

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC139354.10 - phase: 0
(381 letters)
Database: MTGI
36,976 sequences; 27,044,181 total letters
Searching..................................................done
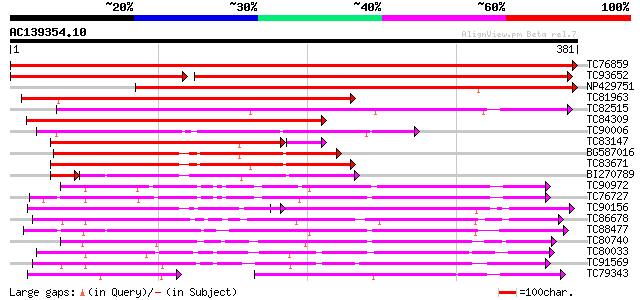
Score E
Sequences producing significant alignments: (bits) Value
TC76859 Enod8.1 [Medicago truncatula]; early nodule-specific pro... 779 0.0
TC93652 Enod8.2 [Medicago truncatula] 418 e-176
NP429751 NP429751|AF463407.1|AAL68830.1 Enod8.3 [Medicago trunca... 500 e-142
TC81963 Enod8-like protein [Medicago truncatula] 303 6e-83
TC82515 similar to GP|3688284|emb|CAA09694.1 lanatoside 15'-O-ac... 268 3e-72
TC84309 similar to GP|9294302|dbj|BAB02204.1 nodulin-like protei... 218 2e-57
TC90006 similar to PIR|A96590|A96590 hypothetical protein T22H22... 197 4e-51
TC83147 similar to PIR|T48618|T48618 early nodule-specific prote... 173 3e-45
BG587016 similar to PIR|T09416|T09 coil protein PO22 microspore... 177 6e-45
TC83671 similar to PIR|T09416|T09416 coil protein PO22 microspo... 173 1e-43
BI270789 similar to PIR|T09416|T09 coil protein PO22 microspore... 143 2e-36
TC90972 similar to GP|10638955|emb|CAB81548. putative proline-ri... 147 7e-36
TC76727 similar to GP|10638955|emb|CAB81548. putative proline-ri... 146 1e-35
TC90156 similar to PIR|B86227|B86227 hypothetical protein [impor... 145 3e-35
TC86678 similar to PIR|E96579|E96579 hypothetical protein T18A20... 140 6e-34
TC88477 similar to GP|21592417|gb|AAM64368.1 lipase/hydrolase p... 125 2e-29
TC80740 similar to GP|10177228|dbj|BAB10602. GDSL-motif lipase/h... 124 5e-29
TC80033 weakly similar to GP|21593518|gb|AAM65485.1 putative GDS... 124 8e-29
TC91569 similar to GP|18252233|gb|AAL61949.1 putative APG protei... 123 1e-28
TC79343 weakly similar to PIR|E86411|E86411 protein F1K23.18 [im... 121 5e-28
>TC76859 Enod8.1 [Medicago truncatula]; early nodule-specific protein
Length = 1350
Score = 779 bits (2011), Expect = 0.0
Identities = 381/381 (100%), Positives = 381/381 (100%)
Frame = +2
Query: 1 MAKIELSRHIPLVTLIVLVLCITPPIFATKNCDFPAIFSFGASNVDTGGLAAAFRAPPSP 60
MAKIELSRHIPLVTLIVLVLCITPPIFATKNCDFPAIFSFGASNVDTGGLAAAFRAPPSP
Sbjct: 47 MAKIELSRHIPLVTLIVLVLCITPPIFATKNCDFPAIFSFGASNVDTGGLAAAFRAPPSP 226
Query: 61 YGETYFHRSTGRFSDGRIILDFIARSFRLPYLSPYLNSLGSNFTHGANFASGGSTINIPK 120
YGETYFHRSTGRFSDGRIILDFIARSFRLPYLSPYLNSLGSNFTHGANFASGGSTINIPK
Sbjct: 227 YGETYFHRSTGRFSDGRIILDFIARSFRLPYLSPYLNSLGSNFTHGANFASGGSTINIPK 406
Query: 121 SILPNGKLSPFSLQIQYIQFKEFISKTKLIRDQGGVFATLIPKEDYFSKALYIFDIGQND 180
SILPNGKLSPFSLQIQYIQFKEFISKTKLIRDQGGVFATLIPKEDYFSKALYIFDIGQND
Sbjct: 407 SILPNGKLSPFSLQIQYIQFKEFISKTKLIRDQGGVFATLIPKEDYFSKALYIFDIGQND 586
Query: 181 LTIGFFGNKTIQQVNATVPDIVNNYIENIKNIYNLGARSFWIHGTGPKGCAPVILANFPS 240
LTIGFFGNKTIQQVNATVPDIVNNYIENIKNIYNLGARSFWIHGTGPKGCAPVILANFPS
Sbjct: 587 LTIGFFGNKTIQQVNATVPDIVNNYIENIKNIYNLGARSFWIHGTGPKGCAPVILANFPS 766
Query: 241 AIKDSYGCAKQYNEVSQYFNFKLKEALAELRSNLSSAAITYVDIYTPKYSLFTNPEKYGF 300
AIKDSYGCAKQYNEVSQYFNFKLKEALAELRSNLSSAAITYVDIYTPKYSLFTNPEKYGF
Sbjct: 767 AIKDSYGCAKQYNEVSQYFNFKLKEALAELRSNLSSAAITYVDIYTPKYSLFTNPEKYGF 946
Query: 301 ELPFVACCGYGGEYNIGVGCGASININGTKIVAGSCKNPSTRIIWDGVHYTEAANEIVFS 360
ELPFVACCGYGGEYNIGVGCGASININGTKIVAGSCKNPSTRIIWDGVHYTEAANEIVFS
Sbjct: 947 ELPFVACCGYGGEYNIGVGCGASININGTKIVAGSCKNPSTRIIWDGVHYTEAANEIVFS 1126
Query: 361 QILTGVFNDPPISLDRACYRK 381
QILTGVFNDPPISLDRACYRK
Sbjct: 1127QILTGVFNDPPISLDRACYRK 1189
>TC93652 Enod8.2 [Medicago truncatula]
Length = 1195
Score = 418 bits (1075), Expect(2) = e-176
Identities = 205/255 (80%), Positives = 220/255 (85%), Gaps = 1/255 (0%)
Frame = +2
Query: 125 NGKLSPFSLQIQ-YIQFKEFISKTKLIRDQGGVFATLIPKEDYFSKALYIFDIGQNDLTI 183
