPMSFVFCINIVLGNVSLRYIPVSFMQTIKSFTPATTVVLQWLVWRKYFDWRIW 137
+ W+ +FP++ I V VS+ + VSF IKS PA +V++ + + F ++
Sbjct: 697 EFWKTLFPVAVAHTIGHVAATVSMSKVAVSFTHIIKSGEPAFSVLVSRFILGETFPVPVY 876
Query: 138 ASLVPIVGGILLTSITELSFNMFGFCAALFGCLATSTKTILAEALLHGYKFDSINTVYHM 197
SL+PI+GG L ++TEL+FNM GF A+ LA + I ++ + G +N +
Sbjct: 877 LSLIPIIGGCALAAVTELNFNMIGFMGAMISNLAFVFRNIFSKKGMKGKSVSGMNYYACL 1056
Query: 198 APFATLIMVFPALLLEGNGILEWFSIHPYPWAA----MIIIFSSGVLAFCLNFSIFYVIH 253
+ + I+ A+ +EG P WAA + L + SIFY ++
Sbjct: 1057SILSLAILTPFAIAVEG----------PAMWAAGYKTALAEIGPQFLWWVAAQSIFYHLY 1206
Query: 254 STTA---------VTFNVAGNLKVAVAVLISWLIFRNPISYMNAVGCAITLVGCTFYGYV 304
+ + +TF++ +K ++ S +IF PI +NA+G AI + G Y
Sbjct: 1207NQVSYMSLDEISPLTFSIGNTMKRISVIVSSIIIFHTPIQPVNALGAAIAVFGTFLYSQA 1386
Query: 305 R--NMISQQ 311
+ N+++++
Sbjct: 1387KQ*NLLTER 1413
>TC85724 homologue to SP|P21727|CPTR_PEA Triose phosphate/phosphate
translocator chloroplast precursor (CTPT) (P36) (E30).
[Garden pea], complete
Length = 1581
Score = 105 bits (261), Expect = 3e-23
Identities = 74/295 (25%), Positives = 133/295 (45%), Gaps = 4/295 (1%)
Frame = +2
Query: 21 WWAFNVTVIIMNKWIFQKLDFKFPLSVSCIHFICSAIGAYVVIKVLKLKPLISVDPQDRW 80
W+ NV I+NK I+ F +P VS IH + + Y ++ P + ++
Sbjct: 434 WYFLNVIFNILNKKIYNY--FPYPYFVSVIHLLVGVV--YCLVSWTVGLPKRAPIDGNQL 601
Query: 81 RRIFPMSFVFCINIVLGNVSLRYIPVSFMQTIKSFTPATTVVLQWLVWRKYFDWRIWASL 140
+ + P++ + V NVS + VSF TIK+ P + + +W SL
Sbjct: 602 KLLIPVAVCHALGHVTSNVSFAAVAVSFTHTIKALEPFFNAAASQFILGQSIPITLWLSL 781
Query: 141 VPIVGGILLTSITELSFNMFGFCAALFGCLATSTKTILAEALLHGYKFDSINTVYHMAPF 200
P+V G+ L S+TELSFN GF +A+ ++ + ++I ++ + DS N +++
Sbjct: 782 APVVLGVSLASLTELSFNWLGFISAMISNISFTYRSIYSKKAM--TDMDSTNVYAYISII 955
Query: 201 ATLIMVFPALLLEGNGILEWFSIHPYPWAAMI----IIFSSGVLAFCLNFSIFYVIHSTT 256
A ++ + PAL++EG +L+ ++ +F G+ N +
Sbjct: 956 ALIVCIPPALIIEGPTLLKTGFADAIAKVGLVKFVSDLFWVGMFYHLYNQVATNTLERVA 1135
Query: 257 AVTFNVAGNLKVAVAVLISWLIFRNPISYMNAVGCAITLVGCTFYGYVRNMISQQ 311
+T V LK + S +IF N IS +G AI + G Y +++ I ++
Sbjct: 1136PLTHAVGNVLKRVFVIGFSIIIFGNKISTQTGIGTAIAIAGVALYSFIKAKIEEE 1300
>TC87823 similar to GP|9295275|gb|AAF86907.1| phosphoenolpyruvate/phosphate
translocator precursor {Mesembryanthemum crystallinum},
partial (74%)
Length = 1552
Score = 102 bits (254), Expect = 2e-22
Identities = 90/315 (28%), Positives = 155/315 (48%), Gaps = 18/315 (5%)
Frame = +3
Query: 14 SLLAILQ-------WWAFNVTVIIMNKWIFQKLDFKFPLSVSCIHFICSAIGAYVVIKVL 66
SLL LQ W+ FN+ I NK + + F P++V+ + F A+G +V +
Sbjct: 378 SLLKTLQLGSLFGLWYLFNIYFNIYNKQVLKACHF--PVTVTVVQF---AVGTVLVTFMW 542
