

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC139343.10 + phase: 0 /pseudo
(1612 letters)
Database: MTGI
36,976 sequences; 27,044,181 total letters
Searching..................................................done
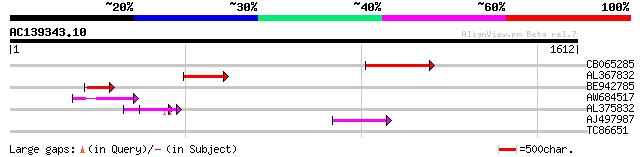
Score E
Sequences producing significant alignments: (bits) Value
CB065285 weakly similar to PIR|A84500|A84 probable retroelement ... 322 6e-88
AL367832 weakly similar to GP|14091845|gb Putative retroelement ... 198 2e-50
BE942785 160 4e-39
AW684517 similar to GP|9927273|dbj Similar to Arabidopsis thalia... 147 2e-35
AL375832 94 4e-19
AJ497987 weakly similar to GP|9927273|dbj Similar to Arabidopsis... 79 1e-14
TC86651 similar to PIR|F84555|F84555 similar to prolyl 4-hydroxy... 31 4.4
>CB065285 weakly similar to PIR|A84500|A84 probable retroelement gag/pol
polyprotein [imported] - Arabidopsis thaliana, partial
(2%)
Length = 592
Score = 322 bits (826), Expect = 6e-88
Identities = 172/195 (88%), Positives = 176/195 (90%)
Frame = -3
Query: 1013 NQKTAKNR*LMKVQIPIASGV*SLMVLSMLMVKELGQSLYPHRGITSLLPPEFCSNVQTI 1072
NQKTAKNR*L+KVQIPIA+GV*SLMVLSMLMVKEL QSLYP RGITSLLPP FCSNVQTI
Sbjct: 590 NQKTAKNR*LVKVQIPIANGV*SLMVLSMLMVKELEQSLYPRRGITSLLPPGFCSNVQTI 411
Query: 1073 WLSMKHVSLGSRKQLT*ESNTSTSMEIPRSSSIR*RVNGRPIMLS*FLIVIMRDVC*HIL 1132
WLSMKHVSLGSRKQLT*ESNTSTSMEI SS R RVNGRP M *FL MR VC* IL
Sbjct: 410 WLSMKHVSLGSRKQLT*ESNTSTSMEILHLSSTRSRVNGRPTMPI*FLTATMRGVC*PIL 231
Query: 1133 QRLSCTIFLVMRTKWLMLLLLYPPCFE*TIGMMCQ*SKCNALKDLHMCLLLGM*SIRLVK 1192
Q+LSCTIFLV RTKWLML LLYPPCFE*TIGMMCQ*SKCNALKDLHMCLLLGM*SIR VK
Sbjct: 230 QKLSCTIFLVTRTKWLMLWLLYPPCFE*TIGMMCQ*SKCNALKDLHMCLLLGM*SIRPVK 51
Query: 1193 MWLTINPGTTTSSSS 1207
MWLTINPGTTTS+SS
Sbjct: 50 MWLTINPGTTTSNSS 6
>AL367832 weakly similar to GP|14091845|gb Putative retroelement {Oryza
sativa}, partial (2%)
Length = 384
Score = 198 bits (503), Expect = 2e-50
Identities = 108/127 (85%), Positives = 112/127 (88%)
Frame = -2
Query: 495 KKRKPFSLTRKRLSSST*VPRRTSEKSRSVLF*RKGLKRRSSNSSGNIRIFLHGRMKICQ 554
KK KPFSLTRKRLS ST*VPRRTSE+SRSVL *RKGLK RSS+SS N IFLH RMKICQ
Sbjct: 383 KKGKPFSLTRKRLSLST*VPRRTSERSRSVLL*RKGLKGRSSSSSENTWIFLHARMKICQ 204
Query: 555 V*IL*LWSIGSPPSLNVLPSGRN*EGLIQIWLSRLRARFKNRLMRVSSLQLSTLNGLPIL 614
V*IL LW+IGS SLNVLPSGRN*EGLIQIWLSRLR RFK+RLMRV S QLS LNGLPIL
Sbjct: 203 V*ILKLWNIGSQQSLNVLPSGRN*EGLIQIWLSRLRVRFKSRLMRVFS*QLSILNGLPIL 24
Query: 615 YLCQRRM 621
YLCQRRM
