

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC138130.19 - phase: 0
(881 letters)
Database: MTGI
36,976 sequences; 27,044,181 total letters
Searching..................................................done
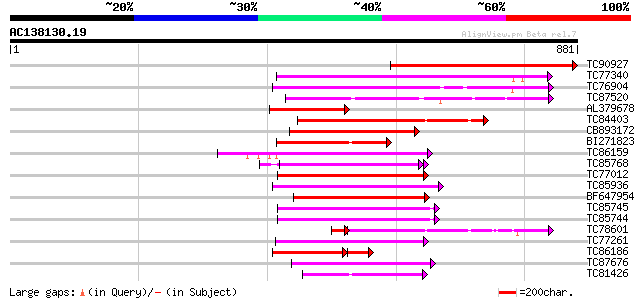
Score E
Sequences producing significant alignments: (bits) Value
TC90927 similar to GP|9279712|dbj|BAB01269.1 cell division prote... 566 e-162
TC77340 SP|Q9BAE0|FTSH_MEDSA Cell division protein ftsH homolog ... 312 4e-85
TC76904 homologue to GP|4325041|gb|AAD17230.1| FtsH-like protein... 301 7e-82
TC87520 similar to GP|9757998|dbj|BAB08420.1 cell division prote... 272 5e-73
AL379678 homologue to GP|2062173|gb| cell division protein FtsH ... 245 6e-65
TC84403 homologue to GP|15021761|gb|AAK77908.1 AAA-metalloprotea... 236 4e-62
CB893172 homologue to PIR|T45642|T45 FtsH metalloproteinase-like... 218 1e-56
BI271823 similar to GP|21592745|gb cell division protein FtsH-li... 196 2e-50
TC86159 homologue to GP|21387177|gb|AAM47992.1 26S proteasome AA... 192 4e-49
TC85768 homologue to SP|P54774|CC48_SOYBN Cell division cycle pr... 190 2e-48
TC77012 homologue to SP|P54776|PRSA_LYCES 26S protease regulator... 188 9e-48
TC85936 homologue to GP|13537115|dbj|BAB40755. AtSUG1 {Arabidops... 179 3e-45
BF647954 homologue to GP|11094192|db 26S proteasome regulatory p... 179 5e-45
TC85745 homologue to GP|17297987|dbj|BAB78491. 26S proteasome re... 177 1e-44
TC85744 homologue to PIR|T08959|T08959 proteinase homolog F19B15... 177 1e-44
TC78601 homologue to GP|15021761|gb|AAK77908.1 AAA-metalloprotea... 144 7e-42
TC77261 homologue to GP|8777330|dbj|BAA96920.1 26S proteasome AA... 167 2e-41
TC86186 similar to GP|4325041|gb|AAD17230.1| FtsH-like protein P... 142 2e-41
TC87676 PIR|G96577|G96577 26S proteasome ATPase subunit [importe... 161 9e-40
TC81426 similar to GP|6016719|gb|AAF01545.1| putative cell divis... 152 7e-37
>TC90927 similar to GP|9279712|dbj|BAB01269.1 cell division protein
FtsH-like {Arabidopsis thaliana}, partial (28%)
Length = 984
Score = 567 bits (1460), Expect = e-162
Identities = 290/290 (100%), Positives = 290/290 (100%)
Frame = +2
Query: 592 GRIEILKVHARKKPIAEDVDYEIVASMTDGMVGAELANIVEVAAINMMRDSRTEVSTDDL 651
GRIEILKVHARKKPIAEDVDYEIVASMTDGMVGAELANIVEVAAINMMRDSRTEVSTDDL
Sbjct: 8 GRIEILKVHARKKPIAEDVDYEIVASMTDGMVGAELANIVEVAAINMMRDSRTEVSTDDL 187
Query: 652 LQAAQMEERGMLDRKERSKEKWEQVAINEAAMAVAAMNLPNFDNIEYITIAPRAGRELGY 711
LQAAQMEERGMLDRKERSKEKWEQVAINEAAMAVAAMNLPNFDNIEYITIAPRAGRELGY
Sbjct: 188 LQAAQMEERGMLDRKERSKEKWEQVAINEAAMAVAAMNLPNFDNIEYITIAPRAGRELGY 367
Query: 712 VRTMLESINFNDGMLTRQSLFDHITVQLAPRAADEMWFGKDQLSTIWAETADNARVAARM 771
VRTMLESINFNDGMLTRQSLFDHITVQLAPRAADEMWFGKDQLSTIWAETADNARVAARM
Sbjct: 368 VRTMLESINFNDGMLTRQSLFDHITVQLAPRAADEMWFGKDQLSTIWAETADNARVAARM 547
Query: 772 YMIGGLSDKYRGVSNFWVTDRINEIDLEAMKILNLCYERAKEILQQNKTLMDTLVNELVV 831
YMIGGLSDKYRGVSNFWVTDRINEIDLEAMKILNLCYERAKEILQQNKTLMDTLVNELVV
Sbjct: 548 YMIGGLSDKYRGVSNFWVTDRINEIDLEAMKILNLCYERAKEILQQNKTLMDTLVNELVV 727
Query: 832 KKTLTKEDIVRLVQLHGHAKPIPISVLDIRDAKHKELQEIASNGKEINES 881
KKTLTKEDIVRLVQLHGHAKPIPISVLDIRDAKHKELQEIASNGKEINES
Sbjct: 728 KKTLTKEDIVRLVQLHGHAKPIPISVLDIRDAKHKELQEIASNGKEINES 877
>TC77340 SP|Q9BAE0|FTSH_MEDSA Cell division protein ftsH homolog chloroplast
precursor (EC 3.4.24.-). [Alfalfa] {Medicago sativa},
complete
Length = 2473
Score = 312 bits (799), Expect = 4e-85
Identities = 183/449 (40%), Positives = 270/449 (59%), Gaps = 20/449 (4%)
Frame = +3
Query: 415 VKFTDVAGLGKIRLELEEIVKFFTHGEMYRRRGVKIPGGILLCGPPGVGKTLLAKAVAGE 474
