

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC138453.15 + phase: 0 /pseudo/partial
(195 letters)
Database: MTGI
36,976 sequences; 27,044,181 total letters
Searching..................................................done
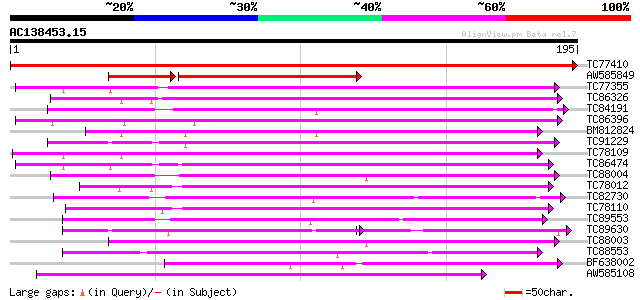
Score E
Sequences producing significant alignments: (bits) Value
TC77410 similar to GP|21392365|gb|AAM48289.1 flavanone 3 beta-hy... 353 2e-98
AW585849 similar to GP|21038956|dbj flavanone 3-hydroxylase {Mal... 132 9e-38
TC77355 weakly similar to PIR|T05552|T05552 SRG1 protein-related... 106 5e-24
TC86326 similar to GP|6016680|gb|AAF01507.1| putative leucoantho... 99 8e-22
TC84191 weakly similar to PIR|T05119|T05119 leucoanthocyanidin d... 96 9e-21
TC86396 similar to GP|10178137|dbj|BAB11549. leucoanthocyanidin ... 94 4e-20
BM812824 similar to GP|22266677|db leucoanthocyanidin dioxgenase... 93 7e-20
TC91229 similar to GP|9294689|dbj|BAB03055.1 gene_id:MHC9.10~sim... 90 5e-19
TC78109 similar to PIR|S44261|S44261 SRG1 protein - Arabidopsis ... 89 8e-19
TC86474 weakly similar to GP|6016680|gb|AAF01507.1| putative leu... 89 1e-18
TC88004 weakly similar to PIR|T04185|T04185 hypothetical protein... 88 2e-18
TC78012 similar to PIR|T05552|T05552 SRG1 protein-related protei... 86 9e-18
TC82730 similar to GP|10177853|dbj|BAB11205. flavanone 3-hydroxy... 86 1e-17
TC78110 similar to PIR|S44261|S44261 SRG1 protein - Arabidopsis ... 83 8e-17
TC89553 weakly similar to SP|P24397|HY6H_HYONI Hyoscyamine 6-dio... 82 1e-16
TC89630 similar to PIR|D86201|D86201 protein F12K11.6 [imported]... 64 2e-16
TC88003 weakly similar to PIR|T04184|T04184 hypothetical protein... 80 4e-16
TC88553 weakly similar to GP|21327037|gb|AAM48133.1 putative fla... 79 9e-16
BF638002 weakly similar to GP|20260150|gb strong similarity to n... 78 2e-15
AW585108 78 2e-15
>TC77410 similar to GP|21392365|gb|AAM48289.1 flavanone 3 beta-hydroxylase
{Solanum tuberosum}, partial (98%)
Length = 1471
Score = 353 bits (907), Expect = 2e-98
Identities = 168/195 (86%), Positives = 181/195 (92%)
Frame = +1
Query: 1 MAPVKTLKYLAQEKTLESSFIRDVSERPKVAYNNFSNEIPIISLAGIDDVDGLRIETCNK 60
MAP +TL YLAQEKTLESSF+R+ ERPKVAYNNFSNEIPIISL GIDD G R E CNK
Sbjct: 58 MAPAQTLTYLAQEKTLESSFVREEDERPKVAYNNFSNEIPIISLDGIDDAGGRRAEICNK 237
Query: 61 IVEACENWGIFQIVDHGVDQKLISEMTRLAKGFFDLPPEEKLRFDLSGGKKGGFIVSSHL 120