NG SPFSLQIQ K FISKT LIRDQGGVFATLIPKEDYFSKALY FDIGQNDL
Sbjct: 410 NGIFSPFSLQIQGTFNSKIFISKTNLIRDQGGVFATLIPKEDYFSKALYTFDIGQNDLIG 589
Query: 184 GFFGNKTIQQVNATVPDIVNNYIENIKNIYNLGARSFWIHGTGPKGCAPVILANFPSAIK 243
G+FGNKTI+QVNATVPDIVNN+I NIKNIYNLGARSFWIH T P GC P ILANFPSAIK
Sbjct: 590 GYFGNKTIKQVNATVPDIVNNFIVNIKNIYNLGARSFWIHSTVPSGCTPTILANFPSAIK 769
Query: 244 DSYGCAKQYNEVSQYFNFKLKEALAELRSNLSSAAITYVDIYTPKYSLFTNPEKYGFELP 303
DSYGCAKQYNEVSQYFN KLK+ALA+LR +L AAITYVDIY+P YSLF NP+KYGFELP
Sbjct: 770 DSYGCAKQYNEVSQYFNLKLKKALAQLRVDLPLAAITYVDIYSPNYSLFQNPKKYGFELP 949
Query: 304 FVACCGYGGEYNIGVGCGASININGTKIVAGSCKNPSTRIIWDGVHYTEAANEIVFSQIL 363
VACCGYGG+YNI VGCG ++NINGTKI AGSCKNPSTRIIWDG H+TE +IVF QI
Sbjct: 950 HVACCGYGGKYNIRVGCGETLNINGTKIEAGSCKNPSTRIIWDGSHFTERRYKIVFDQIS 1129
Query: 364 TGVFNDPPISLDRAC 378
TG F+DPPISL+RAC
Sbjct: 1130TGAFSDPPISLNRAC 1174
Score = 217 bits (552), Expect(2) = e-176
Identities = 106/120 (88%), Positives = 113/120 (93%), Gaps = 1/120 (0%)
Frame = +3
Query: 1 MAKIELSRHIPLVTLIVLVLCI-TPPIFATKNCDFPAIFSFGASNVDTGGLAAAFRAPPS 59
M K EL RH+ LV+LIVL+LCI TPPIFAT+NCDFPAIFSFGASNVDTGGLAAAF+APPS
Sbjct: 33 MEKFELLRHMSLVSLIVLILCIITPPIFATRNCDFPAIFSFGASNVDTGGLAAAFQAPPS 212
Query: 60 PYGETYFHRSTGRFSDGRIILDFIARSFRLPYLSPYLNSLGSNFTHGANFASGGSTINIP 119
PYGETYFHRSTGRFSDGRIILDFIA+SF LPYLSPYLNSLGSNFTHGANFA+GGSTIN P
Sbjct: 213 PYGETYFHRSTGRFSDGRIILDFIAQSFGLPYLSPYLNSLGSNFTHGANFATGGSTINSP 392
>NP429751 NP429751|AF463407.1|AAL68830.1 Enod8.3 [Medicago truncatula]
Length = 902
Score = 500 bits (1288), Expect = e-142
Identities = 241/299 (80%), Positives = 259/299 (86%), Gaps = 2/299 (0%)
Frame = +3
Query: 85 RSFRLPYLSPYLNSLGSNFTHGANFASGGSTINIPKSILPNGKLSPFSLQIQYIQFKEFI 144
+SF LPYLSPYLNSLGSNFTHGANFA+ GSTI IP SI+PNG SPFSLQIQ IQFK+FI
Sbjct: 3 QSFGLPYLSPYLNSLGSNFTHGANFATAGSTIKIPNSIIPNGMFSPFSLQIQSIQFKDFI 182
Query: 145 SKTKLIRDQGGVFATLIPKEDYFSKALYIFDIGQNDLTIGFFGNKTIQQVNATVPDIVNN 204
K K IRDQGGVFATLIPKEDY+SKALY FDIGQNDLT GFFGNKTIQQVN TVPDIV +
Sbjct: 183 PKAKFIRDQGGVFATLIPKEDYYSKALYTFDIGQNDLTAGFFGNKTIQQVNTTVPDIVKS 362
Query: 205 YIENIKNIYNLGARSFWIHGTGPKGCAPVILANFPSAIKDSYGCAKQYNEVSQYFNFKLK 264
+I+NIKNIYNLGARSFWIH TGP GC P+ILANFPSAIKD YGCAKQYNEVSQYFN KLK
Sbjct: 363 FIDNIKNIYNLGARSFWIHNTGPIGCVPLILANFPSAIKDRYGCAKQYNEVSQYFNLKLK 542
Query: 265 EALAELRSNLSSAAITYVDIYTPKYSLFTNPEKYGFELPFVACCGYGGE--YNIGVGCGA 322
EALA+LR +L AAITYVDIY+PKYSLF NP+KYGFELP VACCG GG+ YNI GCGA
Sbjct: 543 EALAQLRKDLPLAAITYVDIYSPKYSLFQNPKKYGFELPLVACCGNGGKYNYNIRAGCGA 722
Query: 323 SININGTKIVAGSCKNPSTRIIWDGVHYTEAANEIVFSQILTGVFNDPPISLDRACYRK 381
+ININGT V GSCK PSTRIIWDG HYTEAAN+IVF QI G F DPPI L+RACY+K
Sbjct: 723 TININGTNTVVGSCKKPSTRIIWDGTHYTEAANKIVFDQISNGAFTDPPIPLNRACYKK 899
>TC81963 Enod8-like protein [Medicago truncatula]
Length = 783
Score = 303 bits (777), Expect = 6e-83
Identities = 148/227 (65%), Positives = 180/227 (79%), Gaps = 3/227 (1%)
Frame = +3
Query: 9 HIPLVTL-IVLVLCITPPIFATKN--CDFPAIFSFGASNVDTGGLAAAFRAPPSPYGETY 65
HI VT +VL + T I ++ N C+F AIF+FG SN DTGGLAA+F AP PYGETY
Sbjct: 99 HISSVTFFVVLSIATTTVIESSSNSECNFRAIFNFGDSNSDTGGLAASFVAPKPPYGETY 278
Query: 66 FHRSTGRFSDGRIILDFIARSFRLPYLSPYLNSLGSNFTHGANFASGGSTINIPKSILPN 125
FHR GRFSDGR+I+DFIA+SF LPYLS YL+SLG+NF+HGANFA+ STI P SI+P
Sbjct: 279 FHRPNGRFSDGRLIVDFIAQSFGLPYLSAYLDSLGTNFSHGANFATTSSTIRPPPSIIPQ 458
Query: 126 GKLSPFSLQIQYIQFKEFISKTKLIRDQGGVFATLIPKEDYFSKALYIFDIGQNDLTIGF 185
G SPF L +QY QF++F +T+ IR QGG+FA+L+PKE+YFSKALY FDIGQNDL GF
Sbjct: 459 GGFSPFYLDVQYTQFRDFKPRTQFIRQQGGLFASLMPKEEYFSKALYTFDIGQNDLGAGF 638
Query: 186 FGNKTIQQVNATVPDIVNNYIENIKNIYNLGARSFWIHGTGPKGCAP 232
FGN TIQQVNA+VP+I+N++ +N+K+IYNLG RSFWIH TGP GC P
