

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
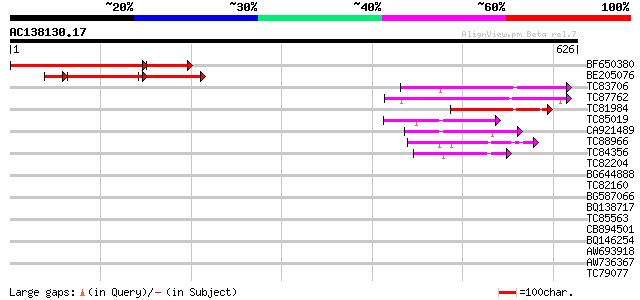
Query= AC138130.17 + phase: 0 /pseudo
(626 letters)
Database: MTGI
36,976 sequences; 27,044,181 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
BF650380 similar to GP|18203774|gb Unknown (protein for MGC:1404... 307 e-104
BE205076 similar to EGAD|59173|6188 85 kDa calcium-independent p... 151 9e-47
TC83706 weakly similar to PIR|T48283|T48283 ankyrin-like protein... 82 5e-16
TC87762 similar to GP|21537142|gb|AAM61483.1 unknown {Arabidopsi... 82 6e-16
TC81984 homologue to GP|12321945|gb|AAG51002.1 ankyrin-like prot... 68 1e-11
TC85019 weakly similar to GP|9293890|dbj|BAB01793.1 emb|CAB70981... 60 2e-09
CA921489 weakly similar to PIR|T47566|T475 hypothetical protein ... 59 6e-09
TC88966 similar to GP|21305941|gb|AAM45816.1 NADH dehydrogenase ... 54 2e-07
TC84356 46 5e-05
TC82204 similar to GP|21537142|gb|AAM61483.1 unknown {Arabidopsi... 39 0.008
BG644888 weakly similar to PIR|F85043|F850 hypothetical protein ... 39 0.008
TC82160 38 0.010
BG587066 similar to PIR|E84725|E84 ankyrin-like protein [importe... 35 0.017
BQ138717 similar to GP|12321945|gb ankyrin-like protein; 93648-9... 37 0.017
TC85563 similar to GP|14279688|gb|AAK58690.1 receptor-like kinas... 37 0.023
CB894501 weakly similar to PIR|F86279|F86 hypothetical protein A... 37 0.030
BQ146254 weakly similar to GP|11994366|dbj gene_id:MDC16.7~unkno... 36 0.051
AW693918 similar to GP|9759271|dbj contains similarity to ankyri... 35 0.066
AW736367 weakly similar to PIR|T01879|T01 hypothetical protein F... 35 0.066
TC79077 similar to GP|17529252|gb|AAL38853.1 unknown protein {Ar... 35 0.086
>BF650380 similar to GP|18203774|gb Unknown (protein for MGC:14049) {Mus
musculus}, partial (1%)
Length = 664
Score = 307 bits (786), Expect(2) = e-104
Identities = 151/152 (99%), Positives = 152/152 (99%)
Frame = +1
Query: 1 MDQEFFDAIKKNDMITFSSIVKEREGILNQKTDDTFSAPLHLASKYGCIEMVSEIVKLCP 60
MDQEFFDAIKKNDMITFSSIVKEREGILNQKTDDTFSAPLHLASKYGCIEMVSEIVKLCP
Sbjct: 46 MDQEFFDAIKKNDMITFSSIVKEREGILNQKTDDTFSAPLHLASKYGCIEMVSEIVKLCP 225
Query: 61 DMVSAENKNMETPIHEACRQENVKVLMLLLEVNPTAACKLNPTCKSAFLVACSHGHLDLV 120
DMVSAENKNMETPIHEACRQENVKVLMLLLEVNPTAACKLNPTCKSAFLVACSHGHLDLV
Sbjct: 226 DMVSAENKNMETPIHEACRQENVKVLMLLLEVNPTAACKLNPTCKSAFLVACSHGHLDLV 405
Query: 121 NLLLNLSEIVGQEVAGFDQACFHVAAVRGHTE 152
NLLLNLSEIVGQEVAGFDQACFHVAAVRGHT+
Sbjct: 406 NLLLNLSEIVGQEVAGFDQACFHVAAVRGHTD 501
Score = 89.4 bits (220), Expect(2) = e-104
Identities = 46/51 (90%), Positives = 46/51 (90%)
Frame = +2
Query: 152 ESC*TSGRILFK*STRKGIQHCIMLVIKDTLRLYGSY*VAIQNLLYNTTTM 202
ESC*TSGRILFK*STRKGIQHCIML IKDTLRL GSY* QNLLYNTTTM
Sbjct: 509 ESC*TSGRILFK*STRKGIQHCIMLXIKDTLRLDGSY*XXXQNLLYNTTTM 661
Score = 29.6 bits (65), Expect = 3.6