IVEACENWGIFQ+VDHGVD KLISEMTR AKGFFDLPPEEKLRFD+SGGKKGGFIVSSHL
Sbjct: 238 IVEACENWGIFQVVDHGVDSKLISEMTRFAKGFFDLPPEEKLRFDMSGGKKGGFIVSSHL 417
Query: 121 QGEPVRDWREMMIYFSYPIKQRDYSRWPNKPEGWKEVTEQYSEKLMSLSCKLLEVLSEAM 180
QGE V+DWRE++ YFSYPI+QRDYSRWP+KPEGWKEVTEQYSEKLM+L+CKLLEVLSEAM
Sbjct: 418 QGEAVKDWRELVTYFSYPIRQRDYSRWPDKPEGWKEVTEQYSEKLMNLACKLLEVLSEAM 597
Query: 181 GLEKEALTKACVDMD 195
GLEK+ALTKACVDMD
Sbjct: 598 GLEKDALTKACVDMD 642
>AW585849 similar to GP|21038956|dbj flavanone 3-hydroxylase {Malus x
domestica}, partial (23%)
Length = 527
Score = 132 bits (332), Expect(2) = 9e-38
Identities = 63/63 (100%), Positives = 63/63 (100%)
Frame = +3
Query: 59 NKIVEACENWGIFQIVDHGVDQKLISEMTRLAKGFFDLPPEEKLRFDLSGGKKGGFIVSS 118
NKIVEACENWGIFQIVDHGVDQKLISEMTRLAKGFFDLPPEEKLRFDLSGGKKGGFIVSS
Sbjct: 75 NKIVEACENWGIFQIVDHGVDQKLISEMTRLAKGFFDLPPEEKLRFDLSGGKKGGFIVSS 254
Query: 119 HLQ 121
HLQ
Sbjct: 255 HLQ 263
Score = 41.2 bits (95), Expect(2) = 9e-38
Identities = 21/23 (91%), Positives = 21/23 (91%)
Frame = +2
Query: 35 FSNEIPIISLAGIDDVDGLRIET 57
FSNEIPIISLAGID V GLRIET
Sbjct: 2 FSNEIPIISLAGIDYVYGLRIET 70
>TC77355 weakly similar to PIR|T05552|T05552 SRG1 protein-related protein
F24A6.150 - Arabidopsis thaliana, partial (44%)
Length = 1359
Score = 106 bits (265), Expect = 5e-24
Identities = 61/190 (32%), Positives = 102/190 (53%), Gaps = 3/190 (1%)
Frame = +1
Query: 3 PVKTLKYLAQEKTLE--SSFIRDVSERPKVAYN-NFSNEIPIISLAGIDDVDGLRIETCN 59
PV ++ + + L+ + ++R+ E KV Y S+EIP+I + + + +E
Sbjct: 82 PVPNVQEMVKVNPLQVPTKYVRNEEEMEKVNYMPQLSSEIPVIDFTQLSNGN---MEELL 252
Query: 60 KIVEACENWGIFQIVDHGVDQKLISEMTRLAKGFFDLPPEEKLRFDLSGGKKGGFIVSSH 119
K+ AC+ WG FQ+V+HGV++ LI M + FF+LP EEK ++ + G+ +
Sbjct: 253 KLEIACKEWGFFQMVNHGVEKDLIQRMRDASNEFFELPMEEKEKYAMLPNDIQGYGQNYV 432
Query: 120 LQGEPVRDWREMMIYFSYPIKQRDYSRWPNKPEGWKEVTEQYSEKLMSLSCKLLEVLSEA 179
+ E DW ++++ YP + R WP G+KE+ E YS ++ + +LL LS
Sbjct: 433 VSEEQTLDWSDVLVLIIYPDRYRKLQFWPKTSHGFKEIIEAYSSEVRRVGEELLSFLSII 612
Query: 180 MGLEKEALTK 189
MGLEK AL +
Sbjct: 613 MGLEKHALAE 642
>TC86326 similar to GP|6016680|gb|AAF01507.1| putative leucoanthocyanidin
dioxygenase {Arabidopsis thaliana}, partial (81%)
Length = 1686
Score = 99.4 bits (246), Expect = 8e-22
Identities = 55/179 (30%), Positives = 99/179 (54%), Gaps = 3/179 (1%)
Frame = +2