V F DVAG + +LEL+E+V F + + Y G KIP G LL GPPG GKTLLA+AVAGE
Sbjct: 999 VTFADVAGADQAKLELQEVVDFLKNPDKYTALGAKIPKGCLLVGPPGTGKTLLARAVAGE 1178
Query: 475 AGVNFFSISASQFVEIYVGVGASRVRSLYQEAKENAPSVVFIDELDAVGRKRGLIKGSGG 534
AG FFS +AS+FVE++VGVGASRVR L+++AK AP +VFIDE+DAVGR+RG G G
Sbjct: 1179 AGTPFFSCAASEFVELFVGVGASRVRDLFEKAKSKAPCIVFIDEIDAVGRQRGAGLGGGN 1358
Query: 535 QERDATLNQLLVCLDGFEGRGEVITIASTNRPDILDPALVRPGRFDRKIFIPKPGFIGRI 594
ER+ T+NQLL +DGF G VI +A+TNRPD+LD AL+RPGRFDR++ + +P GR+
Sbjct: 1359 DEREQTINQLLTEMDGFSGNSGVIVLAATNRPDVLDSALLRPGRFDRQVTVDRPDVAGRV 1538
Query: 595 EILKVHARKKPIAEDVDYEIVASMTDGMVGAELANIVEVAAINMMRDSRTEVSTDDLLQA 654
+IL+VH+R K +A+DVD++ +A T G GA+L N++ AAI R E+S D++ A
Sbjct: 1539 KILQVHSRGKALAKDVDFDKIARRTPGFTGADLQNLMNEAAILAARRDLKEISKDEIADA 1718
Query: 655 AQMEERGMLDRKER--SKEKWEQVAINEAAMAVAAMNLPNFDNIEYITIAPRAGRELGYV 712
+ G ++K S+EK + VA +EA A+ +P +D + I+I PR G+ G
Sbjct: 1719 LERIIAGP-EKKNAVVSEEKKKLVAYHEAGHALVGALMPEYDPVAKISIIPR-GQAGGLT 1892
Query: 713 RTMLESINFNDGMLTRQSLFDHITVQLAPRAADEMWFGKDQLST-IWAETADNARVAARM 771
G+ +R L + + V L R A+E+ FG+D ++T + +RVA +M
Sbjct: 1893 FFAPSEERLESGLYSRSYLENQMAVALGGRVAEEVIFGQDNVTTGASNDFMQVSRVARQM 2072
Query: 772 YMIGGLSDKY------RGVSNFWVTDRINE-----------IDLEAMKILNLCYERAKEI 814
G S K G N ++ +++ +D E ++++ YERA +I
Sbjct: 2073 VERFGFSKKIGQVAIGGGGGNPFLGQQMSSQKDYSMATADIVDKEVRELVDKAYERATQI 2252
Query: 815 LQQNKTLMDTLVNELVVKKTLTKEDIVRL 843
+ + ++ L L+ K+T+ E+ + L
Sbjct: 2253 INTHIDILHKLAQLLIEKETVDGEEFMSL 2339
>TC76904 homologue to GP|4325041|gb|AAD17230.1| FtsH-like protein Pftf
precursor {Nicotiana tabacum}, partial (91%)
Length = 2667
Score = 301 bits (771), Expect = 7e-82
Identities = 187/460 (40%), Positives = 269/460 (57%), Gaps = 24/460 (5%)
Frame = +1
Query: 409 LERGVDVKFTDVAGLGKIRLELEEIVKFFTHGEMYRRRGVKIPGGILLCGPPGVGKTLLA 468
+E V F DVAG+ + + + E+V+F E + G +IP G+LL GPPG GKTLLA
Sbjct: 802 MEPNTGVTFDDVAGVDEAKQDFMEVVEFLKKPERFTSVGARIPKGVLLVGPPGTGKTLLA 981
Query: 469 KAVAGEAGVNFFSISASQFVEIYVGVGASRVRSLYQEAKENAPSVVFIDELDAVGRKRGL 528
KA+AGEAGV FFSIS S+FVE++VG+GASRVR L+++AKENAP +VF+DE+DAVGR+RG
Sbjct: 982 KAIAGEAGVPFFSISGSEFVEMFVGIGASRVRDLFKKAKENAPCIVFVDEIDAVGRQRGT 1161
Query: 529 IKGSGGQERDATLNQLLVCLDGFEGRGEVITIASTNRPDILDPALVRPGRFDRKIFIPKP 588
G G ER+ TLNQLL +DGFEG VI +A+TNR DILD AL+RPGRFDR++ + P
Sbjct: 1162 GIGGGNDEREQTLNQLLTEMDGFEGNTGVIVVAATNRADILDSALLRPGRFDRQVSVDVP 1341
Query: 589 GFIGRIEILKVHARKKPIAEDVDYEIVASMTDGMVGAELANIVEVAAINMMRDSRTEVS- 647
GR EILKVHA K DV E++A T G GA+LAN++ AAI R RT +S
Sbjct: 1342 DVRGRTEILKVHANNKKFDNDVSLEVIAMRTPGFSGADLANLLNEAAILAGRRGRTGISP 1521
Query: 648 --TDDLLQ--AAQMEERGMLDRKERSKEKWEQVAINEAAMAVAAMNLPNFDNIEYITIAP 703
DD + A ME M D K +S VA +E A+ P D ++ +T+ P
Sbjct: 1522 KEIDDSIDRIVAGMEGTLMTDGKSKS-----LVAYHEVGHAICGTLTPGHDAVQKVTLVP 1686
Query: 704 RAGRELGYVRTMLESINFND-GMLTRQSLFDHITVQLAPRAADEMWFGKDQLST-IWAET 761
R G R + I +D ++++Q LF I L RAA+E+ FG+ +++T +
Sbjct: 1687 R-----GQARGLTWFIPSDDPTLISKQQLFARIVGGLGGRAAEEIIFGEPEVTTGAGGDL 1851
Query: 762 ADNARVAARMYMIGGLSD----------------KYRGVSNFWVTDRINE-IDLEAMKIL 804
+A +M + G+SD R ++ +++++ E ID ++
Sbjct: 1852 QQITGIARQMVVTFGMSDIGPWSLMDSSAQSGDVIMRMMARNSMSEKLAEDIDTAVKRLS 2031
Query: 805 NLCYERAKEILQQNKTLMDTLVNELVVKKTLTKEDIVRLV 844
+ YE A ++ N+ +D +V L+ K+TL+ ++ L+
Sbjct: 2032 DEAYEIALTQIRNNREAIDKIVEVLLEKETLSGDEFRALL 2151
>TC87520 similar to GP|9757998|dbj|BAB08420.1 cell division protein FtsH
protease-like {Arabidopsis thaliana}, partial (53%)
Length = 1667
Score = 272 bits (695), Expect = 5e-73