Sbjct: 639 FGNMTIQQVNASVPEIINSFSKNVKDIYNLGGRSFWIHNTGPIGCLP 779
>TC82515 similar to GP|3688284|emb|CAA09694.1 lanatoside
15'-O-acetylesterase {Digitalis lanata}, partial (76%)
Length = 1230
Score = 268 bits (684), Expect = 3e-72
Identities = 148/359 (41%), Positives = 201/359 (55%), Gaps = 12/359 (3%)
Frame = +1
Query: 32 CDFPAIFSFGASNVDTGGLAAAFRAPPSPYGETYFHRSTGRFSDGRIILDFIARSFRLPY 91
CDF IF+FG SN DTGG +AF A P PYG TYF GR SDGR+I+DF+A + LPY
Sbjct: 133 CDFQGIFNFGDSNSDTGGFYSAFPAQPIPYGMTYFKTPVGRSSDGRLIVDFLAEALGLPY 312
Query: 92 LSPYLNSLGSNFTHGANFASGGSTINIPKSILPNGKLSPFSLQIQYIQFKEFISKTKLIR 151
LSPYL S+GS++THGANFA+ ST+ +P + L LSPF+LQIQ Q ++F +K
Sbjct: 313 LSPYLQSIGSDYTHGANFATSASTVLLPTTSLFVSGLSPFALQIQLRQMQQFRAKVHDFH 492
Query: 152 DQGGVFATL------IPKEDYFSKALYIFDIGQNDLTIGFFGNKTIQQVNATVPDIVNNY 205
+ + + IP D F K++Y+F IGQND T + I + +P I+
Sbjct: 493 KRDPLKPSTCASKIKIPSPDIFGKSIYMFYIGQNDFTSKIAASGGINGLKNYLPQIIYQI 672
Query: 206 IENIKNIYNL-GARSFWIHGTGPKGCAPVILANFPSAIKD--SYGCAKQYNEVSQYFNFK 262
IK +Y G R+F + GP GC P L P D +GC YN +N
Sbjct: 673 ASAIKELYYAQGGRTFMVLNLGPVGCYPGYLVELPHTSSDLNEHGCIITYNNAVDDYNKL 852
Query: 263 LKEALAELRSNLSSAAITYVDIYTPKYSLFTNPEKYGFELPFVACCGY-GGEYNIG--VG 319
LKE L + R +LS A++ YVD + LF +P YG + ACCG+ GG+YN
Sbjct: 853 LKETLTQTRKSLSDASLIYVDTNSALMELFRHPTSYGLKHSTKACCGHGGGDYNFDPKAL 1032
Query: 320 CGASININGTKIVAGSCKNPSTRIIWDGVHYTEAANEIVFSQILTGVFNDPPISLDRAC 378
CG ++A +C++P + WDG+H+TEAAN+I+ IL G +DPP L + C
Sbjct: 1033CG--------NMLASACEDPQNYVSWDGIHFTEAANKIIAMAILNGSLSDPPFLLHKLC 1185
>TC84309 similar to GP|9294302|dbj|BAB02204.1 nodulin-like protein protein
{Arabidopsis thaliana}, partial (47%)
Length = 672
Score = 218 bits (556), Expect = 2e-57
Identities = 101/202 (50%), Positives = 145/202 (71%)
Frame = +2
Query: 12 LVTLIVLVLCITPPIFATKNCDFPAIFSFGASNVDTGGLAAAFRAPPSPYGETYFHRSTG 71
+++ ++V+ + P+ +NC FPAIF+FG SN DTGGL+AAF P P G T+F G
Sbjct: 59 VISTFLVVVMVPSPVIGARNCSFPAIFNFGDSNSDTGGLSAAFGQAPPPNGITFFQTPAG 238
Query: 72 RFSDGRIILDFIARSFRLPYLSPYLNSLGSNFTHGANFASGGSTINIPKSILPNGKLSPF 131
RFSDGR+I+DF+A++ LPYLS YL+S+GSNF++GANFA+ GSTI + SP
Sbjct: 239 RFSDGRLIIDFLAQNLSLPYLSAYLDSVGSNFSNGANFATAGSTIRPQNTTKSQSGYSPI 418
Query: 132 SLQIQYIQFKEFISKTKLIRDQGGVFATLIPKEDYFSKALYIFDIGQNDLTIGFFGNKTI 191
SL +Q IQ+ +F +++ L+R +GGVF L+PKE+YFS+ALY FDIGQNDLT G+ N T
Sbjct: 419 SLDVQLIQYSDFKARSILVRKKGGVFMKLLPKEEYFSEALYTFDIGQNDLTAGYKLNMTT 598
Query: 192 QQVNATVPDIVNNYIENIKNIY 213
+QV A +PD++ + + I+++Y
Sbjct: 599 EQVKAYIPDVLGQFSDVIRSVY 664
>TC90006 similar to PIR|A96590|A96590 hypothetical protein T22H22.20
[imported] - Arabidopsis thaliana, partial (63%)
Length = 858
Score = 197 bits (502), Expect = 4e-51
Identities = 116/268 (43%), Positives = 161/268 (59%), Gaps = 11/268 (4%)
Frame = +3
Query: 19 VLCITPPIFATK-----NCDFPAIFSFGASNVDTGGLAAA-FRAPPSPYGETYFHRSTGR 72
++CI IF + DFPA+F+FG SN DTG L A F + P G TYFH +GR
Sbjct: 57 IICIITTIFLLPCAKSIHLDFPAVFNFGDSNSDTGTLVTAGFESLYPPNGHTYFHLPSGR 236
Query: 73 FSDGRIILDFIARSFRLPYLSPYLNSLG-SNFTHGANFASGGSTINIPKSILPNGKLSPF 131
+SDGR+I+DF+ + LP+L+ YL+SLG NF G NFA+ GSTI +P + + PF
Sbjct: 237 YSDGRLIIDFLMDALDLPFLNAYLDSLGLPNFRKGCNFAAAGSTI-LPATA---SSICPF 404
Query: 132 SLQIQYIQFKEFISKT-KLIRDQGGVFATLIPKEDYFSKALYIFDIGQNDLTIGFFGNKT 190
S IQ QF +F ++ +L+ +G F +P ED F K LY+FDIGQNDL G F +KT
Sbjct: 405 SFGIQVSQFLKFKARALELLSGKGRKFDKYVPSEDIFEKGLYMFDIGQNDLA-GAFYSKT 581
Query: 191 IQQVNATVPDIVNNYIENIKNIYNLGARSFWIHGTGPKGCAPVILANF---PSAIKDSYG 247
+ QV A++P I+ + IK +Y+ GAR FWIH TGP GC +A F PS + D G
Sbjct: 582 LDQVLASIPTILLEFESGIKRLYDEGARYFWIHNTGPLGCLAQNVAKFGTDPSKL-DELG 758
Query: 248 CAKQYNEVSQYFNFKLKEALAELRSNLS 275
C +N+ + FN +L ++L ++S
Sbjct: 759 CVSGHNQAVKTFNLQLHALCSKLAGSIS 842