Query: 15 TLESSFIRDVSERPKVAYNNFSN-EIPIISLAGI--DDVDGLRIETCNKIVEACENWGIF 71
++ +I+ ++RP V +++ + IPII L G+ DD+D + +I +AC +WG F
Sbjct: 323 SIPDRYIKPPTDRPIVDTSSYDDINIPIIDLGGLNGDDLD-VHASILKQISDACRDWGFF 499
Query: 72 QIVDHGVDQKLISEMTRLAKGFFDLPPEEKLRFDLSGGKKGGFIVSSHLQGEPVRDWREM 131
QIV+HGV L+ + + FF LP E K ++ S G+ ++ + DW +
Sbjct: 500 QIVNHGVSPDLMDKARETWRQFFHLPMEAKQQYANSPTTYEGYGSRLGVEKGAILDWSDY 679
Query: 132 MIYFSYPIKQRDYSRWPNKPEGWKEVTEQYSEKLMSLSCKLLEVLSEAMGLEKEALTKA 190
P+ +D ++WP P+ +EV ++Y ++L+ LS +L++ LS +GLE++ L A
Sbjct: 680 YFLHYLPVSVKDCNKWPASPQSCREVFDEYGKELVKLSGRLMKALSLNLGLEEKILQNA 856
>TC84191 weakly similar to PIR|T05119|T05119 leucoanthocyanidin dioxygenase
(EC 1.14.11.-) F7H19.60 - Arabidopsis thaliana, partial
(18%)
Length = 718
Score = 95.9 bits (237), Expect = 9e-21
Identities = 56/181 (30%), Positives = 94/181 (50%), Gaps = 2/181 (1%)
Frame = +1
Query: 14 KTLESSFIRDVSERPKVAYNNFSNEIPIISLAGIDDVDGLRIETCNKIVEACENWGIFQI 73
+ L FIR +ERP+ +P+ISL+ ++ KI EA WG F I
Sbjct: 70 RELPPQFIRLANERPENTKAMEGVTVPMISLSQPHNL------LVKKINEAASEWGFFVI 231
Query: 74 VDHGVDQKLISEMTRLAKGFFDLPPEEKLRF--DLSGGKKGGFIVSSHLQGEPVRDWREM 131
DHG+ QKLI + + + FF LP +EK + D S GK G+ E +W +
Sbjct: 232 TDHGISQKLIQSLQDVGQEFFSLPQKEKETYANDPSSGKFDGYGTKMTKNLEQKVEWVDY 411
Query: 132 MIYFSYPIKQRDYSRWPNKPEGWKEVTEQYSEKLMSLSCKLLEVLSEAMGLEKEALTKAC 191
+ P + ++ WP P ++EV ++Y+++++ ++ +LE+LSE + LE + L K+C
Sbjct: 412 YFHLMSPHSKLNFEMWPKSPPSYREVVQEYNKEMLRVTDNILELLSEGLELESKTL-KSC 588
Query: 192 V 192
+
Sbjct: 589 L 591
>TC86396 similar to GP|10178137|dbj|BAB11549. leucoanthocyanidin
dioxygenase-like protein {Arabidopsis thaliana}, partial
(82%)
Length = 1326
Score = 93.6 bits (231), Expect = 4e-20
Identities = 56/196 (28%), Positives = 100/196 (50%), Gaps = 8/196 (4%)
Frame = +2
Query: 3 PVKTLKYLAQE--KTLESSFIRDVSERPKVAYNNFSNE-----IPIISLAGIDDVDGLRI 55
P+ ++ LA+ ++ S +I+ S+RP N+ IP+I L + D +
Sbjct: 125 PIVRVQALAESGLTSIPSCYIKPRSQRPTKTTFATQNDHDHINIPVIDLEHLSSEDPVLR 304
Query: 56 ETCNKIV-EACENWGIFQIVDHGVDQKLISEMTRLAKGFFDLPPEEKLRFDLSGGKKGGF 114
ET K V EAC WG FQ+V+HG+ +L+ + + FF+LP E K F S G+
Sbjct: 305 ETVLKRVSEACREWGFFQVVNHGISHELMESAKEVWREFFNLPLEVKEEFANSPSTYEGY 484
Query: 115 IVSSHLQGEPVRDWREMMIYFSYPIKQRDYSRWPNKPEGWKEVTEQYSEKLMSLSCKLLE 174
++ + DW + S P R+ ++WP P +++ +Y E+++ L ++LE
Sbjct: 485 GSRLGVKKGAILDWSDYFFLHSMPPSLRNQAKWPATPSSLRKIIAEYGEEVVKLGGRMLE 664