Identities = 172/427 (40%), Positives = 257/427 (59%), Gaps = 11/427 (2%)
Frame = +2
Query: 429 ELEEIVKFFTHGEMYRRRGVKIPGGILLCGPPGVGKTLLAKAVAGEAGVNFFSISASQFV 488
ELEE+V++ + + R G K+P GILL G PG GKTLLAKA+AGEAGV FF + S+F
Sbjct: 11 ELEEVVEYLKNPAKFTRLGGKLPKGILLTGAPGTGKTLLAKAIAGEAGVPFFYRAGSEFE 190
Query: 489 EIYVGVGASRVRSLYQEAKENAPSVVFIDELDAVGRKRGLIKGSGGQERDATLNQLLVCL 548
E++VGVGA RVRSL+Q AK+ AP ++FIDE+DAVG R +G TL+QLLV +
Sbjct: 191 EMFVGVGARRVRSLFQAAKKKAPCIIFIDEIDAVGSTRKQWEG----HTKKTLHQLLVEM 358
Query: 549 DGFEGRGEVITIASTNRPDILDPALVRPGRFDRKIFIPKPGFIGRIEILKVHARKKPIAE 608
DGFE +I +A+TN PDILDPAL RPGRFDR I +P P GR EIL+++ KP A+
Sbjct: 359 DGFEQNEGIILMAATNLPDILDPALTRPGRFDRHIVVPNPDVRGRQEILELYLHDKPTAD 538
Query: 609 DVDYEIVASMTDGMVGAELANIVEVAAINMMRDSRTEVSTDDLLQAAQME---ERGMLDR 665
+VD + +A T G GA+LAN+V +AAI + V + L A+Q+E +R ++
Sbjct: 539 NVDIKAIARGTPGFNGADLANLVNIAAI------KAAVEGAEKLTASQLEFAKDRIIMGT 700
Query: 666 KER----SKEKWEQVAINEAAMAVAAMNLPNFDNIEYITIAPRAGRELGYVRTMLESINF 721
+ + S+E + A +E+ A+ A+N I TI PR G LG V + S
Sbjct: 701 ERKTMFVSEESKKLTAYHESGHAIVALNTDGAHPIHKATIMPR-GSALGMVTQLPSS--- 868
Query: 722 NDGMLTRQSLFDHITVQLAPRAADEMWFGKDQLSTIWAETADNARVAARMYMIG--GLSD 779
++ ++++ L + V + R A+E+ FG+D ++T + +A A+ YM+ G+SD
Sbjct: 869 DETSISKKQLLARLDVCMGGRVAEELIFGRDNVTTGASSDLQSATELAQ-YMVSSCGMSD 1045
Query: 780 KYR--GVSNFWVTDRINEIDLEAMKILNLCYERAKEILQQNKTLMDTLVNELVVKKTLTK 837
+ ++ + ID E +K+L Y+R K +L++++ + TL N L+ +TL+
Sbjct: 1046TIGPIHIKERPSSEMQSRIDAEVVKLLRDAYDRVKALLKKHEKALHTLANALLESETLSA 1225
Query: 838 EDIVRLV 844
E+I RL+
Sbjct: 1226EEIRRLL 1246
>AL379678 homologue to GP|2062173|gb| cell division protein FtsH isolog
{Arabidopsis thaliana}, partial (12%)
Length = 376
Score = 245 bits (625), Expect = 6e-65
Identities = 124/124 (100%), Positives = 124/124 (100%)
Frame = +3
Query: 404 RLPQYLERGVDVKFTDVAGLGKIRLELEEIVKFFTHGEMYRRRGVKIPGGILLCGPPGVG 463
RLPQYLERGVDVKFTDVAGLGKIRLELEEIVKFFTHGEMYRRRGVKIPGGILLCGPPGVG
Sbjct: 3 RLPQYLERGVDVKFTDVAGLGKIRLELEEIVKFFTHGEMYRRRGVKIPGGILLCGPPGVG 182
Query: 464 KTLLAKAVAGEAGVNFFSISASQFVEIYVGVGASRVRSLYQEAKENAPSVVFIDELDAVG 523
KTLLAKAVAGEAGVNFFSISASQFVEIYVGVGASRVRSLYQEAKENAPSVVFIDELDAVG
Sbjct: 183 KTLLAKAVAGEAGVNFFSISASQFVEIYVGVGASRVRSLYQEAKENAPSVVFIDELDAVG 362
Query: 524 RKRG 527
RKRG
Sbjct: 363 RKRG 374
>TC84403 homologue to GP|15021761|gb|AAK77908.1 AAA-metalloprotease FtsH
{Pisum sativum}, partial (36%)
Length = 897
Score = 236 bits (601), Expect = 4e-62
Identities = 140/303 (46%), Positives = 191/303 (62%), Gaps = 5/303 (1%)
Frame = +3
Query: 447 GVKIPGGILLCGPPGVGKTLLAKAVAGEAGVNFFSISASQFVEIYVGVGASRVRSLYQEA 506
G KIP G LL GPPG GKTLLAKA AGE+GV F SIS S F+E++VGVG+SRVR+L++EA
Sbjct: 3 GAKIPKGALLVGPPGTGKTLLAKATAGESGVPFLSISGSDFLELFVGVGSSRVRNLFKEA 182
Query: 507 KENAPSVVFIDELDAVGRKRGLIKGS-GGQERDATLNQLLVCLDGFEGRGEVITIASTNR 565
++ APS+VFIDE+DA+GR RG G ER++TLNQLLV +DGF V+ +A TNR
Sbjct: 183 RKCAPSIVFIDEIDAIGRARGSRGGDRANDERESTLNQLLVEMDGFGTTAGVVVLAGTNR 362
Query: 566 PDILDPALVRPGRFDRKIFIPKPGFIGRIEILKVHARKKPIAEDVDY--EIVASMTDGMV 623
PD+LD AL+RPGRFDR+I I KP GR +I +++ + + + Y +AS+T G
Sbjct: 363 PDVLDKALLRPGRFDRQITIDKPDIKGRDQIFQIYLKGIKLDHEPSYYSHKLASLTPGFA 542
Query: 624 GAELANIVEVAAINMMRDSRTEVSTDDLLQAAQMEERGMLDRKER--SKEKWEQVAINEA 681
GA++AN+ AA+ R V T D +AA G L++K R SK + VA +EA
Sbjct: 543 GADIANVCNEAALIAARTEEAHV-THDHFEAAIDRIIGGLEKKNRVISKLQRRTVAYHEA 719
Query: 682 AMAVAAMNLPNFDNIEYITIAPRAGRELGYVRTMLESINFNDGMLTRQSLFDHITVQLAP 741
AVA L + D + +TI PR LG+ + + + ++T++ LFD + L
Sbjct: 720 GHAVAGWFLEHTDPLLKVTIVPRGTSALGFA----QYVPNENLLMTKEQLFDRTCMTLGG 887
Query: 742 RAA 744