Query: 67 KL----KPLISVDPQDRWRRIFPMSFVFCINIVLGNVSLRYIPVSFMQTIKSFTPATTVV 122
L +P I+ IFP++ V + + N+SL + VSF TIK+ P +V+
Sbjct: 543 ALNLYKRPKIT---GAMLAAIFPLAIVHTLGNLFTNMSLGKVAVSFTHTIKAMEPFFSVI 713
Query: 123 LQWL-VWRKYFDWRIWASLVPIVGGILLTSITELSFNMFGFCAALFGCLATSTKTILAEA 181
L + + + W I SLVPIVGG+ L SITE SFN GF +A+ + ++ +L++
Sbjct: 714 LSAMFLGERPTPWVI-GSLVPIVGGVALASITEASFNWAGFASAMASNVTNQSRNVLSKK 890
Query: 182 LL--HGYKFDSINTVYHMAPFATLIMVFP-ALLLEGNGILEWFSIHPYPWAAMIIIFSSG 238
++ D+I T++ + + ++ P A+ +EG + + + S
Sbjct: 891 VMVKQEESLDNI-TLFSIITIMSFFLLAPAAIFMEGVKFTPAY-LQSAGLDVRQVYTRSL 1064
Query: 239 VLAFCLNF---SIFYVIHSTTAVTFNVAGNLKVAVAVLISWLIFRNPISYMNAVGCAITL 295
+ A C + + ++ + VT +V +K V ++ S +IF+ P+S +NA G AI L
Sbjct: 1065LAALCFHAYQQVSYMILQRVSPVTHSVGNCVKRVVVIVSSVIIFKTPVSPVNAFGTAIAL 1244
Query: 296 VGCTFYGYVRNMISQ 310
G FY V+ + S+
Sbjct: 1245AGVFFYSRVKRIKSK 1289
>TC90493 similar to GP|13937218|gb|AAK50101.1 AT5g17630/K10A8_110
{Arabidopsis thaliana}, partial (70%)
Length = 1081
Score = 99.0 bits (245), Expect = 2e-21
Identities = 80/297 (26%), Positives = 140/297 (46%), Gaps = 7/297 (2%)
Frame = +1
Query: 12 VRSLLAILQWWAF-NVTVIIMNKWIFQKLDFKFPLSVSCIHFICSAIGAYVVIKVLKLKP 70
++ L + +W F N+ I NK + F FP ++ +I +V+ LKL+P
Sbjct: 160 LKKLALVFGFWYFQNIVFNIYNKKVLNI--FSFPWLLASFQLFVGSIWM-LVLWSLKLQP 330
Query: 71 LISVDPQDRWRRIFPMSFVFCINIVLGNVSLRYIPVSFMQTIKSFTPATTVVLQWLVWRK 130
+ + + P F I + VS + VSF IKS P +V+ ++ +
Sbjct: 331 CPKISKPFIFALLGPALF-HTIGHISACVSFSKVAVSFTHVIKSAEPVFSVIFSSVLGDR 507
Query: 131 YFDWRIWASLVPIVGGILLTSITELSFNMFGFCAALFGCLATSTKTILAEALLHGYK-FD 189
Y ++W S++PIV G L ++TE+SFN+ G AL + + I ++ L +K D
Sbjct: 508 Y-PIQVWLSILPIVLGCSLAAVTEVSFNVGGLWCALISNVGFVLRNIYSKKSLQNFKEVD 684
Query: 190 SINTVYHMAPFATLIMVFP-ALLLEGN----GILEWFSIHPYPWAAMIIIFSSGVLAFCL 244
+N +Y + + +FP A+ +EG+ G + P I + SG+
Sbjct: 685 GLN-LYGWITILSFMYLFPVAIFVEGSQWIPGYYKALEAIGTPSTFYIWVLVSGLFYHLY 861
Query: 245 NFSIFYVIHSTTAVTFNVAGNLKVAVAVLISWLIFRNPISYMNAVGCAITLVGCTFY 301
N S + + + +TF+V +K V ++ S L+FRNP+ +N +G AI ++G Y
Sbjct: 862 NQSSYQALDEISPLTFSVGNTMKRVVVIVSSILVFRNPVRPLNGLGSAIAILGTFLY 1032
>TC89249 similar to GP|13486667|dbj|BAB39904. contains ESTs D48306(S14443)
D24269(R1613) AU076096(E20048)~similar to Arabidopsis
thaliana, partial (78%)
Length = 1295
Score = 95.1 bits (235), Expect = 3e-20
Identities = 80/322 (24%), Positives = 146/322 (44%), Gaps = 4/322 (1%)
Frame = +2
Query: 23 AFNVTVIIMNKWIFQKLDFKFPLSVSCIHFICSAIGAYVVIKVLKLKPLISVDPQDRWRR 82
A +V+++I NK + L F F +++ H + + +V ++ L P D +