>TC83147 similar to PIR|T48618|T48618 early nodule-specific protein-like -
Arabidopsis thaliana, partial (43%)
Length = 653
Score = 173 bits (439), Expect(2) = 3e-45
Identities = 84/160 (52%), Positives = 115/160 (71%), Gaps = 2/160 (1%)
Frame = +1
Query: 28 ATKNCDFPAIFSFGASNVDTGGLAAAFRAPPSPYGETYFHRSTGRFSDGRIILDFIARSF 87
++ +C FPAI++FG SN DTGG++AAF PSPYG+ +FH+ GR SDGR+ILD+IA
Sbjct: 70 SSPSCSFPAIYNFGDSNSDTGGISAAFVPIPSPYGQGFFHKPFGRDSDGRVILDYIADKL 249
Query: 88 RLPYLSPYLNSLGSNFTHGANFASGGSTINIPKSILPNGKLSPFSLQIQYIQFKEFISKT 147
+ PYLS YLNSLG+N+ HGANFA+GGSTI + +SPFSL IQ +QF +F ++T
Sbjct: 250 KWPYLSAYLNSLGTNYRHGANFATGGSTIRKQNETIFQYGISPFSLDIQIVQFNQFKART 429
Query: 148 KLIRDQ--GGVFATLIPKEDYFSKALYIFDIGQNDLTIGF 185
K + + + + +P + F+KALY FDIGQNDL++GF
Sbjct: 430 KQLYQEANNSLERSKLPVPEEFAKALYTFDIGQNDLSVGF 549
Score = 26.2 bits (56), Expect(2) = 3e-45
Identities = 10/27 (37%), Positives = 14/27 (51%)
Frame = +2
Query: 187 GNKTIQQVNATVPDIVNNYIENIKNIY 213
G Q+ T+PDIVN ++ IY
Sbjct: 551 GKMNFDQIRETMPDIVNQLASAVQKIY 631
>BG587016 similar to PIR|T09416|T09 coil protein PO22
microspore/pollen-specific - alfalfa, partial (55%)
Length = 696
Score = 177 bits (449), Expect = 6e-45
Identities = 96/200 (48%), Positives = 129/200 (64%), Gaps = 6/200 (3%)
Frame = +3
Query: 30 KNCDFPAIFSFGASNVDTGGLAAAFRAPPSPYGETYFHRSTGRFSDGRIILDFIARSFRL 89
K C++PAI++FG SN DTG A + A P G +YF +TGR SDGR+I+DFI+ +L
Sbjct: 114 KKCEYPAIYNFGDSNSDTGAANAIYTAVTPPNGISYFGSTTGRASDGRLIIDFISEELKL 293
Query: 90 PYLSPYLNSLGSNFTHGANFASGGSTINIPKSILPNGKLSPFSLQIQYIQFKEFISKTKL 149
PYLS YLNS+GSN+ HGANFA GG+ SI P G SP L +Q QF F S TK+
Sbjct: 294 PYLSAYLNSIGSNYRHGANFAVGGA------SIRPGG-YSPIFLGLQVSQFILFKSHTKI 452
Query: 150 IRDQ------GGVFATLIPKEDYFSKALYIFDIGQNDLTIGFFGNKTIQQVNATVPDIVN 203
+ +Q F + +P+ + FSKALY DIGQNDL IG N + +QV ++PDI++
Sbjct: 453 LFNQLSDNRTESPFKSGLPRNEEFSKALYTIDIGQNDLAIG-LQNTSEEQVKRSIPDILS 629
Query: 204 NYIENIKNIYNLGARSFWIH 223
+ + ++ +YN GAR FWIH
Sbjct: 630 QFSQAVQQLYNEGARVFWIH 689
>TC83671 similar to PIR|T09416|T09416 coil protein PO22
microspore/pollen-specific - alfalfa, partial (60%)
Length = 671
Score = 173 bits (438), Expect = 1e-43
Identities = 96/211 (45%), Positives = 132/211 (62%), Gaps = 6/211 (2%)
Frame = +3
Query: 28 ATKNCDFPAIFSFGASNVDTGGLAAAFRAPPSPYGETYFHRSTGRFSDGRIILDFIARSF 87
++ C +PAI++FG SN DTG + A + + P G T+F +GR SDGR+I+D I
Sbjct: 39 SSDKCVYPAIYNFGDSNSDTGTIYATYTSVQPPNGITFFGNISGRASDGRLIIDSITEEL 218
Query: 88 RLPYLSPYLNSLGSNFTHGANFASGGSTINIPKSILPNGKLSPFSLQIQYIQFKEFISKT 147
+LPYLS YLNS+GSN+ HGANFA G+ SI P G F+L +Q QF F S T
Sbjct: 219 KLPYLSAYLNSVGSNYRHGANFAVSGA------SIRPRG-YHLFNLGLQVSQFILFKSHT 377
Query: 148 KLIRDQGGVFATL------IPKEDYFSKALYIFDIGQNDLTIGFFGNKTIQQVNATVPDI 201
K++ +Q T +P+ + FSKALY DIGQNDL G F + +QV ++P+I
Sbjct: 378 KILFNQLSNNRTEPSLKSGLPRPEDFSKALYTIDIGQNDLAHG-FQYTSEEQVQRSIPEI 554
Query: 202 VNNYIENIKNIYNLGARSFWIHGTGPKGCAP 232
++N+ +++K +YN GAR FWIH TGP GC P
Sbjct: 555 LSNFSQSVKQLYNEGARVFWIHNTGPIGCLP 647
>BI270789 similar to PIR|T09416|T09 coil protein PO22
microspore/pollen-specific - alfalfa, partial (62%)
Length = 657
Score = 143 bits (361), Expect(2) = 2e-36
Identities = 79/190 (41%), Positives = 114/190 (59%), Gaps = 2/190 (1%)
Frame = +1
Query: 48 GGLAAAFRAPPSPYGETYFHRSTGRFSDGRIILDFIARSFRLPYLSPYLNSLGSNFTHGA 107
G A F P G + F +GR SDGR+I+D+I ++PYLS YLNS+GSN+ +GA
Sbjct: 76 GTAYATFLCNQPPNGIS-FGNISGRASDGRLIIDYITEELKVPYLSAYLNSVGSNYRYGA 252
Query: 108 NFASGGSTINIPKSILPNGKLSPFSLQIQYIQFKEFISKTKLIRDQG--GVFATLIPKED 165
NFA+GG+ SI P SPF L +Q QF +F S T+++ + G + +P+ +
Sbjct: 253 NFAAGGA------SIRPGSGFSPFHLGLQVDQFIQFKSHTRILFNNGTEPSLKSGLPRPE 414
Query: 166 YFSKALYIFDIGQNDLTIGFFGNKTIQQVNATVPDIVNNYIENIKNIYNLGARSFWIHGT 225