RAA
Sbjct: 888 RAA 896
>CB893172 homologue to PIR|T45642|T45 FtsH metalloproteinase-like protein -
Arabidopsis thaliana, partial (25%)
Length = 661
Score = 218 bits (554), Expect = 1e-56
Identities = 116/205 (56%), Positives = 145/205 (70%), Gaps = 3/205 (1%)
Frame = +3
Query: 435 KFFTHGEMYRRRGVKIPGGILLCGPPGVGKTLLAKAVAGEAGVNFFSISASQFVEIYVGV 494
+F + + Y R G + P G+LL G PG GKTLLAKAVAGEA V F S SAS+FVE+YVG+
Sbjct: 36 EFLRNPDRYARLGARPPRGVLLVGLPGTGKTLLAKAVAGEADVPFISCSASEFVELYVGM 215
Query: 495 GASRVRSLYQEAKENAPSVVFIDELDAVGRKR-GLIKGSGGQERDATLNQLLVCLDGFEG 553
GASRVR L+ AK+ APS++FIDE+DAV + R G + G ER+ TLNQLL +DGF+
Sbjct: 216 GASRVRDLFARAKKEAPSIIFIDEIDAVAKSRDGKFRIVGNDEREQTLNQLLTEMDGFDS 395
Query: 554 RGEVITIASTNRPDILDPALVRPGRFDRKIFIPKPGFIGRIEILKVHARKK--PIAEDVD 611
VI +A+TNR D+LDPAL RPGRFDR + + P IGR ILKVH KK P+A+DV
Sbjct: 396 NSPVIVLAATNRADVLDPALRRPGRFDRIVMVETPDRIGRESILKVHVSKKELPLAKDVY 575
Query: 612 YEIVASMTDGMVGAELANIVEVAAI 636
+ASMT G GA+LAN+V AA+
Sbjct: 576 IGDIASMTTGFTGADLANLVNEAAL 650
>BI271823 similar to GP|21592745|gb cell division protein FtsH-like protein
{Arabidopsis thaliana}, partial (33%)
Length = 714
Score = 196 bits (499), Expect = 2e-50
Identities = 101/179 (56%), Positives = 129/179 (71%)
Frame = +3
Query: 415 VKFTDVAGLGKIRLELEEIVKFFTHGEMYRRRGVKIPGGILLCGPPGVGKTLLAKAVAGE 474
V F DV G+ ++EL EIV Y++ G K+P G+LL GPPG GKTLLA+AVAGE
Sbjct: 90 VGFEDVQGVDSAKVELMEIVSCLQGDINYQKLGAKLPRGVLLVGPPGTGKTLLARAVAGE 269
Query: 475 AGVNFFSISASQFVEIYVGVGASRVRSLYQEAKENAPSVVFIDELDAVGRKRGLIKGSGG 534
AGV FF++SAS+FVE++VG GA+R+R L+ A++ APS++FIDELDAVG KRG
Sbjct: 270 AGVPFFTVSASEFVEMFVGRGAARIRDLFSRARKFAPSIIFIDELDAVGGKRG---RGFN 440
Query: 535 QERDATLNQLLVCLDGFEGRGEVITIASTNRPDILDPALVRPGRFDRKIFIPKPGFIGR 593
+ERD TLNQLL +DGFE V+ IA+TNRP+ LDPAL RPGRF K+F+ +P GR
Sbjct: 441 EERDQTLNQLLTEMDGFESEIRVVVIAATNRPEALDPALCRPGRFSXKVFVGEPDEEGR 617
>TC86159 homologue to GP|21387177|gb|AAM47992.1 26S proteasome AAA-ATPase
subunit RPT4a-like protein {Arabidopsis thaliana},
complete
Length = 1640
Score = 192 bits (489), Expect = 4e-49
Identities = 124/367 (33%), Positives = 197/367 (52%), Gaps = 33/367 (8%)
Frame = +2
Query: 323 RTVVVSYKK---QKKDYEDRIKIQKADAEERRK-MREMEAEMGWSEAGGD---------E 369
R V Y+K Q K+ E R++ + + +K + E ++ ++ G +
Sbjct: 104 RNAVAEYRKKLLQHKELESRVRSVRENLRASKKEFNKTEDDLKSLQSVGQIIGEVLRPLD 283
Query: 370 DESELVKEGEENPYL-----KMTKEFMKSGARVRRAQN-----RRLPQYLERGV------ 413
+E +VK Y+ K+ KE + SG RV R LP+ ++ V
Sbjct: 284 NERLIVKASSGPRYVVGCRSKVDKEKLTSGTRVVLDMTTLTIMRALPREVDPVVYNMLHE 463
Query: 414 ---DVKFTDVAGLGKIRLELEEIVKF-FTHGEMYRRRGVKIPGGILLCGPPGVGKTLLAK 469
++ ++ V GL EL E ++ + E++ R G+K P G+LL GPPG GKTLLA+
Sbjct: 464 DPGNISYSAVGGLSDQIRELRESIELPLMNPELFLRVGIKPPKGVLLYGPPGTGKTLLAR 643
Query: 470 AVAGEAGVNFFSISASQFVEIYVGVGASRVRSLYQEAKENAPSVVFIDELDAVGRKRGLI 529
A+A NF + +S ++ Y+G A +R ++ A+++ P ++F+DE+DA+G +R
Sbjct: 644 AIASNIDANFLKVVSSAIIDKYIGESARLIREMFGYARDHQPCIIFMDEIDAIGGRRFSE 823
Query: 530 KGSGGQERDATLNQLLVCLDGFEGRGEVITIASTNRPDILDPALVRPGRFDRKIFIPKPG 589
S +E TL +LL LDGF+ G+V I +TNRPD+LDPAL+RPGR DRKI IP P
Sbjct: 824 GTSADREIQRTLMELLNQLDGFDQLGKVKMIMATNRPDVLDPALLRPGRLDRKIEIPLPN 1003
Query: 590 FIGRIEILKVHARKKPIAEDVDYEIVASMTDGMVGAELANIVEVAAINMMRDSRTEVSTD 649
R+EILK+HA ++DYE V + +G GA+L N+ A ++ +R R V +
Sbjct: 1004EQSRMEILKIHAAGIAKHGEIDYEAVVKLAEGFNGADLRNVCTEAGMSAIRAERDYVIHE 1183
Query: 650 DLLQAAQ 656
D ++A +
Sbjct: 1184DFMKAVR 1204
>TC85768 homologue to SP|P54774|CC48_SOYBN Cell division cycle protein 48
homolog (Valosin containing protein homolog) (VCP).