Query: 175 VLSEAMGLEKEALTKA 190
++S +GL+++ L A
Sbjct: 665 LMSTNLGLKEDYLMNA 712
>BM812824 similar to GP|22266677|db leucoanthocyanidin dioxgenase {Vitis
labrusca x Vitis vinifera}, partial (53%)
Length = 581
Score = 92.8 bits (229), Expect = 7e-20
Identities = 53/170 (31%), Positives = 87/170 (51%), Gaps = 13/170 (7%)
Frame = +1
Query: 27 RPKVAYNNFSN----------EIPIISLAGIDDVDGLRIETCN-KIVEACENWGIFQIVD 75
RPK N N ++P I L I+ D + C K+ +A E WG+ +V+
Sbjct: 67 RPKEELANIGNIFDEEKKEGPQVPTIDLKEINSSDEIVRGKCREKLKKAAEEWGVMHLVN 246
Query: 76 HGVDQKLISEMTRLAKGFFDLPPEEKLRF--DLSGGKKGGFIVSSHLQGEPVRDWREMMI 133
HG+ LI+ + + + FF+LP EEK ++ D S GK G+ +W +
Sbjct: 247 HGISDDLINRLKKAGETFFELPVEEKEKYANDQSSGKIQGYGSKLANNASGQLEWEDYFF 426
Query: 134 YFSYPIKQRDYSRWPNKPEGWKEVTEQYSEKLMSLSCKLLEVLSEAMGLE 183
+ +P +RD S WP P + +VT +Y+++L L+ K++EVLS +GL+
Sbjct: 427 HCIFPEDKRDLSIWPKTPADYTKVTSEYAKELRVLASKIMEVLSLELGLK 576
>TC91229 similar to GP|9294689|dbj|BAB03055.1 gene_id:MHC9.10~similar to
oxidases {Arabidopsis thaliana}, partial (86%)
Length = 1198
Score = 90.1 bits (222), Expect = 5e-19
Identities = 52/179 (29%), Positives = 91/179 (50%), Gaps = 3/179 (1%)
Frame = +2
Query: 14 KTLESSFIRDVSERPKVAYNNFSNEIPIISLAGIDDVDGLRIETCN---KIVEACENWGI 70
KT+ F+RD++ERP + ++++P+I + G + E N K+ ACE WG
Sbjct: 92 KTVPQRFVRDMTERPTLLSPQ-NSDMPVIDFSKFSK--GNKEEVFNELSKLSTACEEWGF 262
Query: 71 FQIVDHGVDQKLISEMTRLAKGFFDLPPEEKLRFDLSGGKKGGFIVSSHLQGEPVRDWRE 130
FQ+++H +D L+ + +++ FF LP +EK ++ ++ G G+ + + DW
Sbjct: 263 FQVINHEIDLNLMENIEDMSREFFMLPLDEKQKYPMAPGTVQGYGQAFVFSEDQKLDWCN 442
Query: 131 MMIYFSYPIKQRDYSRWPNKPEGWKEVTEQYSEKLMSLSCKLLEVLSEAMGLEKEALTK 189
M P R+ + WP KP E E YS K+ L LL+ ++ + LE++ K
Sbjct: 443 MFALGIEPHYIRNSNLWPKKPARLSETIELYSRKIRKLCQNLLKYIALGLSLEEDVFEK 619
>TC78109 similar to PIR|S44261|S44261 SRG1 protein - Arabidopsis thaliana,
partial (48%)
Length = 1314
Score = 89.4 bits (220), Expect = 8e-19
Identities = 49/187 (26%), Positives = 102/187 (54%), Gaps = 5/187 (2%)
Frame = +3
Query: 2 APVKTLKYLAQEKTLE--SSFIRDVSERPKVAYNNFSN---EIPIISLAGIDDVDGLRIE 56
+PV ++ LA+E + ++R +RP ++ S ++P+I L+ + L
Sbjct: 66 SPVPFVQELAKEALTKVPDRYVRPHHDRPILSSTTTSTPLPQLPVIDLSKLLSSHDLNEP 245
Query: 57 TCNKIVEACENWGIFQIVDHGVDQKLISEMTRLAKGFFDLPPEEKLRFDLSGGKKGGFIV 116
K+ AC+ WG FQ+++HGV + L+ + + A+ FF+LP EEK +F + G G+