F ALY DIG NDL GF + + +QV + P+I+ ++ + +K +YN+ AR FWIH
Sbjct: 415 DFCTALYTIDIGLNDLASGFL-HASEEQVQMSFPEILGHFSKAVKQLYNVXARVFWIHNV 591
Query: 226 GPKGCAPVIL 235
GP GC P I+
Sbjct: 592 GPVGCCPSII 621
Score = 26.9 bits (58), Expect(2) = 2e-36
Identities = 10/20 (50%), Positives = 15/20 (75%)
Frame = +3
Query: 28 ATKNCDFPAIFSFGASNVDT 47
++ C +PAI++FG SN DT
Sbjct: 15 SSHECVYPAIYNFGDSNSDT 74
>TC90972 similar to GP|10638955|emb|CAB81548. putative proline-rich protein
APG isolog {Cicer arietinum}, partial (88%)
Length = 1258
Score = 147 bits (371), Expect = 7e-36
Identities = 112/343 (32%), Positives = 160/343 (45%), Gaps = 14/343 (4%)
Frame = +3
Query: 35 PAIFSFGASNVDTGG---LAAAFRAPPSPYGETYF-HRSTGRFSDGRIILDFIA-----R 85
PAI +FG S VD G L F+A PYG + H+ TGRF +G++ D A +
Sbjct: 105 PAIVTFGDSAVDVGNNDYLFTLFKANYPPYGRDFVXHKPTGRFCNGKLATDITAETLGFK 284
Query: 86 SFRLPYLSPYLNSLGSNFTHGANFASGGSTINIPKSILPNGKLSPFSLQIQYIQFKEFIS 145
S+ YLSP + G N GANFAS S + +IL + P S Q++Y +KE+
Sbjct: 285 SYAPAYLSP--QATGKNLLIGANFASAASGYDEKAAILNHA--IPLSQQLKY--YKEY-- 440
Query: 146 KTKLIRDQGGVFATLIPKEDYFSKALYIFDIGQNDLTIGFFGNKTIQQVNATVPD----- 200
++KL + G A I K ALY+ G +D ++ N I +V PD
Sbjct: 441 QSKLSKIAGSKKAASIIKG-----ALYLLSGGSSDFIQNYYVNPLINKV--VTPDQYSAY 599
Query: 201 IVNNYIENIKNIYNLGARSFWIHGTGPKGCAPVILANFPSAIKDSYGCAKQYNEVSQYFN 260
+V+ Y +K++Y LGAR + P GC P F K GC + N +Q FN
Sbjct: 600 LVDTYSSFVKDLYKLGARKIGVTSLPPLGCLPATRTLFGFHEK---GCVTRINNDAQGFN 770
Query: 261 FKLKEALAELRSNLSSAAITYVDIYTPKYSLFTNPEKYGFELPFVACCGYGGEYNIGVGC 320
K+ A +L+ L I +IY P Y L +P K+GF CCG G + C
Sbjct: 771 KKINSATVKLQKQLPGLKIVVFNIYKPLYELVQSPSKFGFAEARKGCCGTGIVETTSLLC 950
Query: 321 GASININGTKIVAGSCKNPSTRIIWDGVHYTEAANEIVFSQIL 363
G+C N + + WD VH +EAAN+I+ ++
Sbjct: 951 NQK--------SLGTCSNATQYVFWDSVHPSEAANQILADALI 1055
>TC76727 similar to GP|10638955|emb|CAB81548. putative proline-rich protein
APG isolog {Cicer arietinum}, partial (92%)
Length = 1919
Score = 146 bits (369), Expect = 1e-35
Identities = 115/364 (31%), Positives = 167/364 (45%), Gaps = 14/364 (3%)
Frame = -2
Query: 14 TLIVLVLC--ITPPIFATKNCDFPAIFSFGASNVDTGG---LAAAFRAPPSPYGETYFHR 68
TL++L++ +T FA PAI +FG S VD G L F+A PYG + ++
Sbjct: 1156 TLVLLIVSCFLTCGSFAQDTL-VPAIMTFGDSAVDVGNNDYLPTLFKANYPPYGRDFTNK 980
Query: 69 S-TGRFSDGRIILDFIAR-----SFRLPYLSPYLNSLGSNFTHGANFASGGSTINIPKSI 122
TGRF +G++ DF A SF YLSP + G N GANFAS S + +
Sbjct: 979 QPTGRFCNGKLATDFTAETLGFTSFAPAYLSPQAS--GKNLLLGANFASAASGYDEKAAT 806
Query: 123 LPNGKLSPFSLQIQYIQFKEFISKTKLIRDQGGVFATLIPKEDYFSKALYIFDIGQNDLT 182
L + P S Q++Y FKE+ + KL + G A I K+ +LY+ G +D
Sbjct: 805 LNHA--IPLSQQLEY--FKEY--QGKLAQVAGSKKAASIIKD-----SLYVLSAGSSDFV 659
Query: 183 IGFFGNKTIQQ---VNATVPDIVNNYIENIKNIYNLGARSFWIHGTGPKGCAPVILANFP 239
++ N I Q V+ +++++ IK +Y LGAR + P GC P F
Sbjct: 658 QNYYTNPWINQAITVDQYSSYLLDSFTNFIKGVYGLGARKIGVTSLPPLGCLPAARTLFG 479
Query: 240 SAIKDSYGCAKQYNEVSQYFNFKLKEALAELRSNLSSAAITYVDIYTPKYSLFTNPEKYG 299
GC + N +Q FN K+ A + L+ L I DIY P Y L NP +G
Sbjct: 478 Y---HENGCVARINTDAQGFNKKVSSAASNLQKQLPGLKIVIFDIYKPLYDLVQNPSNFG 308
Query: 300 FELPFVACCGYGGEYNIGVGCGASININGTKIVAGSCKNPSTRIIWDGVHYTEAANEIVF 359
F CCG G + C G+C N + + WD VH +EAAN+++
Sbjct: 307 FAEAGKGCCGTGLVETTSLLCNPK--------SLGTCSNATQYVFWDSVHPSEAANQVLA 152
Query: 360 SQIL 363
++
Sbjct: 151 DNLI 140
>TC90156 similar to PIR|B86227|B86227 hypothetical protein [imported] -
Arabidopsis thaliana, partial (84%)
Length = 1421
Score = 145 bits (365), Expect = 3e-35
Identities = 76/207 (36%), Positives = 109/207 (51%), Gaps = 3/207 (1%)
Frame = +1
Query: 176 IGQNDLTIGFFGNKTIQQVNATVPDIVNNYIENIKNIYNLGARSFWIHGTGPKGCAPVIL 235
+G+ L I N + QV +P ++ +K++YN G R FW+H TGP GC P ++
