

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC139354.3 + phase: 0
(282 letters)
Database: MTGI
36,976 sequences; 27,044,181 total letters
Searching..................................................done
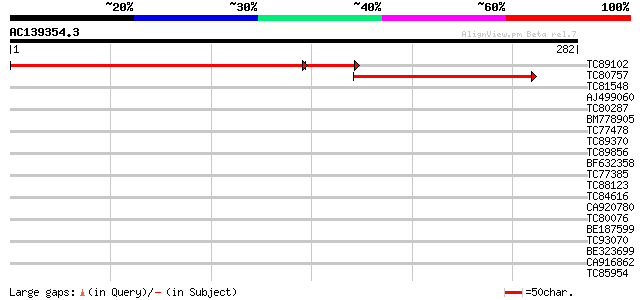
Score E
Sequences producing significant alignments: (bits) Value
TC89102 similar to PIR|F96506|F96506 hypothetical protein T12C22... 305 5e-95
TC80757 similar to GP|8978268|dbj|BAA98159.1 gene_id:K2I5.7~pir|... 74 6e-14
TC81548 weakly similar to GP|1931651|gb|AAB65486.1| membrane-ass... 36 0.015
AJ499060 weakly similar to GP|13122418|dbj putative glycin-rich ... 35 0.025
TC80287 weakly similar to GP|12667516|gb|AAG61157.1 DBC2 {Homo s... 35 0.025
BM778905 similar to SP|Q05046|CH62_ Chaperonin CPN60-2 mitochon... 32 0.27
TC77478 similar to GP|9884651|dbj|BAB11979.1 MGDG synthase type ... 31 0.47
TC89370 weakly similar to PIR|T01856|T01856 hypothetical protein... 31 0.61
TC89856 homologue to GP|4583990|emb|CAB40540.1 cyclin D3 {Medica... 31 0.61
BF632358 weakly similar to GP|19352296|gb MULTIPLE BANDED ANTIGE... 30 0.80
TC77385 similar to PIR|T04685|T04685 hypothetical protein F4B14.... 30 0.80
TC88123 similar to GP|8777484|dbj|BAA97064.1 contains similarity... 30 0.80
TC84616 similar to PIR|F96767|F96767 proteinase IV F2P9.14 [impo... 30 1.0
CA920780 similar to GP|21689829|gb| unknown protein {Arabidopsis... 27 1.3
TC80076 similar to GP|10177459|dbj|BAB10850. gene_id:MQB2.13~unk... 30 1.4
BE187599 homologue to GP|6176566|gb|A orphan nuclear receptor TE... 30 1.4
TC93070 weakly similar to GP|17065230|gb|AAL32769.1 Unknown prot... 29 1.8
BE323699 homologue to PIR|T39712|T397 hypothetical protein SPBC1... 29 1.8
CA916862 GP|12805231|gb Similar to mesenchyme homeobox 2 {Mus mu... 29 1.8
TC85954 homologue to GP|10719739|gb|AAG17942.1 putative pyridoxi... 29 1.8
>TC89102 similar to PIR|F96506|F96506 hypothetical protein T12C22.4
[imported] - Arabidopsis thaliana, partial (16%)
Length = 666
Score = 305 bits (782), Expect(2) = 5e-95
Identities = 148/148 (100%), Positives = 148/148 (100%)
Frame = +1
Query: 1 MRTHNAVEMAKTVLEVADVAWTAVEYHHHHHRHHDPVVTHDDKYDDKDNNCPSDQDLEAL 60
MRTHNAVEMAKTVLEVADVAWTAVEYHHHHHRHHDPVVTHDDKYDDKDNNCPSDQDLEAL
Sbjct: 139 MRTHNAVEMAKTVLEVADVAWTAVEYHHHHHRHHDPVVTHDDKYDDKDNNCPSDQDLEAL 318
Query: 61 KLENRRLRNLLDQNLKLLNNLSESNSFLSNCPPDLHVRLAATVRSDEYLTRLKVLQEETA 120
KLENRRLRNLLDQNLKLLNNLSESNSFLSNCPPDLHVRLAATVRSDEYLTRLKVLQEETA
Sbjct: 319 KLENRRLRNLLDQNLKLLNNLSESNSFLSNCPPDLHVRLAATVRSDEYLTRLKVLQEETA 498
Query: 121 SGGNQFPFKEATEVDYQSADILINVDSK 148
SGGNQFPFKEATEVDYQSADILINVDSK
Sbjct: 499 SGGNQFPFKEATEVDYQSADILINVDSK 582
Score = 60.1 bits (144), Expect(2) = 5e-95
Identities = 24/27 (88%), Positives = 24/27 (88%)
Frame = +3
Query: 148 KEPSWWVWVADETDPINFEECSGIDDE 174
K P WWVWVADETDPINF ECSGIDDE
Sbjct: 582 KNPVWWVWVADETDPINFXECSGIDDE 662
>TC80757 similar to GP|8978268|dbj|BAA98159.1
gene_id:K2I5.7~pir||T05575~similar to unknown protein
{Arabidopsis thaliana}, partial (26%)