[Soybean], partial (95%)
Length = 2785
Score = 190 bits (483), Expect = 2e-48
Identities = 102/263 (38%), Positives = 158/263 (59%), Gaps = 1/263 (0%)
Frame = +2
Query: 389 EFMKSGARVRRAQNRRLPQYLERGVDVKFTDVAGLGKIRLELEEIVKF-FTHGEMYRRRG 447
E G ++R RL + V + DV G+ K ++ E+V+ H ++++ G
Sbjct: 650 EIFCEGEPIKREDENRLDE-------VGYDDVGGVRKQMAQIRELVELPLRHPQLFKSIG 808
Query: 448 VKIPGGILLCGPPGVGKTLLAKAVAGEAGVNFFSISASQFVEIYVGVGASRVRSLYQEAK 507
VK P GILL GPPG GKTL+A+AVA E G FF I+ + + G S +R ++EA+
Sbjct: 809 VKPPKGILLYGPPGSGKTLIARAVANETGAFFFCINGPEIMSKLAGESESNLRKAFEEAE 988
Query: 508 ENAPSVVFIDELDAVGRKRGLIKGSGGQERDATLNQLLVCLDGFEGRGEVITIASTNRPD 567
+NAPS++FIDE+D++ KR + + G+ ++QLL +DG + R VI + +TNRP+
Sbjct: 989 KNAPSIIFIDEIDSIAPKR---EKTHGEVERRIVSQLLTLMDGLKSRAHVIVMGATNRPN 1159
Query: 568 ILDPALVRPGRFDRKIFIPKPGFIGRIEILKVHARKKPIAEDVDYEIVASMTDGMVGAEL 627
+DPAL R GRFDR+I I P +GR+E+L++H + +AEDVD E ++ T G VGA+L
Sbjct: 1160SIDPALRRFGRFDREIDIGVPDEVGRLEVLRIHTKNMKLAEDVDLEKISKETHGYVGADL 1339
Query: 628 ANIVEVAAINMMRDSRTEVSTDD 650
A + AA+ +R+ + +D
Sbjct: 1340AALCTEAALQCIREKMDVIDLED 1408
Score = 181 bits (459), Expect = 1e-45
Identities = 93/225 (41%), Positives = 135/225 (59%), Gaps = 1/225 (0%)
Frame = +2
Query: 419 DVAGLGKIRLELEEIVKF-FTHGEMYRRRGVKIPGGILLCGPPGVGKTLLAKAVAGEAGV 477
D+ GL ++ EL+E V++ H E + + G+ G+L GPPG GKTLLAKA+A E
Sbjct: 1538 DIGGLENVKRELQETVQYPVEHPEKFEKFGMSPSKGVLFYGPPGCGKTLLAKAIANECQA 1717
Query: 478 NFFSISASQFVEIYVGVGASRVRSLYQEAKENAPSVVFIDELDAVGRKRGLIKGSGGQER 537
NF SI + + ++ G + VR ++ +A+ +AP V+F DELD++ +RG G G
Sbjct: 1718 NFISIKGPELLTMWFGESEANVREIFDKARGSAPCVLFFDELDSIATQRGSSVGDAGGAA 1897
Query: 538 DATLNQLLVCLDGFEGRGEVITIASTNRPDILDPALVRPGRFDRKIFIPKPGFIGRIEIL 597
D LNQLL +DG + V I +TNRPDI+DPAL+RPGR D+ I+IP P R +I
Sbjct: 1898 DRVLNQLLTEMDGMSAKKTVFIIGATNRPDIIDPALLRPGRLDQLIYIPLPDEDSRHQIF 2077
Query: 598 KVHARKKPIAEDVDYEIVASMTDGMVGAELANIVEVAAINMMRDS 642
K RK PI++DVD +A T G GA++ I + A +R++
Sbjct: 2078 KACLRKSPISKDVDIRALAKYTQGFSGADITEICQRACKYAIREN 2212
>TC77012 homologue to SP|P54776|PRSA_LYCES 26S protease regulatory subunit
6A homolog (TAT-binding protein homolog 1) (TBP-1)
(Mg(2+), complete
Length = 1793
Score = 188 bits (477), Expect = 9e-48
Identities = 102/235 (43%), Positives = 145/235 (61%), Gaps = 1/235 (0%)
Frame = +1
Query: 417 FTDVAGLGKIRLEL-EEIVKFFTHGEMYRRRGVKIPGGILLCGPPGVGKTLLAKAVAGEA 475
+ D+ GL K EL E IV TH E +++ G++ P G+LL GPPG GKTL+A+A A +
Sbjct: 637 YNDIGGLEKQIQELVEAIVLPMTHKERFQKLGIRPPKGVLLYGPPGTGKTLMARACAAQT 816
Query: 476 GVNFFSISASQFVEIYVGVGASRVRSLYQEAKENAPSVVFIDELDAVGRKRGLIKGSGGQ 535
F ++ Q V++++G GA VR +Q AKE +P ++FIDE+DA+G KR + SG +
Sbjct: 817 NATFLKLAGPQLVQMFIGDGAKLVRDAFQLAKEKSPCIIFIDEIDAIGTKRFDSEVSGDR 996
Query: 536 ERDATLNQLLVCLDGFEGRGEVITIASTNRPDILDPALVRPGRFDRKIFIPKPGFIGRIE 595
E T+ +LL LDGF + IA+TNR DILDPAL+R GR DRKI P P R
Sbjct: 997 EVQRTMLELLNQLDGFSSDDRIKVIAATNRADILDPALMRSGRLDRKIEFPHPTEEARAR 1176
Query: 596 ILKVHARKKPIAEDVDYEIVASMTDGMVGAELANIVEVAAINMMRDSRTEVSTDD 650
IL++H+RK + DV++E +A TD GA+L + A + +R TEV+ +D
Sbjct: 1177ILQIHSRKMNVHPDVNFEELARSTDDFNGAQLKAVCVEAGMLALRRDATEVNHED 1341
>TC85936 homologue to GP|13537115|dbj|BAB40755. AtSUG1 {Arabidopsis
thaliana}, partial (98%)
Length = 1633
Score = 179 bits (455), Expect = 3e-45
Identities = 97/268 (36%), Positives = 155/268 (57%), Gaps = 2/268 (0%)
Frame = +1
Query: 409 LERGVDVKFTDVAGLGKIRLELEEIVKF-FTHGEMYRRRGVKIPGGILLCGPPGVGKTLL 467
+E+ D + + GL + E++E+++ H E++ G+ P G+LL GPPG GKTLL
Sbjct: 595 VEKVPDSTYDMIGGLDQQIKEIKEVIELPIKHPELFESLGIAQPKGVLLYGPPGTGKTLL 774