Sbjct: 246 ELKKLHYACKEWGFFQLINHGVSESLMENVKKGAEEFFNLPMEEKKKFGQTEGDVEGYGQ 425
Query: 117 SSHLQGEPVRDWREMMIYFSYPIKQRDYSRWPNKPEGWKEVTEQYSEKLMSLSCKLLEVL 176
+ + E DW +M++ F+ P +R + N P ++ E Y EK+ +L+ ++++++
Sbjct: 426 AFVVSEEQKLDWADMLVLFTLPPHKRKPHLFSNIPLPFRVDLENYCEKMRTLAIQIMDLM 605
Query: 177 SEAMGLE 183
+ ++ ++
Sbjct: 606 AHSLAVD 626
>TC86474 weakly similar to GP|6016680|gb|AAF01507.1| putative
leucoanthocyanidin dioxygenase {Arabidopsis thaliana},
partial (20%)
Length = 1395
Score = 88.6 bits (218), Expect = 1e-18
Identities = 56/188 (29%), Positives = 96/188 (50%), Gaps = 3/188 (1%)
Frame = +1
Query: 3 PVKTLKYLAQEKTLE--SSFIRDVSERPKVAYN-NFSNEIPIISLAGIDDVDGLRIETCN 59
PV ++ ++ L+ ++R E KV Y +FS+++P+I + G + E
Sbjct: 118 PVPNIQETVRKNPLKVPERYVRSEEEIEKVLYMPHFSSQVPVIDFGLLSH--GNKNELL- 288
Query: 60 KIVEACENWGIFQIVDHGVDQKLISEMTRLAKGFFDLPPEEKLRFDLSGGKKGGFIVSSH 119
K+ AC+ WG FQIV+HG++ L+ + + FFDL EEK ++ + G+ +S
Sbjct: 289 KLDIACKEWGFFQIVNHGMEIDLMQRLKDVVAEFFDLSIEEKDKYAMPPDDIQGYGHTSV 468
Query: 120 LQGEPVRDWREMMIYFSYPIKQRDYSRWPNKPEGWKEVTEQYSEKLMSLSCKLLEVLSEA 179
+ + + DW + +I YP + R WP PE K+ E YS ++ + +L+ LS
Sbjct: 469 VSEKQILDWCDQLILLVYPTRFRKPQFWPETPEKLKDTIEAYSSEIKRVGEELINSLSLI 648
Query: 180 MGLEKEAL 187
GLE+ L
Sbjct: 649 FGLEEHVL 672
>TC88004 weakly similar to PIR|T04185|T04185 hypothetical protein F7L13.80 -
Arabidopsis thaliana, partial (17%)
Length = 932
Score = 88.2 bits (217), Expect = 2e-18
Identities = 57/178 (32%), Positives = 96/178 (53%), Gaps = 3/178 (1%)
Frame = +2
Query: 15 TLESSFIRDVSERPKVAYNNFSNEIPIISLAGIDDVDGLRIETCNKIVEACENWGIFQIV 74
+L FI +RP ++ FS+ IPII L+ +D V +KI +ACE +G FQIV
Sbjct: 203 SLTPDFILPEHKRPHLSEVTFSDSIPIIDLSHLDVV--------HKISKACEEFGFFQIV 358
Query: 75 DHGVDQKLISEMTRLAKGFFDLPPEEKLRFDLSGGKKGGFIVSSHLQ---GEPVRDWREM 131
+HGV ++ ++M + FF+L PEE+ + K +V+ L+ GE V+ W E
Sbjct: 359 NHGVPDQVCTKMMKAITNFFELAPEERKHLSSTDHTKNVRLVNYLLRVDGGETVKLWSEN 538
Query: 132 MIYFSYPIKQRDYSRWPNKPEGWKEVTEQYSEKLMSLSCKLLEVLSEAMGLEKEALTK 189
++ YP+ N ++E +Y++++ SL +LL ++S +GLE++ L K
Sbjct: 539 FVHPWYPMDDIISLLPENIGTQYREAFTEYAKEISSLVRRLLSLISIGLGLEEDFLLK 712
>TC78012 similar to PIR|T05552|T05552 SRG1 protein-related protein F24A6.150
- Arabidopsis thaliana, partial (22%)
Length = 784
Score = 85.9 bits (211), Expect = 9e-18
Identities = 48/165 (29%), Positives = 87/165 (52%), Gaps = 2/165 (1%)
Frame = +2