Length = 969
Score = 73.9 bits (180), Expect = 6e-14
Identities = 31/91 (34%), Positives = 56/91 (61%)
Frame = +3
Query: 172 DDESYLIISEEHVVDGVANFMARCIMSNPKARNMSPEELQNNLSKAFAGTSKLEKVLDIW 231
D+E Y+++ ++ +VDG+A FMA ++S K ++++P +LQ+ LSK F+ K K+ W
Sbjct: 603 DEEDYVLVRQDDIVDGIACFMAAYLLSLKKTKDLTPSQLQDALSKTFSVKKKKGKLRKAW 782
Query: 232 AAGKLFYALSTWGLALAGLYQTRSLLKVAAK 262
K+ Y +++WG G+YQ +L+ K
Sbjct: 783 DGSKVIYNVASWGATAIGIYQNPVILRAGTK 875
>TC81548 weakly similar to GP|1931651|gb|AAB65486.1| membrane-associated
salt-inducible protein isolog; 88078-84012 {Arabidopsis
thaliana}, partial (23%)
Length = 882
Score = 36.2 bits (82), Expect = 0.015
Identities = 18/70 (25%), Positives = 35/70 (49%), Gaps = 6/70 (8%)
Frame = +1
Query: 24 VEYHHHHHRHH------DPVVTHDDKYDDKDNNCPSDQDLEALKLENRRLRNLLDQNLKL 77
+ +HHHHH HH ++ H ++ + +++N + DL + +L + D N
Sbjct: 73 IHHHHHHHHHHQFLLLLQQLIFHQEQENHQNSNFNTXSDLTSTLTSVGQLLTMKDLN-AT 249
Query: 78 LNNLSESNSF 87
L++ + SN F
Sbjct: 250 LHHFANSNKF 279
>AJ499060 weakly similar to GP|13122418|dbj putative glycin-rich protein
{Oryza sativa (japonica cultivar-group)}, partial (6%)
Length = 463
Score = 35.4 bits (80), Expect = 0.025
Identities = 20/62 (32%), Positives = 31/62 (49%), Gaps = 2/62 (3%)
Frame = +3
Query: 5 NAVEMAKTVLEVADVAWTAVEY--HHHHHRHHDPVVTHDDKYDDKDNNCPSDQDLEALKL 62
N + + +L+V+ +A A+E HHH H HH P T + PS D+ +L L
Sbjct: 90 NVLVLVLALLQVSYLAVLAIEVSTHHHSHPHHPPTPTPAPAHP------PSSYDIHSLSL 251
Query: 63 EN 64
+N
Sbjct: 252 DN 257
>TC80287 weakly similar to GP|12667516|gb|AAG61157.1 DBC2 {Homo sapiens},
partial (4%)
Length = 779
Score = 35.4 bits (80), Expect = 0.025
Identities = 25/71 (35%), Positives = 33/71 (46%), Gaps = 10/71 (14%)
Frame = -3
Query: 24 VEYHHHHH-RHHDPVVTHDDKYDDKD-NNCPSDQDLE--------ALKLENRRLRNLLDQ 73
VE HHHHH + + V HDD DD D + D DL+ KL +R+R L +
Sbjct: 486 VENHHHHHVGYSEEQVNHDDDDDDDDESEDDGDDDLDDELVPWDIGNKLGRQRMRKLGKR 307
Query: 74 NLKLLNNLSES 84
+NN S
Sbjct: 306 VSSKMNNSKRS 274
Score = 31.2 bits (69), Expect = 0.47
Identities = 10/21 (47%), Positives = 14/21 (66%)
Frame = +3
Query: 26 YHHHHHRHHDPVVTHDDKYDD 46
+HHHHH HH +V H ++ D
Sbjct: 408 HHHHHHHHHG*LVLHCTQHGD 470
>BM778905 similar to SP|Q05046|CH62_ Chaperonin CPN60-2 mitochondrial
precursor (HSP60-2). [Pumpkin Winter squash] {Cucurbita
maxima}, partial (9%)
Length = 456
Score = 32.0 bits (71), Expect = 0.27
Identities = 9/12 (75%), Positives = 10/12 (83%)
Frame = +2
Query: 26 YHHHHHRHHDPV 37
YHHHHH HH P+
Sbjct: 329 YHHHHHHHHQPL 364
>TC77478 similar to GP|9884651|dbj|BAB11979.1 MGDG synthase type A {Glycine
max}, partial (85%)
Length = 2229
Score = 31.2 bits (69), Expect = 0.47
Identities = 23/69 (33%), Positives = 40/69 (57%), Gaps = 4/69 (5%)
Frame = -2
Query: 87 FLSNCPPDLHVRLAATVRSDEYLTRLKVLQE---ETASGGNQFPFKEATEVDYQSADI-L 142
FL+ PP L +L+ TVR D+ TR+++L+E E + G + A+EV ++ D+
Sbjct: 671 FLNL*PPPLLFKLSETVRFDDDATRVRLLKER*LERSEAG-----EHASEVRLRNDDVSS 507
Query: 143 INVDSKEPS 151
+N +EP+
Sbjct: 506 VNRPIREPN 480