Sbjct: 535 LGKMTLLIHLPKNLSYVQVIKRIPTVITEIENAVKSLYNEGGRKFWVHNTGPFGCLPKLI 714
Query: 236 ANFPSAIKDSYGCAKQYNEVSQYFNFKLKEALAELRSNLSSAAITYVDIYTPKYSLFTNP 295
A DS+GC YN ++ FN L + +LR+ L A + YVDIY K L TN
Sbjct: 715 ALSXKKDLDSFGCLSSYNSAARLFNEALYHSSQKLRTELKDATLVYVDIYAIKNDLITNA 894
Query: 296 EKYGFELPFVACCGYGG---EYNIGVGCGASININGTKIVAGSCKNPSTRIIWDGVHYTE 352
KYGF P + CCG+GG ++ V CG G ++ C S + WDG+HYTE
Sbjct: 895 TKYGFTNPLMVCCGFGGPPYNFDARVTCGQP----GYQV----CDEGSRYVSWDGIHYTE 1050
Query: 353 AANEIVFSQILTGVFNDPPISLDRACY 379
AAN + S+IL+ ++ P I C+
Sbjct: 1051AANTWIASKILSTAYSTPRIPFGFFCH 1131
Score = 120 bits (301), Expect = 9e-28
Identities = 75/177 (42%), Positives = 105/177 (58%), Gaps = 4/177 (2%)
Frame = +3
Query: 13 VTLIVLVLCITPPIFATKNCDF-PAI-FSFGASNVDTGGLAAAFRAPPS-PYGETYFHRS 69
+T+ + ++ + + C PA+ F FG SN DTGGL + P + P G T+FHRS
Sbjct: 66 MTICIFFSSVSLALSVSSGCSSKPAVVFVFGDSNSDTGGLVSGLGFPVNLPNGRTFFHRS 245
Query: 70 TGRFSDGRIILDFIARSFRLPYLSPYLNSL-GSNFTHGANFASGGSTINIPKSILPNGKL 128
TGR SDGR+++DF+ +S +L+PYL+S+ GS FT+GANFA GS S LP K
Sbjct: 246 TGRLSDGRLVIDFLCQSLNTRFLTPYLDSMSGSTFTNGANFAVVGS------STLP--KY 401
Query: 129 SPFSLQIQYIQFKEFISKTKLIRDQGGVFATLIPKEDYFSKALYIFDIGQNDLTIGF 185
PFSL IQ +QF+ F +++ + G A + + F ALY+ DIGQNDL F
Sbjct: 402 LPFSLNIQVMQFQHFKARSLQLATSG---AKNMINDQGFRDALYLIDIGQNDLADSF 563
>TC86678 similar to PIR|E96579|E96579 hypothetical protein T18A20.15
[imported] - Arabidopsis thaliana, partial (43%)
Length = 1341
Score = 140 bits (354), Expect = 6e-34
Identities = 113/374 (30%), Positives = 168/374 (44%), Gaps = 17/374 (4%)
Frame = +2
Query: 16 IVLVLCITPPIFATKNCDF---PAIFSFGASNVDTGG-----LAAAFRAPPSPYGETYFH 67
+ L++ I F +K + A+F FG S +D G +A PYGETYF+
Sbjct: 113 VFLIIAIISQTFGSKTDYYRSNKALFIFGDSFLDAGNNNYINTTTFDQANFLPYGETYFN 292
Query: 68 RSTGRFSDGRIILDFIARSFRLPYLSPYLNSLGSNFTHGANFASGGSTINIPKSILPNGK 127
TGRFSDGR+I DFIA +P + P+L + + +G NFASGG+ + G
Sbjct: 293 FPTGRFSDGRLISDFIAEYVNIPLVPPFLQPDNNKYYNGVNFASGGAGALVETF---QGS 463
Query: 128 LSPFSLQIQYIQFKEFISKTKLIRDQGGVFATLIPKEDYFSKALYIFDIGQNDLTIGFFG 187
+ PF + Q I FK+ T +R + G + S A+Y+F IG ND F
Sbjct: 464 VIPF--KTQAINFKKV---TTWLRHKLG----SSDSKTLLSNAVYMFSIGSNDYLSPFLT 616
Query: 188 NKTI---QQVNATVPDIVNNYIENIKNIYNLGARSFWIHGTGPKGCAPVILANFPSAIKD 244
N + V ++ N+ IK I+ GA+ F I P GC P + I
Sbjct: 617 NSDVLKHYSHTEYVAMVIGNFTSTIKEIHKRGAKKFVILNLPPLGCLP------GTRIIQ 778
Query: 245 SYG---CAKQYNEVSQYFNFKLKEALAELRSNLSSAAITYVDIYTPKYSLFTNPEKYGFE 301
S G C ++ + ++ N L E L EL+ L + D + + +P KYGF+
Sbjct: 779 SQGKGSCLEELSSLASIHNQALYEVLLELQKQLRGFKFSLYDFNSDLSHMINHPLKYGFK 958
Query: 302 LPFVACCGYG---GEYNIGVGCGASININGTKIVAGSCKNPSTRIIWDGVHYTEAANEIV 358
ACCG G GEY+ CG ++ C P+ + WD H TE+A + +
Sbjct: 959 EGKSACCGSGPFRGEYS----CGGKRGEKHFEL----CDKPNESVFWDSYHLTESAYKQL 1114
Query: 359 FSQILTGVFNDPPI 372
+Q+ + N I
Sbjct: 1115AAQMWSPTGNSHTI 1156
>TC88477 similar to GP|21592417|gb|AAM64368.1 lipase/hydrolase putative
{Arabidopsis thaliana}, partial (92%)
Length = 1415
Score = 125 bits (315), Expect = 2e-29
Identities = 104/378 (27%), Positives = 158/378 (41%), Gaps = 12/378 (3%)
Frame = +2
Query: 10 IPLVTLIVLVLCITPPIFATKNCDFPAIFSFGASNVDTG---GLAAAFRAPPSPYGETYF 66
I + L+V+VL + + A P F FG S VD G GL + RA PYG F
Sbjct: 110 INMFALLVVVLGLWSGVGADPQV--PWYFIFGDSLVDNGNNNGLQSLARADYLPYGID-F 280
Query: 67 HRSTGRFSDGRIILDFIARSFRLP-YLSPYLNSLGSNFTHGANFASGGSTINIPKSILPN 125
TGRFS+G+ +D IA Y+ PY ++ G N+AS + I
Sbjct: 281 GGPTGRFSNGKTTVDAIAELLGFDDYIPPYASASDDAILKGVNYASAAAGIREETGRQLG 460
Query: 126 GKLSPFSLQIQYIQFKEFISKTKLIRDQGGVFATLIPKEDYFSKALYIFDIGQNDLTIGF 185
+LS FS Q+Q Q + + T + SK +Y +G ND +
Sbjct: 461 ARLS-FSAQVQNYQ--------STVSQVVNILGTEDQAASHLSKCIYSIGLGSNDYLNNY 613