Query: 468 AKAVAGEAGVNFFSISASQFVEIYVGVGASRVRSLYQEAKENAPSVVFIDELDAVGRKRG 527
A+AVA F +S S+ V+ Y+G G+ VR L+ A+E+APS++F+DE+D++G R
Sbjct: 775 ARAVAHHTDCTFIRVSGSELVQKYIGEGSRMVRELFVMAREHAPSIIFMDEIDSIGSARM 954
Query: 528 LI-KGSGGQERDATLNQLLVCLDGFEGRGEVITIASTNRPDILDPALVRPGRFDRKIFIP 586
G+G E T+ +LL LDGFE ++ + +TNR DILD AL+RPGR DRKI P
Sbjct: 955 ESGSGNGDSEVQRTMLELLNQLDGFEASNKIKVLMATNRIDILDQALLRPGRIDRKIEFP 1134
Query: 587 KPGFIGRIEILKVHARKKPIAEDVDYEIVASMTDGMVGAELANIVEVAAINMMRDSRTEV 646
P R +ILK+H+R+ + +D + +A +G GAEL + A + +R+ R V
Sbjct: 1135NPNEESRRDILKIHSRRMNLMRGIDLKKIAEKMNGASGAELKAVCTEAGMFALRERRVHV 1314
Query: 647 STDDLLQAAQMEERGMLDRKERSKEKWE 674
+ +D A + ++ ++ W+
Sbjct: 1315TQEDFEMAVAKVMKKETEKNMSLRKLWK 1398
>BF647954 homologue to GP|11094192|db 26S proteasome regulatory particle
triple-A ATPase subunit4 {Oryza sativa (japonica
cultivar-group)}, partial (53%)
Length = 649
Score = 179 bits (453), Expect = 5e-45
Identities = 91/212 (42%), Positives = 134/212 (62%)
Frame = +2
Query: 441 EMYRRRGVKIPGGILLCGPPGVGKTLLAKAVAGEAGVNFFSISASQFVEIYVGVGASRVR 500
E++ R G+K P G+LL GPPG GKTLLA+A+A NF + +S ++ Y+G + +R
Sbjct: 11 ELFLRVGIKPPKGVLLYGPPGTGKTLLARAIASNIDANFLKVVSSAIIDKYIGESSRLIR 190
Query: 501 SLYQEAKENAPSVVFIDELDAVGRKRGLIKGSGGQERDATLNQLLVCLDGFEGRGEVITI 560
++ A+++ P ++F+DE+DA+G +R S +E TL +LL LDGF+ G+V I
Sbjct: 191 EMFGYARDHQPCIIFMDEIDAIGGRRFSEGTSADREIQRTLMELLNQLDGFDQLGKVKII 370
Query: 561 ASTNRPDILDPALVRPGRFDRKIFIPKPGFIGRIEILKVHARKKPIAEDVDYEIVASMTD 620
+TNRPD+LDPAL+RPGR DRKI IP P R+EILK+HA ++DYE V + +
Sbjct: 371 MATNRPDVLDPALLRPGRLDRKIEIPLPNEQSRMEILKIHAAGIAKHGEIDYEAVVKLAE 550
Query: 621 GMVGAELANIVEVAAINMMRDSRTEVSTDDLL 652
G GA+L NI A ++ +R R V +D +
Sbjct: 551 GFNGADLRNICTEAGMSAIRAERDYVIHEDFM 646
>TC85745 homologue to GP|17297987|dbj|BAB78491. 26S proteasome regulatory
particle triple-A ATPase subunit2b {Oryza sativa
(japonica cultivar-group), partial (98%)
Length = 1696
Score = 177 bits (450), Expect = 1e-44
Identities = 100/252 (39%), Positives = 147/252 (57%), Gaps = 1/252 (0%)
Frame = +3
Query: 417 FTDVAGLGKIRLELEEIVKF-FTHGEMYRRRGVKIPGGILLCGPPGVGKTLLAKAVAGEA 475
+ D+ GL E++E V+ TH E+Y G+K P G++L G PG GKTLLAKAVA
Sbjct: 648 YADIGGLDAQIQEIKEAVELPLTHPELYEDIGIKPPKGVILYGEPGTGKTLLAKAVANST 827
Query: 476 GVNFFSISASQFVEIYVGVGASRVRSLYQEAKENAPSVVFIDELDAVGRKRGLIKGSGGQ 535
F + S+ ++ Y+G G VR L++ A + +P++VFIDE+DAVG KR G +
Sbjct: 828 SATFLRVVGSELIQKYLGDGPKLVRELFRVADDLSPAIVFIDEIDAVGTKRYDAHSGGER 1007
Query: 536 ERDATLNQLLVCLDGFEGRGEVITIASTNRPDILDPALVRPGRFDRKIFIPKPGFIGRIE 595
E T+ +LL LDGF+ RG+V I +TNR + LDPAL+RPGR DRKI P P R
Sbjct: 1008EIQRTMLELLNQLDGFDSRGDVKVILATNRIESLDPALLRPGRIDRKIEFPLPDIKTRRR 1187
Query: 596 ILKVHARKKPIAEDVDYEIVASMTDGMVGAELANIVEVAAINMMRDSRTEVSTDDLLQAA 655
I ++H + +A+DV+ E D GA++ I A + +R+ R +V+ D +A
Sbjct: 1188IFQIHTSRMTLADDVNLEEFVMTKDEFSGADIKAICTEAGLLALRERRMKVTHPDFKKA- 1364
Query: 656 QMEERGMLDRKE 667
+++ M +KE
Sbjct: 1365--KDKVMFKKKE 1394
>TC85744 homologue to PIR|T08959|T08959 proteinase homolog F19B15.70 -
Arabidopsis thaliana, complete
Length = 1629
Score = 177 bits (450), Expect = 1e-44
Identities = 101/252 (40%), Positives = 146/252 (57%), Gaps = 1/252 (0%)
Frame = +2
Query: 417 FTDVAGLGKIRLELEEIVKF-FTHGEMYRRRGVKIPGGILLCGPPGVGKTLLAKAVAGEA 475
+ D+ GL E++E V+ TH E+Y G+K P G++L G PG GKTLLAKAVA
Sbjct: 650 YADIGGLDAQIQEIKEAVELPLTHPELYEDIGIKPPKGVILYGEPGTGKTLLAKAVANST 829
Query: 476 GVNFFSISASQFVEIYVGVGASRVRSLYQEAKENAPSVVFIDELDAVGRKRGLIKGSGGQ 535