Query: 25 SERPKVAYNNFS-NEIPIISLAGI-DDVDGLRIETCNKIVEACENWGIFQIVDHGVDQKL 82
++ P V + S ++P+I L + + DG +E K AC+ WG FQ+++HGV+ L
Sbjct: 197 NQEPSVVSSTISLPQVPVIDLKKLLSEEDGTELE---KFDHACKEWGFFQLINHGVNPLL 367
Query: 83 ISEMTRLAKGFFDLPPEEKLRFDLSGGKKGGFIVSSHLQGEPVRDWREMMIYFSYPIKQR 142
+ +L + FF+LP EEK G GF + E +W ++ ++P R
Sbjct: 368 VENFKKLVQDFFNLPVEEKKILSQKPGDMEGFGQMFVVSDEHKLEWADLFYIITHPSYMR 547
Query: 143 DYSRWPNKPEGWKEVTEQYSEKLMSLSCKLLEVLSEAMGLEKEAL 187
+ + +P+ P+ ++E E YS +L L ++E +S+A+ ++K L
Sbjct: 548 NPNLFPSIPQPFRENLEMYSLELKKLCVPIVEFMSKALKIQKNEL 682
>TC82730 similar to GP|10177853|dbj|BAB11205. flavanone 3-hydroxylase-like
protein {Arabidopsis thaliana}, partial (81%)
Length = 1406
Score = 85.5 bits (210), Expect = 1e-17
Identities = 53/177 (29%), Positives = 89/177 (49%), Gaps = 1/177 (0%)
Frame = +3
Query: 16 LESSFIRDVSERPKVAYNNFSNEIPIISLAGIDDVDGLRIETCNKIVEACENWGIFQIVD 75
L S+IR S+RP ++ + +PII L + R + +I EAC ++G FQ+V+
Sbjct: 144 LPESYIRPESDRPCLSQVSEFENVPIIDLGSHN-----RTQIVQQIGEACSSYGFFQVVN 308
Query: 76 HGVDQKLISEMTRLAKGFFDLPPEEKLR-FDLSGGKKGGFIVSSHLQGEPVRDWREMMIY 134
HGV + + + +A FF LP EEK++ + K S ++ E V +WR+ +
Sbjct: 309 HGVPLEELKKTAEVAYDFFKLPVEEKMKLYSDDPTKTMRLSTSFNVNKEEVHNWRDYLRL 488
Query: 135 FSYPIKQRDYSRWPNKPEGWKEVTEQYSEKLMSLSCKLLEVLSEAMGLEKEALTKAC 191
YP+ WP+ P +KE Y +++ L ++ E +S +G K+ L K C
Sbjct: 489 HCYPL-DNYVPEWPSNPTSFKETVANYCKEVRELGLRIEEYISVELG-PKKRLLKEC 653
>TC78110 similar to PIR|S44261|S44261 SRG1 protein - Arabidopsis thaliana,
partial (53%)
Length = 1303
Score = 82.8 bits (203), Expect = 8e-17
Identities = 45/172 (26%), Positives = 88/172 (51%), Gaps = 4/172 (2%)
Frame = +2
Query: 20 FIRDVSERPKVAYNNFSNEIPIISLAGIDDVD----GLRIETCNKIVEACENWGIFQIVD 75
++R +RP ++ E+P+I + + D GL ++ K+ AC+ WG FQ+++
Sbjct: 152 YVRPHHDRPIISTTTPLLELPVIDFSKLFSQDLTIKGLELD---KLHSACKEWGFFQLIN 322
Query: 76 HGVDQKLISEMTRLAKGFFDLPPEEKLRFDLSGGKKGGFIVSSHLQGEPVRDWREMMIYF 135
HGV L+ + AK F++LP EEK +F G G+ + + E DW +M
Sbjct: 323 HGVSTSLVENVKMGAKEFYNLPIEEKKKFSQKEGDVEGYGQAFVMSEEQKLDWADMFFMI 502
Query: 136 SYPIKQRDYSRWPNKPEGWKEVTEQYSEKLMSLSCKLLEVLSEAMGLEKEAL 187
+ P R +P P +++ E YS +L L+ ++++ ++ A+ ++ + +
Sbjct: 503 TLPSHMRKPHLFPKLPLPFRDDLETYSAELKKLAIQIIDFMANALKVDAKEI 658
>TC89553 weakly similar to SP|P24397|HY6H_HYONI Hyoscyamine 6-dioxygenase
(EC 1.14.11.11) (Hyoscyamine 6-beta- hydroxylase).