Query: 186 FGNKTIQQVNATVPD-----IVNNYIENIKNIYNLGARSFWIHGTGPKGCAPVILANFPS 240
F + + PD ++ +Y E ++ +YN AR + G G GC+P LA +
Sbjct: 614 FMPQFYNTHDQYTPDEYADDLIQSYTEQLRTLYNN*ARKMVLFGIGQIGCSPNELATRSA 793
Query: 241 AIKDSYGCAKQYNEVSQYFNFKLKEALAELRSNLSSAAITYVDIYTPKYSLFTNPEKYGF 300
D C ++ N +Q FN KLK + + + L + + YV+ Y + +NP YGF
Sbjct: 794 ---DGVTCVEEINSANQIFNNKLKGLVDQFNNQLPDSKVIYVNSYGIFQDIISNPSAYGF 964
Query: 301 ELPFVACCGYG---GEYNIGVGCGASININGTKIVAGSCKNPSTRIIWDGVHYTEAANEI 357
+ CCG G G++ + C+N + WD H TEA N +
Sbjct: 965 SVTNAGCCGVGRNNGQFT-------------CLPLQTPCENRREYLFWDAFHPTEAGNVV 1105
Query: 358 VFSQILTGVFNDPPISLD 375
V + + D +D
Sbjct: 1106VAQRAYSAQSPDDAYPID 1159
>TC80740 similar to GP|10177228|dbj|BAB10602. GDSL-motif
lipase/hydrolase-like protein {Arabidopsis thaliana},
partial (94%)
Length = 1167
Score = 124 bits (312), Expect = 5e-29
Identities = 109/344 (31%), Positives = 157/344 (44%), Gaps = 11/344 (3%)
Frame = +3
Query: 35 PAIFSFGASNVDT---GGLAAAFRAPPSPYGETYFHRS-TGRFSDGRIILDFIARSFRLP 90
PA+F FG S VD L ++ PYG + ++ TGRF +G++ DF A +
Sbjct: 186 PALFIFGDSVVDARNNNNLYTIVKSNFPPYGRDFNNQMPTGRFCNGKLAADFTAENLGFT 365
Query: 91 YLSP-YLN--SLGSNFTHGANFASGGSTINIPKSILPNGKLSPFSLQIQYIQFKEFISKT 147
P YLN N +GANFASG S P + L + SL+ Q +KE +
Sbjct: 366 TYPPAYLNLQEKRKNLLNGANFASGASGYFDPTAKLYHA----ISLEQQLEHYKE--CQN 527
Query: 148 KLIRDQGGVFATLIPKEDYFSKALYIFDIGQNDLTIGFFGNKTIQQVNAT--VPDIV-NN 204
L+ G A+ I S A+Y+ G +D ++ N + +V DI+ +
Sbjct: 528 ILVGVAGKSNASSI-----ISGAIYLVRAGSSDFVQNYYINPLLYKVFTADQFSDILMQH 692
Query: 205 YIENIKNIYNLGARSFWIHGTGPKGCAPVILANFPSAIKDSYGCAKQYNEVSQYFNFKLK 264
Y I+N+Y LGAR + P GC P + F S S C + N + FN KL
Sbjct: 693 YTIFIQNLYALGARKIGVTTLPPLGCLPAAITLFGS---HSNECVDRLNNDALNFNTKLN 863
Query: 265 EALAELRSNLSSAAITYVDIYTPKYSLFTNPEKYGFELPFVACCGYGGEYNIGVGCGASI 324
L+ LS+ + +DIY P + L T P + GF ACCG G SI
Sbjct: 864 TTSQNLQKELSNLTLAVLDIYQPLHDLVTKPTENGFYEARKACCGTG-------LIETSI 1022
Query: 325 NINGTKIVAGSCKNPSTRIIWDGVHYTEAANEIVFSQIL-TGVF 367
N I G+C N + + WDG H +EAAN ++ +L +G+F
Sbjct: 1023LCNKDSI--GTCANATEYVFWDGFHTSEAANNVLADDLLISGIF 1148
>TC80033 weakly similar to GP|21593518|gb|AAM65485.1 putative GDSL-motif
lipase/hydrolase {Arabidopsis thaliana}, partial (89%)
Length = 1164
Score = 124 bits (310), Expect = 8e-29
Identities = 104/365 (28%), Positives = 156/365 (42%), Gaps = 17/365 (4%)
Frame = +1
Query: 19 VLCITPPIFATKNCDFPAIFSFGASNVDTGG---LAAAFRAPPSPYGETYFH-RSTGRFS 74
+LC+ + PAI FG S+VD G + R+ PYG + ++TGRFS
Sbjct: 82 ILCLLLFHLNKVSAKVPAIIVFGDSSVDAGNNNFIPTVARSNFQPYGRDFQGGKATGRFS 261
Query: 75 DGRIILDFIARSFRLP-----YLSPYLNSLGSNFTHGANFASGGSTI-NIPKSILPNGKL 128
+GRI DFIA SF + YL P N S+F G +FAS + N +L +
Sbjct: 262 NGRIPTDFIAESFGIKESVPAYLDPKYNI--SDFATGVSFASAATGYDNATSDVL---SV 426
Query: 129 SPFSLQIQYIQFKEFISKTKLIRDQGGVFATLIPKEDYFSKALYIFDIGQNDLTIGFF-- 186
P Q++Y +K++ + T+ S+++++ +G ND ++
Sbjct: 427 IPLWKQLEY--YKDYQKNLSSYLGEAKAKETI-------SESVHLMSMGTNDFLENYYTM 579
Query: 187 ----GNKTIQQVNATVPDIVNNYIENIKNIYNLGARSFWIHGTGPKGCAPVI-LANFPSA 241
T QQ + I N+I +N+Y LGAR + G P GC P+ NF
Sbjct: 580 PGRASQYTPQQYQTFLAGIAENFI---RNLYALGARKISLGGLPPMGCLPLERTTNFMG- 747
Query: 242 IKDSYGCAKQYNEVSQYFNFKLKEALAELRSNLSSAAITYVDIYTPKYSLFTNPEKYGFE 301
GC +N ++ FN KLK +L L + + + Y + P+ YGFE
Sbjct: 748 ---QNGCVANFNNIALEFNDKLKNITTKLNQELPDMKLVFSNPYYIMLHIIKKPDLYGFE 918
Query: 302 LPFVACCGYGGEYNIGVGCGASININGTKIVAGSCKNPSTRIIWDGVHYTEAANEIVFSQ 361
VACC G + +G C SC + S + WD H TE N IV
Sbjct: 919 SASVACCA-TGMFEMGYACSRGSMF--------SCTDASKFVFWDSFHPTEKTNNIVAKY 1071
Query: 362 ILTGV 366
++ V
Sbjct: 1072VVEHV 1086
>TC91569 similar to GP|18252233|gb|AAL61949.1 putative APG protein
{Arabidopsis thaliana}, partial (60%)
Length = 1398