>TC89370 weakly similar to PIR|T01856|T01856 hypothetical protein F9D12.19 -
Arabidopsis thaliana, partial (15%)
Length = 710
Score = 30.8 bits (68), Expect = 0.61
Identities = 9/13 (69%), Positives = 10/13 (76%)
Frame = +2
Query: 27 HHHHHRHHDPVVT 39
HHHHH HH P +T
Sbjct: 161 HHHHHHHHHPSIT 199
Score = 27.3 bits (59), Expect = 6.8
Identities = 7/12 (58%), Positives = 9/12 (74%)
Frame = +2
Query: 26 YHHHHHRHHDPV 37
+HHHHH HH +
Sbjct: 161 HHHHHHHHHPSI 196
>TC89856 homologue to GP|4583990|emb|CAB40540.1 cyclin D3 {Medicago sativa},
partial (41%)
Length = 689
Score = 30.8 bits (68), Expect = 0.61
Identities = 11/28 (39%), Positives = 18/28 (64%), Gaps = 4/28 (14%)
Frame = +1
Query: 27 HHHHHRH----HDPVVTHDDKYDDKDNN 50
HHHH +H D + H++K++D DN+
Sbjct: 217 HHHHQQHTSSLFDALYCHEEKWEDDDND 300
Score = 30.4 bits (67), Expect = 0.80
Identities = 11/28 (39%), Positives = 13/28 (46%)
Frame = -2
Query: 29 HHHRHHDPVVTHDDKYDDKDNNCPSDQD 56
HHH H P+ HD K K+ C D
Sbjct: 304 HHHYHRLPIFLHDSKEHQKEKKCVVGDD 221
>BF632358 weakly similar to GP|19352296|gb MULTIPLE BANDED ANTIGEN
(FRAGMENT). 6/101 {Dictyostelium discoideum}, partial
(3%)
Length = 578
Score = 30.4 bits (67), Expect = 0.80
Identities = 9/16 (56%), Positives = 12/16 (74%)
Frame = -1
Query: 26 YHHHHHRHHDPVVTHD 41
+HHHHH +H P+ HD
Sbjct: 530 HHHHHHHYHYPLQHHD 483
>TC77385 similar to PIR|T04685|T04685 hypothetical protein F4B14.20 -
Arabidopsis thaliana, partial (90%)
Length = 1244
Score = 30.4 bits (67), Expect = 0.80
Identities = 8/18 (44%), Positives = 13/18 (71%)
Frame = +3
Query: 23 AVEYHHHHHRHHDPVVTH 40
+++ HHH+H HHD + H
Sbjct: 138 SIQIHHHNHNHHDSIHNH 191
>TC88123 similar to GP|8777484|dbj|BAA97064.1 contains similarity to
RNA-binding protein~gene_id:K15M2.15 {Arabidopsis
thaliana}, partial (11%)
Length = 815
Score = 30.4 bits (67), Expect = 0.80
Identities = 9/21 (42%), Positives = 15/21 (70%)
Frame = -2
Query: 26 YHHHHHRHHDPVVTHDDKYDD 46
+HHHH+RH P + H + ++D
Sbjct: 148 HHHHHYRHSQPPLAHCNLHND 86
>TC84616 similar to PIR|F96767|F96767 proteinase IV F2P9.14 [imported] -
Arabidopsis thaliana, partial (12%)
Length = 665
Score = 30.0 bits (66), Expect = 1.0
Identities = 9/12 (75%), Positives = 9/12 (75%)
Frame = +3
Query: 23 AVEYHHHHHRHH 34
A YHHHHH HH
Sbjct: 135 ATPYHHHHHHHH 170
>CA920780 similar to GP|21689829|gb| unknown protein {Arabidopsis thaliana},
partial (27%)
Length = 780
Score = 27.3 bits (59), Expect(2) = 1.3
Identities = 13/33 (39%), Positives = 18/33 (54%), Gaps = 8/33 (24%)
Frame = +3
Query: 26 YHHH-----HHRHHDPVVT---HDDKYDDKDNN 50
+HHH HRH+ P T DK++D+ NN
Sbjct: 525 FHHHSLQLRRHRHYSPPDTLQLQQDKHNDEHNN 623
Score = 20.8 bits (42), Expect(2) = 1.3
Identities = 6/27 (22%), Positives = 16/27 (59%)
Frame = +3
Query: 8 EMAKTVLEVADVAWTAVEYHHHHHRHH 34
E+++ ++ ++++ +HH HH H
Sbjct: 405 EVSRMLILLSNLIMHLCVHHHQHHDTH 485
>TC80076 similar to GP|10177459|dbj|BAB10850. gene_id:MQB2.13~unknown
protein {Arabidopsis thaliana}, partial (21%)
Length = 910
Score = 29.6 bits (65), Expect = 1.4
Identities = 8/15 (53%), Positives = 10/15 (66%)