[Henbane], partial (49%)
Length = 1119
Score = 82.4 bits (202), Expect = 1e-16
Identities = 51/170 (30%), Positives = 90/170 (52%), Gaps = 3/170 (1%)
Frame = +1
Query: 19 SFIRDVSERPKVAYNNFSNEIPIISLAGIDDVDGLRIETCNKIVEACENWGIFQIVDHGV 78
SF++ + RP N S IP+I L G D T +I++A E +G FQ+++HGV
Sbjct: 70 SFVQPLECRPGKVTNPSSKTIPLIDLGGHDHA-----HTILQILKASEEYGFFQVINHGV 234
Query: 79 DQKLISEMTRLAKGFFDLPPEEKL---RFDLSGGKKGGFIVSSHLQGEPVRDWREMMIYF 135
+ L+ E + + F +PP+EK+ D +G + S + + + V+ W++ + +
Sbjct: 235 SKDLMDEAFNIFQEFHAMPPKEKISECSRDPNGINCKIYASSENYKIDAVQYWKDTLTH- 411
Query: 136 SYPIKQRDYSRWPNKPEGWKEVTEQYSEKLMSLSCKLLEVLSEAMGLEKE 185
P WP KP ++EV +Y+++L L ++LE+L E +GL+ E
Sbjct: 412 PCPPSGEFMEFWPQKPPKYREVVGKYTQELNKLGHEILEMLCEGLGLKLE 561
>TC89630 similar to PIR|D86201|D86201 protein F12K11.6 [imported] -
Arabidopsis thaliana, partial (7%)
Length = 680
Score = 63.5 bits (153), Expect(2) = 2e-16
Identities = 37/105 (35%), Positives = 59/105 (55%), Gaps = 1/105 (0%)
Frame = +3
Query: 19 SFIRDVSERPKVAYNNFSNEIPIISLAGIDDVDGL-RIETCNKIVEACENWGIFQIVDHG 77
SF ++ ++AYNN + +PII LA ID+ D + + +K+ EACE WG FQ+V+HG
Sbjct: 153 SFFHHQPDKNEIAYNN-CHVLPIIDLADIDNKDPIIHQKIIDKLKEACETWGFFQVVNHG 329
Query: 78 VDQKLISEMTRLAKGFFDLPPEEKLRFDLSGGKKGGFIVSSHLQG 122
+ ++ E+ K F + PE K + KG F +S++ G
Sbjct: 330 IPLNILEELKDGIKRFHEQDPEVKKDL-YTTDPKGPFTYNSNIGG 461
Score = 38.1 bits (87), Expect(2) = 2e-16
Identities = 24/75 (32%), Positives = 39/75 (52%), Gaps = 1/75 (1%)
Frame = +1
Query: 120 LQGEPVRDWREMMIYFSYPIKQRDYSRWPNKPEGWKEVTEQYSEKLMSLSCKLLEVLSEA 179
L P +WR+ I F P D + + P +++ +YS+++M+L L E+LSEA
Sbjct: 463 LSNSPALNWRDTFICFLAP----DTPKPEDFPLVCRDILIEYSKQMMNLGFLLFELLSEA 630
Query: 180 MGLEKEAL-TKACVD 193
+GL L CV+
Sbjct: 631 LGLNPNHLKDMGCVE 675
>TC88003 weakly similar to PIR|T04184|T04184 hypothetical protein F7L13.70 -
Arabidopsis thaliana, partial (26%)
Length = 1209
Score = 80.5 bits (197), Expect = 4e-16
Identities = 49/158 (31%), Positives = 84/158 (53%), Gaps = 3/158 (1%)
Frame = +2
Query: 35 FSNEIPIISLAGIDDVDGLRIETCNKIVEACENWGIFQIVDHGVDQKLISEMTRLAKGFF 94
+ + IPI L+ D + +E +KI +ACE +G FQIV+HGV ++ ++M + FF
Sbjct: 116 YLDSIPISDLSYCDGNNPSSLEVIHKISKACEEFGFFQIVNHGVPDQVCTKMMKAITNFF 295
Query: 95 DLPPEEKLRFDLSGGKKGGFIVSSHLQ---GEPVRDWREMMIYFSYPIKQRDYSRWPNKP 151
+L PEE+ + K + + +LQ GE V+ W E + YPI
Sbjct: 296 ELAPEEREHLSSTDNTKNVRLFNYYLQVDGGEKVKLWSECFAHPWYPIDDIIQLLPEKIG 475
Query: 152 EGWKEVTEQYSEKLMSLSCKLLEVLSEAMGLEKEALTK 189
++E +Y++++ SL +LL ++S +GLE++ L K
Sbjct: 476 TQYREAFTEYAKEVGSLVRRLLSLISIGLGLEEDCLLK 589
>TC88553 weakly similar to GP|21327037|gb|AAM48133.1 putative flavanone
3-hydroxylase {Saussurea medusa}, partial (26%)
Length = 1017
Score = 79.3 bits (194), Expect = 9e-16
Identities = 48/167 (28%), Positives = 82/167 (48%), Gaps = 2/167 (1%)
Frame = +1