Sbjct: 23 YLCQRRM 3
>BE942785
Length = 460
Score = 160 bits (405), Expect = 4e-39
Identities = 77/85 (90%), Positives = 78/85 (91%)
Frame = -1
Query: 214 ICKKWKACYHSWRGSIPG*PAIIFLLHRGRVCRGDRFSRINCRRYGA*EGWNCHGFFEGC 273
ICK W ACYHSWRGSI *PAIIFLLHR RVCRGDRFSRINCRRYGA EGWNC+GFFEGC
Sbjct: 385 ICK-WXACYHSWRGSILD*PAIIFLLHRSRVCRGDRFSRINCRRYGAQEGWNCNGFFEGC 209
Query: 274 TESRPRGSSCRLGQVNTAS*EQA*G 298
TESRPR SSCRLGQVNTAS*EQA*G
Sbjct: 208 TESRPRESSCRLGQVNTAS*EQA*G 134
>AW684517 similar to GP|9927273|dbj Similar to Arabidopsis thaliana
chromosome II BAC F26H6; putative retroelement pol
polyprotein, partial (1%)
Length = 488
Score = 147 bits (372), Expect = 2e-35
Identities = 88/187 (47%), Positives = 104/187 (55%)
Frame = +3
Query: 179 HVSGHGY*CLLQLSLGETMDTRRGSSDLYFAPKVEICKKWKACYHSWRGSIPG*PAIIFL 238
HVSGHGY C + LS G+T+DT R +LY P+VEIC++W
Sbjct: 3 HVSGHGYQCFV*LSPGKTLDT*RWRGNLYPTPEVEICEEW-------------------- 122
Query: 239 LHRGRVCRGDRFSRINCRRYGA*EGWNCHGFFEGCTESRPRGSSCRLGQVNTAS*EQA*G 298
+C G+ FSR+ R A EGW+C+GFFEG E G+ CRLGQVN *E A G
Sbjct: 123 -----IC*GNCFSRVVYGRCRAQEGWSCNGFFEGRPEGCSGGTGCRLGQVNPTL*E*AKG 287
Query: 299 RSWLLPNFRNFHRGIL*CWVCQRNHRRSNRI*PEACICYTRRHCERLGCH*HSLNHACF* 358
RS +LPN H *C VCQ R I A +C TRRHCERLGC * SLNHAC
Sbjct: 288 RSQILPNIGCLHGNFS*CRVCQYTC*RGGSICS*ALVCDTRRHCERLGCR*CSLNHACVX 467
Query: 359 VIYLVA* 365
+ LVA*
Sbjct: 468 XMCLVA* 488
>AL375832
Length = 535
Score = 94.0 bits (232), Expect = 4e-19
Identities = 63/127 (49%), Positives = 73/127 (56%), Gaps = 7/127 (5%)
Frame = +1
Query: 369 FFKITILSLCPKRE*YFL*GHVCFGIAVD**KICRFFHDCFFSAFSLGNNGNARKNQNIC 428
F KITILSLCPKRE*YFL*G VCFGIAV * KI F FF NGNARKN N+C
Sbjct: 154 FSKITILSLCPKRE*YFL*GRVCFGIAV**IKIVVSFTTAFFLVPFSWKNGNARKNLNVC 333
Query: 429 IFLLISKICIKKPPSKIIQSFYAD*TII-------NPLGTVILHSLRILSSQCMKPKMRK 481
IFLLISK C+KK K ++ + + II P+ L S+ M + +
Sbjct: 334 IFLLISKTCMKKKKYKKQKNSF*NHPIIICRLNHNKPVEHSNSMPLPNFDSRYMNRR*GR 513
Query: 482 VMTYHMR 488
VM Y MR
Sbjct: 514 VMIYRMR 534
Score = 86.7 bits (213), Expect = 7e-17
Identities = 65/147 (44%), Positives = 69/147 (46%), Gaps = 7/147 (4%)
Frame = +2
Query: 325 RSNRI*PEACICYTRRHCERLGCH*HSLNHACF*VIYLVA*VSCFFKITILSLCPKRE*Y 384
RSN I +ACICYTRRHCE LG H*HS NHACF V+ LVA CF K
Sbjct: 23 RSNWIWSQACICYTRRHCE*LGRH*HSFNHACFRVMRLVALALCFQK*QSSRSAQSESDI 202
Query: 385 FL*GHVCFGIAVD**KICRFFHDCFFSAFSLGNNGNARKNQNICIFLLISKIC------- 437
F G + D *K+ FF LG K K