Sbjct: 194 ASSVSIVICNKALMSNLGFPFATTLTSWHLMVTFCTLHVALRF----NLFEAKPVDM-KT 358
Query: 83 IFPMSFVFCINIVLGNVSLRYIPVSFMQTIKSFTPATTVVLQWLVWRKYFDWRIWASLVP 142
+ + ++I N+SL + + F Q K TV+L+ L +K I SL
Sbjct: 359 VMLFGILNGVSIGFLNLSLGFNSIGFYQMTKLAIIPFTVMLETLFLKKQVSAGILNSLYF 538
Query: 143 -IVGGILLTSITELSFNMFGFCAALFGCLATSTKTILAEALLHGYKFDSINTVYHMAPFA 201
++ G+ + SIT+L N G +L + T IL + S +Y APF
Sbjct: 539 FLLVGVGIASITDLQLNFVGTILSLLAIITTCVGQILTNTIQKKLNVTSTQLLYQSAPFQ 718
Query: 202 TLIMVFPALLLEGNGILEWFSIHPYPWAAMIIIF--SSGVLAFCLNFSIFYVIHSTTAVT 259
I+ L++ +L ++ Y ++ +++ F S V+A +NFS F VI T+ VT
Sbjct: 719 AAILFVSGPLVDR--MLTKQNVFAYKYSPLVLTFIIMSCVIAGSVNFSTFLVIGKTSPVT 892
Query: 260 FNVAGNLKVAVAVLISWLIFRNPISYMNAVGCAITLVGCTFYGYVRNMISQQPAVPGTPR 319
+ V G+LK + + + + +P + N +G + + G Y Y +++ V P
Sbjct: 893 YQVLGHLKTCLVLGFGYTLLHDPFTERNIIGILVAVFGMGLYSYFCTQENKKKLVVDPPL 1072
Query: 320 TPRTPRSKMELLPLVNDK-LDD 340
+ + + K PL+N K +DD
Sbjct: 1073SSQV-KDKDINSPLLNGKNMDD 1135
>TC83975 similar to GP|14596173|gb|AAK68814.1 Similar to
glucose-6-phosphate/phosphate-translocator {Arabidopsis
thaliana}, partial (48%)
Length = 671
Score = 79.7 bits (195), Expect = 1e-15
Identities = 47/149 (31%), Positives = 79/149 (52%), Gaps = 3/149 (2%)
Frame = +3
Query: 21 WWAFNVTVIIMNKWIFQKLDFKFPLSVSCIHFICSAIGAYVVIKVLKLKPLISVDPQ--- 77
WWA NV I NK + F +P S + ++ A +I ++ ++ P+
Sbjct: 186 WWALNVVFNIYNKKVLNA--FPYPWLTSTL-----SLAAGSLIMLISWATRVAEAPKVNL 344
Query: 78 DRWRRIFPMSFVFCINIVLGNVSLRYIPVSFMQTIKSFTPATTVVLQWLVWRKYFDWRIW 137
+ W+ +FP++ I V VS+ + VSF IKS PA +V++ + + F +++
Sbjct: 345 EFWKALFPVAVAHTIGHVAATVSMSKVAVSFTHIIKSGEPAFSVLVSKFLLGEAFPLQVY 524
Query: 138 ASLVPIVGGILLTSITELSFNMFGFCAAL 166
SL+PI+GG L ++TEL+FNM GF A+
Sbjct: 525 LSLLPIIGGCALAAVTELNFNMIGFMGAM 611
>AW690832 similar to GP|6056196|gb| unknown protein {Arabidopsis thaliana},
partial (28%)
Length = 389
Score = 75.1 bits (183), Expect = 4e-14
Identities = 32/92 (34%), Positives = 58/92 (62%)
Frame = +3
Query: 21 WWAFNVTVIIMNKWIFQKLDFKFPLSVSCIHFICSAIGAYVVIKVLKLKPLISVDPQDRW 80
W++ N+ VI++NK++ FKFP+ ++ H AI +Y+ I K+ P + + ++
Sbjct: 108 WYSSNIGVILLNKYLISNYGFKFPIFLTMCHMTACAIFSYISIVFFKIVPQQMIKSRSQF 287
Query: 81 RRIFPMSFVFCINIVLGNVSLRYIPVSFMQTI 112
++ +SFVFC ++V GN+SL+Y+ VSF Q +
Sbjct: 288 LKVATLSFVFCGSVVGGNISLKYLAVSFNQAV 383
>TC90765 similar to GP|6056196|gb|AAF02813.1| unknown protein {Arabidopsis
thaliana}, partial (36%)
Length = 658
Score = 70.9 bits (172), Expect = 7e-13
Identities = 41/135 (30%), Positives = 73/135 (53%), Gaps = 5/135 (3%)
Frame = +3