Score = 123 bits (308), Expect = 1e-28
Identities = 107/364 (29%), Positives = 169/364 (46%), Gaps = 16/364 (4%)
Frame = +1
Query: 16 IVLVLCITPPIFATKNCD--FPAIFSFGASNVDTGG---LAAAFRAPPSPYGETYFH-RS 69
IV ++ IT I T N + PA+ FG S+VD+G ++ ++ PYG R
Sbjct: 67 IVWLILITQMIMVTCNNENYVPAVIVFGDSSVDSGNNNMISTFLKSNFRPYGRDIDGGRP 246
Query: 70 TGRFSDGRIILDFIARSFRLPYLSP-YLNSLGS--NFTHGANFASGGSTINIPKSILPNG 126
TGRFS+GRI DFI+ +F + L P YL+ + +F G FAS G+ + S + N
Sbjct: 247 TGRFSNGRIPPDFISEAFGIKSLIPAYLDPAYTIDDFVTGVCFASAGTGYDNATSAILN- 423
Query: 127 KLSPFSLQIQYIQFKEFISKTKL-IRDQGGVFATLIPKEDYFSKALYIFDIGQNDLTIGF 185
+ P ++++ +KE+ K K I ++ + + S+ALYI +G ND +
Sbjct: 424 -VIPLWKEVEF--YKEYQDKLKAHIGEEKSI--------EIISEALYIISLGTNDFLGNY 570
Query: 186 FG------NKTIQQVNATVPDIVNNYIENIKNIYNLGARSFWIHGTGPKGCAPVILANFP 239
+G TI Q + I N+I + +Y+LGAR I G P GC P L
Sbjct: 571 YGFTTLRFRYTISQYQDYLIGIAENFI---RQLYSLGARKLAITGLIPMGCLP--LERAI 735
Query: 240 SAIKDSYGCAKQYNEVSQYFNFKLKEALAELRSNLSSAAITYVDIYTPKYSLFTNPEKYG 299
+ + C ++YN V+ FN KL+ +++L L ++Y + T P YG
Sbjct: 736 NIFGGFHRCYEKYNIVALEFNVKLENMISKLNKELPQLKALSANVYDLFNDIITRPSFYG 915
Query: 300 FELPFVACCGYGGEYNIGVGCGASININGTKIVAGSCKNPSTRIIWDGVHYTEAANEIVF 359
E ACC G ++ K+ +CK+ S + WD H TE N I+
Sbjct: 916 IEEVEKACCSTG---------TIEMSYLCNKMNLMTCKDASKYMFWDAFHPTEKTNRIIS 1068
Query: 360 SQIL 363
+ ++
Sbjct: 1069NYLI 1080
>TC79343 weakly similar to PIR|E86411|E86411 protein F1K23.18 [imported] -
Arabidopsis thaliana, partial (52%)
Length = 1420
Score = 121 bits (303), Expect = 5e-28
Identities = 74/214 (34%), Positives = 107/214 (49%), Gaps = 5/214 (2%)
Frame = +3
Query: 165 DYFSKALYIF-DIGQNDLTIGFFGNKTIQQVNATVPDIVNNYIENIKNIYNLGARSFWIH 223
+ F+ +L++ +IG ND F ++I ++ VP +++ I + +LGAR+ I
Sbjct: 594 EVFANSLFLMGEIGGNDFNYPLFIRRSIVEIKTYVPHVISAITSAINELIDLGARTLMIP 773
Query: 224 GTGPKGCAPVILANFPSAIK---DSYGCAKQYNEVSQYFNFKLKEALAELRSNLSSAAIT 280
G P GC + L + + K DS GC K NE ++++N +L+ L LR A I
Sbjct: 774 GNFPLGCNVIYLTKYETTDKSQYDSAGCLKWLNEFAEFYNQELQYELHRLRRIHPHATII 953
Query: 281 YVDIYTPKYSLFTNPEKYGFELPFVACCGYGGEYNIGVG-CGASININGTKIVAGSCKNP 339
Y D Y L+ NP K+GF CCG GG YN G G CG K +C +P
Sbjct: 954 YADYYNALLPLYQNPTKFGF-TGLKNCCGMGGSYNFGSGSCG--------KPGVFACDDP 1106
Query: 340 STRIIWDGVHYTEAANEIVFSQILTGVFNDPPIS 373
S I WDGVH TEAA ++ I+ G + P S
Sbjct: 1107SQYIGWDGVHLTEAAYRLIADGIINGPCSVPQFS 1208
Score = 84.3 bits (207), Expect = 7e-17
Identities = 48/114 (42%), Positives = 67/114 (58%), Gaps = 11/114 (9%)
Frame = +2
Query: 13 VTL-IVLVLCITPPIFATKNCDFPAIFSFGASNVDTGGLAAAFRAPPS-----PYGETYF 66
VTL ++LV+ ++ P+F + +IFSFG S DTG L + + P PYG+TYF
Sbjct: 125 VTLQLLLVITVSAPLFTAACSSYSSIFSFGDSIADTGNLYLSSQPPSDHCFFPPYGQTYF 304
Query: 67 HRSTGRFSDGRIILDFIARSFRLPYLSPYLNSLG-----SNFTHGANFASGGST 115
H +GR SDGR+I+DFIA S +P + PYL ++ GANFA G+T
Sbjct: 305 HHPSGRCSDGRLIIDFIAESLGIPMVKPYLGIKNGVLEDNSAKEGANFAVIGAT 466
Database: MTGI
Posted date: Oct 22, 2004 3:39 PM
Number of letters in database: 27,044,181
Number of sequences in database: 36,976
Lambda K H
0.322 0.141 0.428
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 12,667,578
Number of Sequences: 36976
Number of extensions: 187646
Number of successful extensions: 1037
Number of sequences better than 10.0: 97
Number of HSP's better than 10.0 without gapping: 943
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 948
length of query: 381
length of database: 9,014,727
effective HSP length: 98
effective length of query: 283
effective length of database: 5,391,079
effective search space: 1525675357
effective search space used: 1525675357
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.9 bits)
S2: 59 (27.3 bits)
Medicago: description of AC139354.10