F + S+ ++ Y+G G VR L++ A + +PS+VFIDE+DAVG KR G +
Sbjct: 830 SATFLRVVGSELIQKYLGDGPKLVRELFRVADDLSPSIVFIDEIDAVGTKRYDAHSGGER 1009
Query: 536 ERDATLNQLLVCLDGFEGRGEVITIASTNRPDILDPALVRPGRFDRKIFIPKPGFIGRIE 595
E T+ +LL LDGF+ RG+V I +TNR + LDPAL+RPGR DRKI P P R
Sbjct: 1010EIQRTMLELLNQLDGFDSRGDVKVILATNRIESLDPALLRPGRIDRKIEFPLPDIKTRRR 1189
Query: 596 ILKVHARKKPIAEDVDYEIVASMTDGMVGAELANIVEVAAINMMRDSRTEVSTDDLLQAA 655
I +H + +A+DV+ E D GA++ I A + +R+ R +V+ D +A
Sbjct: 1190IFTIHTSRMTLADDVNLEEFVMTKDEFSGADIKAICTEAGLLALRERRMKVTHADFKKA- 1366
Query: 656 QMEERGMLDRKE 667
+++ M +KE
Sbjct: 1367--KDKVMFKKKE 1396
>TC78601 homologue to GP|15021761|gb|AAK77908.1 AAA-metalloprotease FtsH
{Pisum sativum}, partial (49%)
Length = 1338
Score = 144 bits (362), Expect(2) = 7e-42
Identities = 113/336 (33%), Positives = 177/336 (52%), Gaps = 17/336 (5%)
Frame = +1
Query: 526 RGLIKGSGGQERDATLNQLLVCLDGFEGRGEVITIASTNRPDILDPALVRPGRFDRKIFI 585
RG GS ER++TLNQLLV +DGF V+ +A TNR DILD AL+RPGRFDR I I
Sbjct: 85 RGGFSGSN-DERESTLNQLLVEMDGFGTTAGVVVLAGTNRADILDNALLRPGRFDRTISI 261
Query: 586 PKPGFIGRIEILKVHARKKPIAEDVDY--EIVASMTDGMVGAELANIVEVAAINMMRDSR 643
P GR +I +++ +K + + Y + +A++T G GA++AN+ AA+ R
Sbjct: 262 DVPDIKGRDQIFQIYLKKIKLDHEPSYYSQRLAALTPGFAGADIANVCNEAALIAARTDE 441
Query: 644 TEVSTDDLLQAAQMEERGMLDRKER--SKEKWEQVAINEAAMAVAAMNLPNFDNIEYITI 701
++V T D +AA G L++K R SK + VA +EA AVA L + + + +TI
Sbjct: 442 SQV-TMDHFEAAIDRIIGGLEKKNRVISKRERRTVAYHEAGHAVAGWFLEHCEPLLKVTI 618
Query: 702 APRAGRELGYVRTMLESINFNDGMLTRQSLFDHITVQLAPRAADEMWFGKDQLSTIWAET 761
PR LG+ + + + + T++ L D + L RAA+++ G +ST
Sbjct: 619 VPRGTAALGFA----QYVPSENLLRTKEQLLDMTCMTLGGRAAEQVLIG--AIST--GAQ 774
Query: 762 ADNARVAARMY---MIGGLSDKYRGVSNF----------WVTDRINEIDLEAMKILNLCY 808
D +V Y I G S+K G+ +F + D N ID E +N Y
Sbjct: 775 NDLEKVTKMTYAQVAIYGFSEKV-GLLSFPQNEDQFGKPYSGDTGNIIDQEVRDWVNHAY 951
Query: 809 ERAKEILQQNKTLMDTLVNELVVKKTLTKEDIVRLV 844
ER ++++++K + + L+ K+ L +ED+VR++
Sbjct: 952 ERTVQLIEEHKEKLAQIAELLLEKEVLHQEDLVRIL 1059
Score = 46.2 bits (108), Expect(2) = 7e-42
Identities = 19/27 (70%), Positives = 26/27 (95%)
Frame = +3
Query: 501 SLYQEAKENAPSVVFIDELDAVGRKRG 527
+L+QEA++ APS+VFIDE+DA+GRKRG
Sbjct: 3 NLFQEARQCAPSIVFIDEIDAIGRKRG 83
>TC77261 homologue to GP|8777330|dbj|BAA96920.1 26S proteasome AAA-ATPase
subunit RPT3 {Arabidopsis thaliana}, partial (94%)
Length = 1575
Score = 167 bits (422), Expect = 2e-41
Identities = 93/238 (39%), Positives = 139/238 (58%), Gaps = 1/238 (0%)
Frame = +2
Query: 414 DVKFTDVAGLGKIRLELEEIVKF-FTHGEMYRRRGVKIPGGILLCGPPGVGKTLLAKAVA 472
DV + D+ G + E+ E V+ TH E+Y++ G+ P G+LL GPPG GKT+LAKAVA
Sbjct: 563 DVTYNDIGGCDIQKQEIREAVELPLTHHELYKQIGIDPPRGVLLYGPPGTGKTMLAKAVA 742
Query: 473 GEAGVNFFSISASQFVEIYVGVGASRVRSLYQEAKENAPSVVFIDELDAVGRKRGLIKGS 532
F + S+FV+ Y+G G VR +++ AKENAP+++FIDE+DA+ R +
Sbjct: 743 NHTTAAFIRVVGSEFVQKYLGEGPRMVRDVFRLAKENAPAIIFIDEVDAIATARFDAQTG 922
Query: 533 GGQERDATLNQLLVCLDGFEGRGEVITIASTNRPDILDPALVRPGRFDRKIFIPKPGFIG 592
+E L +LL +DGF+ V I +TNR D LDPAL+RPGR DRKI P P
Sbjct: 923 ADREVQRILMELLNQMDGFDQTVNVKVIMATNRADTLDPALLRPGRLDRKIEFPLPDRRQ 1102
Query: 593 RIEILKVHARKKPIAEDVDYEIVASMTDGMVGAELANIVEVAAINMMRDSRTEVSTDD 650
+ + +V K ++++VD E S D + AE++ I + A ++ +R +R + D
Sbjct: 1103KRLVFQVCTAKMNLSDEVDLEDYVSRPDKISAAEISAICQEAGMHAVRKNRYVILPKD 1276
>TC86186 similar to GP|4325041|gb|AAD17230.1| FtsH-like protein Pftf
precursor {Nicotiana tabacum}, partial (47%)
Length = 1303
Score = 142 bits (359), Expect(2) = 2e-41
Identities = 68/117 (58%), Positives = 90/117 (76%)