Query: 185 GYKFDSINTVYHMAPFATLIMVFPALLLEGNGILEWFSIHPYPWAAMIIIFSSGVLAFCL 244
G K +S+N + +M+P A + ++ + +E N + S+ +++F + A+
Sbjct: 18 GEKLNSMNLLLYMSPIAVVFLLPAVVFMEPNVLDITLSLGKEHKFMGVLLFLNSAAAYGA 197
Query: 245 NFSIFYVIHSTTAVTFNVAGNLKVAVAVLISWLIFRNPISYMNAVGCAITLVGCTFYGYV 304
N + F V T+A+T V GN K AVAV+IS L+F+NP++++ G ++T++G YG
Sbjct: 198 NLTNFLVTKHTSALTLQVLGNAKGAVAVVISILLFQNPVTFIGMAGYSVTVMGVIAYGET 377
Query: 305 RNM-----ISQQPAV 314
+ IS P V
Sbjct: 378 KRRFR*TNISHSPIV 422
>TC83497
Length = 1131
Score = 69.7 bits (169), Expect = 2e-12
Identities = 39/166 (23%), Positives = 86/166 (51%), Gaps = 3/166 (1%)
Frame = +1
Query: 41 FKFPLSVSCIHFICSAIGAYVVIKVLKLKPLISVDPQDRWRRIFPMSF--VFCINIVLGN 98
F +P ++ +H C+++G Y +++ ++ R + ++F +F NI + N
Sbjct: 541 FPYPWLLTSLHATCASLGCYGLLQ----GGYFTMSHLGRRENLILLAFSLLFTTNIAVSN 708
Query: 99 VSLRYIPVSFMQTIKSFTPATTVVLQWLVWRKYFDWRIWASLVPIVGGILLTSITELSFN 158
+SL + V+F Q +++ P TV++ +++ + ++ + +LVPI+ G LT++ E +F
Sbjct: 709 LSLAMVSVAFYQVLRTTVPVFTVLIYRVLFGRTYENMTYLTLVPIMIGAALTTVGEYTFT 888
Query: 159 MFGFCAALFGCLATSTKTILAEALLHG-YKFDSINTVYHMAPFATL 203
GF G + + KT+ ++ G ++ + M+PFA +
Sbjct: 889 DLGFLLTFAGVMLAAVKTVATNRIMTGPLALPAMEVLLRMSPFAAM 1026
>BG588458 similar to GP|9295275|gb| phosphoenolpyruvate/phosphate
translocator precursor {Mesembryanthemum crystallinum},
partial (48%)
Length = 780
Score = 66.6 bits (161), Expect = 1e-11
Identities = 38/102 (37%), Positives = 61/102 (59%), Gaps = 1/102 (0%)
Frame = +2
Query: 83 IFPMSFVFCINIVLGNVSLRYIPVSFMQTIKSFTPATTVVLQWL-VWRKYFDWRIWASLV 141
IFP++ V + + N+SL + VSF TIK+ P +V+L + + + W I SLV
Sbjct: 335 IFPLAIVHTLGNLFTNMSLGKVAVSFTHTIKAMEPFFSVILSAMFLGERPTPWVI-GSLV 511
Query: 142 PIVGGILLTSITELSFNMFGFCAALFGCLATSTKTILAEALL 183
PIVGG+ L SITE SFN GF +A+ + ++ +L++ ++
Sbjct: 512 PIVGGVALASITEASFNWAGFASAMASNVTNQSRNVLSKKVM 637
>AJ498298 similar to PIR|T05349|T05 hypothetical protein F8B4.90 -
Arabidopsis thaliana, partial (54%)
Length = 600
Score = 62.0 bits (149), Expect = 3e-10
Identities = 38/175 (21%), Positives = 87/175 (49%), Gaps = 2/175 (1%)
Frame = +2
Query: 11 VVRSLLAILQWWAFNVTVIIMNKWIFQK--LDFKFPLSVSCIHFICSAIGAYVVIKVLKL 68
++ S + W + TVI+ NK+I K ++ FP+S++ IH A A ++++V K
Sbjct: 65 ILLSYTYVAIWIFLSFTVIVYNKYILDKKMYNWPFPISLTMIHMSFCATLAILLVRVFKF 244
Query: 69 KPLISVDPQDRWRRIFPMSFVFCINIVLGNVSLRYIPVSFMQTIKSFTPATTVVLQWLVW 128
+S+ + + + P+ ++ +++ L N + Y+ VSF+Q +K+ P + +
Sbjct: 245 VEPVSMSREVYFSSVVPIGALYSLSLWLSNSAYIYLSVSFIQMLKALMPVAVYSIGVGLR 424
Query: 129 RKYFDWRIWASLVPIVGGILLTSITELSFNMFGFCAALFGCLATSTKTILAEALL 183
++ + +++ I G+ + + E F+ +G L +T+ ++ + LL