Query: 19 SFIRDVSERPKVAYNNFSNEIPIISLAGIDDVDGLRIETCNKIVEACENWGIFQIVDHGV 78
++I RP FSN IP+I L+ + G R KI++A E +G FQ+++HG+
Sbjct: 205 NYIFPPESRPGNVKIPFSNSIPVIDLS--EACKGDRSNIIKKIIKAAEEFGFFQVINHGI 378
Query: 79 DQKLISEMTRLAKGFFDLPPEEKLRFDLSGGKK--GGFIVSSHLQGEPVRDWREMMIYFS 136
+ E + K F +P E K + K F S + E V WR+ + +
Sbjct: 379 SMDQMKETMSIFKEIFHMPDEYKKNLCTNDPSKPCKMFTSSFNYATEKVHLWRDSLRHPC 558
Query: 137 YPIKQRDYSRWPNKPEGWKEVTEQYSEKLMSLSCKLLEVLSEAMGLE 183
YP++Q + WP P ++E +S ++ L +L+ ++SE +GL+
Sbjct: 559 YPLEQWQH-LWPENPTTYRECVGDFSNEVKELGSRLMNLISEGLGLK 696
>BF638002 weakly similar to GP|20260150|gb strong similarity to naringenin
3-dioxygenase {Arabidopsis thaliana}, partial (40%)
Length = 652
Score = 78.2 bits (191), Expect = 2e-15
Identities = 41/144 (28%), Positives = 73/144 (50%), Gaps = 7/144 (4%)
Frame = +3
Query: 54 RIETCNKIVEACENWGIFQIVDHGVDQKLISEMTRLAKGFFD-LPPEEKLRFDLSGGKKG 112
R T I AC WG F + +HG+ L+ + FF+ P EKL++ + G
Sbjct: 222 RNSTLESISHACRGWGAFHVTNHGIPPSLLDAVRNSGLTFFNNCPMSEKLKYSCTAGTAA 401
Query: 113 G------FIVSSHLQGEPVRDWREMMIYFSYPIKQRDYSRWPNKPEGWKEVTEQYSEKLM 166
+VSS+ V DWR+ + ++P+ +R+ WP+ ++E+ YS+++
Sbjct: 402 SEGYGSRMLVSSN--DHEVLDWRDYFDHHTFPLSRRNPINWPDFTSDYREIMVNYSDEMK 575
Query: 167 SLSCKLLEVLSEAMGLEKEALTKA 190
L+ KLL ++SE++GL+ + A
Sbjct: 576 ILAQKLLALISESLGLQSSCIEDA 647
>AW585108
Length = 541
Score = 78.2 bits (191), Expect = 2e-15
Identities = 62/155 (40%), Positives = 74/155 (47%)
Frame = +2
Query: 10 LAQEKTLESSFIRDVSERPKVAYNNFSNEIPIISLAGIDDVDGLRIETCNKIVEACENWG 69
L K E S + E K +NN SNE G DDV G R+ET NK E EN G
Sbjct: 26 LRLRKDSELSPACEEDECLKAVHNNPSNETLTTLSDGTDDVGGCRVETRNKTAEVRENRG 205
Query: 70 IFQIVDHGVDQKLISEMTRLAKGFFDLPPEEKLRFDLSGGKKGGFIVSSHLQGEPVRDWR 129
D+G D K E KG D EEK D GGKKGG S+ GE +D R
Sbjct: 206 TP*AADYGADLKSTLETICPVKGPPDPLLEEKSWPDTLGGKKGGPTALSYS*GEVAKDRR 385
Query: 130 EMMIYFSYPIKQRDYSRWPNKPEGWKEVTEQYSEK 164
E I+ + + RD+ R +K EG KE E +SEK
Sbjct: 386 ESAIHPLHLTR*RDHLRRLDKLEGRKEAIE*HSEK 490
Database: MTGI
Posted date: Oct 22, 2004 3:39 PM
Number of letters in database: 27,044,181
Number of sequences in database: 36,976
Lambda K H
0.318 0.136 0.405
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 5,961,583
Number of Sequences: 36976
Number of extensions: 79638
Number of successful extensions: 470
Number of sequences better than 10.0: 136
Number of HSP's better than 10.0 without gapping: 442
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 443
length of query: 195
length of database: 9,014,727
effective HSP length: 91
effective length of query: 104
effective length of database: 5,649,911
effective search space: 587590744
effective search space used: 587590744
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.3 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.7 bits)
S2: 56 (26.2 bits)
Medicago: description of AC138453.15