Sbjct: 203 FCKGVFVSALPFDE*KLSFLSRLLFF*CLFLGKMVMPEKT*TSVFSY*FQKPA*KKKNTK 382
Query: 438 IKKPPSKIIQSFYAD*TIINPLGTVIL 464
KK SKIIQS YA *TIINPL TVIL
Sbjct: 383 NKKTLSKIIQSLYAG*TIINPLNTVIL 463
>AJ497987 weakly similar to GP|9927273|dbj Similar to Arabidopsis thaliana
chromosome II BAC F26H6; putative retroelement pol
polyprotein, partial (1%)
Length = 636
Score = 79.3 bits (194), Expect = 1e-14
Identities = 68/169 (40%), Positives = 83/169 (48%)
Frame = -3
Query: 917 RLVVLWPGPLNACVIIW*VIQLG*YPEWIR*SISLRKLLLLGRLHAGRCFCLNMILCSKL 976
R VVL PG +A VII + LG Y +WI+ S LR L LH GRC+ NMI +
Sbjct: 634 RHVVL*PGLPSASVII*LITLLGWYLKWIQSSTYLRSPL*QEGLHGGRCYYRNMISSTVP 455
Query: 977 KRQSKVAFLPIILLTNLLMIINQLSSISPMKRSCI*NQKTAKNR*LMKVQIPIASGV*SL 1036
+RQ K FL I LTN L I+ S MKR CI* K + KV I I GV* L
Sbjct: 454 RRQLKAVFLLITWLTNHLKTIDLSSLTFLMKRLCI*R*KIVTSHYSEKVLIQIQCGV*YL 275
Query: 1037 MVLSMLMVKELGQSLYPHRGITSLLPPEFCSNVQTIWLSMKHVSLGSRK 1085
M LSM GQ + +TSL + +T L+ K S S +
Sbjct: 274 MGLSMYTATA*GQYSLLLKVLTSLSQQDSGLTARTTSLNTKPASWASTR 128
>TC86651 similar to PIR|F84555|F84555 similar to prolyl 4-hydroxylase alpha
subunit [imported] - Arabidopsis thaliana, partial (85%)
Length = 1329
Score = 30.8 bits (68), Expect = 4.4
Identities = 15/37 (40%), Positives = 23/37 (61%)
Frame = -3
Query: 959 RLHAGRCFCLNMILCSKLKRQSKVAFLPIILLTNLLM 995
++H RC+ L ILC K + KV + +ILL+N +M
Sbjct: 727 KVHQYRCYALTPILCVKFIK--KVVIVRLILLSNFIM 623
Database: MTGI
Posted date: Oct 22, 2004 3:39 PM
Number of letters in database: 27,044,181
Number of sequences in database: 36,976
Lambda K H
0.352 0.154 0.547
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 59,426,180
Number of Sequences: 36976
Number of extensions: 1013108
Number of successful extensions: 11389
Number of sequences better than 10.0: 14
Number of HSP's better than 10.0 without gapping: 3495
Number of HSP's successfully gapped in prelim test: 490
Number of HSP's that attempted gapping in prelim test: 7626
Number of HSP's gapped (non-prelim): 4680
length of query: 1612
length of database: 9,014,727
effective HSP length: 109
effective length of query: 1503
effective length of database: 4,984,343
effective search space: 7491467529
effective search space used: 7491467529
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 14 ( 7.1 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 37 (21.5 bits)
S2: 65 (29.6 bits)
Medicago: description of AC139343.10