Sbjct: 425 KESYKNDTMFNMLSISMGVAVAAYGEARFDTWGVILQLGAVAFEATRLVMIQILL 589
>BG582969 similar to PIR|T05401|T054 hypothetical protein F10M6.90 -
Arabidopsis thaliana, partial (18%)
Length = 783
Score = 52.0 bits (123), Expect = 3e-07
Identities = 43/158 (27%), Positives = 72/158 (45%), Gaps = 5/158 (3%)
Frame = +2
Query: 153 TELSFNMFGFCAALFGCLATSTKTILAEALLHGYKFDSINTVYHMAPFATLIMVFPALLL 212
++LSF+ +G+ LA T I + K +N+ M + I+ P LL+
Sbjct: 20 SKLSFDTYGYSVVF---LANVTTAIYLATIARIGKTSGLNSFGLM--WCNGILCGPVLLI 184
Query: 213 EG--NGILEWFSIHPYPWAA--MIIIFSSGVLAFCLNFSIFYVIHSTTAVTFNVAGNLKV 268
G L+ PY ++ ++I+ S +LAF LN+SIF +A+T + GN+K
Sbjct: 185 WTFIRGDLKTTIDFPYLFSPGFLVILLFSCILAFFLNYSIFLNTTLNSALTQTICGNMKD 364
Query: 269 AVAVLISWLIFRN-PISYMNAVGCAITLVGCTFYGYVR 305
+ W+IF P + N +G + G Y Y +
Sbjct: 365 LFTIGFGWIIFGGLPFDFWNVIGQFLGFTGSGLYAYFK 478
>TC78277 homologue to GP|12321869|gb|AAG50965.1 integral membrane protein
putative; 85705-84183 {Arabidopsis thaliana}, partial
(15%)
Length = 632
Score = 45.4 bits (106), Expect = 3e-05
Identities = 22/45 (48%), Positives = 29/45 (63%)
Frame = +1
Query: 257 AVTFNVAGNLKVAVAVLISWLIFRNPISYMNAVGCAITLVGCTFY 301
A+T V GN K AVAV++S LIFRNP+S +G +T+ G Y
Sbjct: 25 ALTLQVLGNAKGAVAVVVSILIFRNPVSVTGMMGYGLTVFGVILY 159
>TC81116 similar to PIR|T05401|T05401 hypothetical protein F10M6.90 -
Arabidopsis thaliana, partial (14%)
Length = 1199
Score = 38.5 bits (88), Expect = 0.004
Identities = 18/48 (37%), Positives = 28/48 (57%)
Frame = +1
Query: 232 IIIFSSGVLAFCLNFSIFYVIHSTTAVTFNVAGNLKVAVAVLISWLIF 279
+I+ S +LAF LN+SIF +A+T + GN+K + W+IF
Sbjct: 589 VILLFSCILAFFLNYSIFLNTTLNSALTQTICGNMKDLFTIGFGWIIF 732
Database: MTGI
Posted date: Oct 22, 2004 3:39 PM
Number of letters in database: 27,044,181
Number of sequences in database: 36,976
Lambda K H
0.331 0.142 0.464
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 13,710,149
Number of Sequences: 36976
Number of extensions: 231757
Number of successful extensions: 1923
Number of sequences better than 10.0: 62
Number of HSP's better than 10.0 without gapping: 1866
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 1909
length of query: 342
length of database: 9,014,727
effective HSP length: 97
effective length of query: 245
effective length of database: 5,428,055
effective search space: 1329873475
effective search space used: 1329873475
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 15 ( 7.2 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 40 (21.9 bits)
S2: 59 (27.3 bits)
Medicago: description of AC139354.6