Frame = +2
Query: 20 AWTAVEYHHHHHRHH 34
+W +HHHHH HH
Sbjct: 140 SWQPNHHHHHHHHHH 184
>BE187599 homologue to GP|6176566|gb|A orphan nuclear receptor TECdeltaC
short isoform {Mus musculus}, partial (2%)
Length = 586
Score = 29.6 bits (65), Expect = 1.4
Identities = 8/11 (72%), Positives = 9/11 (81%)
Frame = -3
Query: 26 YHHHHHRHHDP 36
+HHHHH HH P
Sbjct: 311 HHHHHHHHHQP 279
Score = 28.9 bits (63), Expect = 2.3
Identities = 8/9 (88%), Positives = 8/9 (88%)
Frame = -3
Query: 26 YHHHHHRHH 34
YHHHHH HH
Sbjct: 314 YHHHHHHHH 288
>TC93070 weakly similar to GP|17065230|gb|AAL32769.1 Unknown protein
{Arabidopsis thaliana}, partial (19%)
Length = 679
Score = 29.3 bits (64), Expect = 1.8
Identities = 8/11 (72%), Positives = 9/11 (81%)
Frame = -1
Query: 26 YHHHHHRHHDP 36
+HHHHH HH P
Sbjct: 319 HHHHHHHHHHP 287
Score = 28.1 bits (61), Expect = 4.0
Identities = 8/17 (47%), Positives = 11/17 (64%)
Frame = -1
Query: 24 VEYHHHHHRHHDPVVTH 40
+ +HHHHH HH + H
Sbjct: 322 LHHHHHHHHHHPHLHLH 272
>BE323699 homologue to PIR|T39712|T397 hypothetical protein SPBC17D11.01 -
fission yeast (Schizosaccharomyces pombe), partial (4%)
Length = 418
Score = 29.3 bits (64), Expect = 1.8
Identities = 10/20 (50%), Positives = 12/20 (60%)
Frame = +3
Query: 26 YHHHHHRHHDPVVTHDDKYD 45
+HHHHH HH H +YD
Sbjct: 201 HHHHHHHHHQH--QHQQQYD 254
>CA916862 GP|12805231|gb Similar to mesenchyme homeobox 2 {Mus musculus},
partial (5%)
Length = 727
Score = 29.3 bits (64), Expect = 1.8
Identities = 10/18 (55%), Positives = 11/18 (60%)
Frame = +2
Query: 17 ADVAWTAVEYHHHHHRHH 34
A A T +HHHHH HH
Sbjct: 392 ASTAATPTIHHHHHHHHH 445
>TC85954 homologue to GP|10719739|gb|AAG17942.1 putative pyridoxine
biosynthetic enzyme {Phaseolus vulgaris}, partial (94%)
Length = 1373
Score = 29.3 bits (64), Expect = 1.8
Identities = 8/10 (80%), Positives = 9/10 (90%)
Frame = -3
Query: 26 YHHHHHRHHD 35
+HHHHH HHD
Sbjct: 1047 HHHHHHHHHD 1018
Score = 28.9 bits (63), Expect = 2.3
Identities = 10/25 (40%), Positives = 14/25 (56%)
Frame = -3
Query: 18 DVAWTAVEYHHHHHRHHDPVVTHDD 42
+++ +HHHHH HH HDD
Sbjct: 1077 EISMPKENHHHHHHHHHH----HDD 1015
Database: MTGI
Posted date: Oct 22, 2004 3:39 PM
Number of letters in database: 27,044,181
Number of sequences in database: 36,976
Lambda K H
0.315 0.131 0.386
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 9,272,288
Number of Sequences: 36976
Number of extensions: 135659
Number of successful extensions: 3526
Number of sequences better than 10.0: 120
Number of HSP's better than 10.0 without gapping: 2027
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 2894
length of query: 282
length of database: 9,014,727
effective HSP length: 95
effective length of query: 187
effective length of database: 5,502,007
effective search space: 1028875309
effective search space used: 1028875309
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.3 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.6 bits)
S2: 58 (26.9 bits)
Medicago: description of AC139354.3