Frame = +2
Query: 409 LERGVDVKFTDVAGLGKIRLELEEIVKFFTHGEMYRRRGVKIPGGILLCGPPGVGKTLLA 468
+E V F DVAG+ + + + E+V+F E + G +IP G+LL GPPG GKTLLA
Sbjct: 827 MEPNTGVTFDDVAGVDEAKQDFMEVVEFLKKPERFTAIGARIPKGVLLIGPPGTGKTLLA 1006
Query: 469 KAVAGEAGVNFFSISASQFVEIYVGVGASRVRSLYQEAKENAPSVVFIDELDAVGRK 525
KA+AGEAGV FFSIS S+FVE++VG+GASRVR L+++AKENAP +VF+DE+DAVGR+
Sbjct: 1007KAIAGEAGVPFFSISGSEFVEMFVGIGASRVRDLFKKAKENAPCIVFVDEIDAVGRQ 1177
Score = 45.4 bits (106), Expect(2) = 2e-41
Identities = 23/40 (57%), Positives = 26/40 (64%)
Frame = +3
Query: 526 RGLIKGSGGQERDATLNQLLVCLDGFEGRGEVITIASTNR 565
RG G G ER+ TLNQLL +DGFEG +I IA TNR
Sbjct: 1179 RGTGIGGGNDEREHTLNQLLTEMDGFEGNTGIIVIAPTNR 1298
>TC87676 PIR|G96577|G96577 26S proteasome ATPase subunit [imported] -
Arabidopsis thaliana, partial (56%)
Length = 963
Score = 161 bits (408), Expect = 9e-40
Identities = 85/223 (38%), Positives = 123/223 (55%)
Frame = +2
Query: 439 HGEMYRRRGVKIPGGILLCGPPGVGKTLLAKAVAGEAGVNFFSISASQFVEIYVGVGASR 498
H E + + G+ P G+L GPPG GKTLLA+AVA F + S+ V+ YVG GA
Sbjct: 17 HPEKFVKLGIDPPKGVLCYGPPGTGKTLLARAVANRTDACFIRVIGSELVQKYVGEGARM 196
Query: 499 VRSLYQEAKENAPSVVFIDELDAVGRKRGLIKGSGGQERDATLNQLLVCLDGFEGRGEVI 558
VR L+Q A+ +VF DE+DA+G R G E T+ +++ LDGF+ RG +
Sbjct: 197 VRELFQMARSKKACIVFFDEVDAIGGARFDDGVGGDNEVQRTMLEIVNQLDGFDARGNIK 376
Query: 559 TIASTNRPDILDPALVRPGRFDRKIFIPKPGFIGRIEILKVHARKKPIAEDVDYEIVASM 618
+ +TNRPD LDPAL+RPGR DRK+ P R +I K+H R D+ +E++A +
Sbjct: 377 VLMATNRPDTLDPALLRPGRLDRKVEFGLPDLESRTQIFKIHTRTMNCERDIRFELLARL 556
Query: 619 TDGMVGAELANIVEVAAINMMRDSRTEVSTDDLLQAAQMEERG 661
GA++ ++ A + +R R V+ D L A +G
Sbjct: 557 CPNSTGADIRSVCTEAGMYAIRARRKTVTEKDFLDAVNKVIKG 685
>TC81426 similar to GP|6016719|gb|AAF01545.1| putative cell division control
protein {Arabidopsis thaliana}, partial (21%)
Length = 965
Score = 152 bits (383), Expect = 7e-37
Identities = 85/195 (43%), Positives = 115/195 (58%), Gaps = 2/195 (1%)
Frame = +2
Query: 456 LCGPPGVGKTLLAKAVAGEAGVNFFSISASQFVEIYVGVGASRVRSLYQEAKENAPSVVF 515
L GPPG GKTL+AKAVA EAG NF I + + YVG VR+L+ A+ AP V+F
Sbjct: 2 LFGPPGCGKTLIAKAVANEAGANFIHIKGPELLNKYVGESELAVRTLFNRARTCAPCVLF 181
Query: 516 IDELDAVGRKRGLIKGSGGQERDATLNQLLVCLDGFEGRGEVITIASTNRPDILDPALVR 575
DE+DA+ KRG GG + LNQLL+ LDG E R V I +TNRPD++DPAL+R
Sbjct: 182 FDEVDALTTKRGK---EGGWVIERLLNQLLIELDGAEQRRGVFVIGATNRPDVMDPALLR 352
Query: 576 PGRFDRKIFIPKPGFIGRIEILKVHARKKPIAEDVDYEIVASM--TDGMVGAELANIVEV 633
PGRF + +++P P R+ ILK AR K I VD + M + + GA+LA ++
Sbjct: 353 PGRFGKLLYVPLPSPDDRVLILKALARNKHIDSSVDLSAIGRMDACENLSGADLAELMNE 532
Query: 634 AAINMMRDSRTEVST 648
A + + + + T
Sbjct: 533 AVMAALDEKLASIET 577
Database: MTGI
Posted date: Oct 22, 2004 3:39 PM
Number of letters in database: 27,044,181
Number of sequences in database: 36,976
Lambda K H
0.317 0.134 0.375
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 21,827,121
Number of Sequences: 36976
Number of extensions: 264908
Number of successful extensions: 1948
Number of sequences better than 10.0: 137
Number of HSP's better than 10.0 without gapping: 1879
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 1918
length of query: 881
length of database: 9,014,727
effective HSP length: 105
effective length of query: 776
effective length of database: 5,132,247
effective search space: 3982623672
effective search space used: 3982623672
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.3 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.7 bits)
S2: 63 (28.9 bits)
Medicago: description of AC138130.19