Identities = 17/75 (22%), Positives = 36/75 (47%)
Frame = +1
Query: 27 ILNQKTDDTFSAPLHLASKYGCIEMVSEIVKLCPDMVSAENKNMETPIHEACRQENVKVL 86
I+ Q+ A H+A+ G ++V E++ PD++ ++ T +H A + + ++
Sbjct: 430 IVGQEVAGFDQACFHVAAVRGHTDVVRELLNKWPDLIQVIDEKGNTALHHAXYKGHFEIG 609
Query: 87 MLLLEVNPTAACKLN 101
+LL A + N
Sbjct: 610 WILLXXXSKLALQYN 654
>BE205076 similar to EGAD|59173|6188 85 kDa calcium-independent phospholipase
A2 {Mus musculus}, partial (2%)
Length = 549
Score = 151 bits (382), Expect(2) = 9e-47
Identities = 74/88 (84%), Positives = 79/88 (89%)
Frame = +1
Query: 65 AENKNMETPIHEACRQENVKVLMLLLEVNPTAACKLNPTCKSAFLVACSHGHLDLVNLLL 124
AE++NMETPIHEACRQENVKVLMLLLEVN TAA LNPTCKSAF VAC HGHLDLVNLLL
Sbjct: 82 AESENMETPIHEACRQENVKVLMLLLEVNATAARTLNPTCKSAFFVACIHGHLDLVNLLL 261
Query: 125 NLSEIVGQEVAGFDQACFHVAAVRGHTE 152
NLSEIV +AGFDQACFH+AA RGHT+
Sbjct: 262 NLSEIVEPGLAGFDQACFHIAASRGHTD 345
Score = 97.8 bits (242), Expect = 1e-20
Identities = 54/74 (72%), Positives = 56/74 (74%)
Frame = +2
Query: 143 HVAAVRGHTESC*TSGRILFK*STRKGIQHCIMLVIKDTLRLYGSY*VAIQNLLYNTTTM 202
H A + ESC*T G+I K*STR GI HCIMLVIKDT RLYGSY* AIQ LLYNTTTM
Sbjct: 326 HQEATQILLESC*TGGQIFLK*STRMGILHCIMLVIKDTERLYGSY*NAIQMLLYNTTTM 505
Query: 203 VIHHCILQ*LRGKS 216
VIHHC LQ* KS
Sbjct: 506 VIHHCTLQ**TEKS 547
Score = 54.3 bits (129), Expect(2) = 9e-47
Identities = 24/26 (92%), Positives = 25/26 (95%)
Frame = +3
Query: 39 PLHLASKYGCIEMVSEIVKLCPDMVS 64
PLHLA KYGCIEMVSEIV+LCPDMVS
Sbjct: 3 PLHLACKYGCIEMVSEIVRLCPDMVS 80
Score = 35.0 bits (79), Expect = 0.086
Identities = 16/64 (25%), Positives = 35/64 (54%)
Frame = +1
Query: 38 APLHLASKYGCIEMVSEIVKLCPDMVSAENKNMETPIHEACRQENVKVLMLLLEVNPTAA 97
A H+A+ G ++V E++ PD+ ++N + +H AC + + + + +LL+ + A
Sbjct: 307 ACFHIAASRGHTDIVRELLNRWPDLSQVIDENGNSALHHACNKGHRETVWILLKRDSNVA 486
Query: 98 CKLN 101
+ N
Sbjct: 487 LQYN 498
>TC83706 weakly similar to PIR|T48283|T48283 ankyrin-like protein -
Arabidopsis thaliana, partial (34%)
Length = 746
Score = 82.4 bits (202), Expect = 5e-16
Identities = 56/197 (28%), Positives = 97/197 (48%), Gaps = 8/197 (4%)
Frame = +1
Query: 432 KHYHEMHKEALLNARNTIVLVAVLIATVTFAAGISPPGGVYQE------GPKKGISMAGE 485
K ++M E L NA N+ +VAVLIATV FAA + PG Q G G +
Sbjct: 55 KRINKMQAEGLNNAINSNTVVAVLIATVAFAAIFTVPGQYPQNTKNLAPGMSPGEANIAP 234
Query: 486 TSAFKVFAISNIIALFTSLSVVIVLVSIIPFRRKPQTILLTIAHKVMWVAVAFMGTGYVA 545
F +F I + ALF SL+VVIV S++ R+ + + + +K+MW+A + ++A
Sbjct: 235 NIEFLIFVIFDSTALFISLAVVIVQTSVVVIEREAKKQMTAVINKLMWIACVLISVAFLA 414
Query: 546 ATWVILPHNQEMQWLSVVLLALGGGFLGTIFISLSVMLVEHWLRKSSWRKKR--KESGDG 603
+++++ +E L++ ALG + +L ++ H L S R +R S
Sbjct: 415 MSYIVVGDQKE---LAIAATALGTVIMAATLGTLCYWVIAHRLEASRLRSQRTMTSSRHS 585
Query: 604 TAESDKESEDSDFQSSY 620
S + ++++++ Y
Sbjct: 586 LTMSGMSASENEYKTVY 636
>TC87762 similar to GP|21537142|gb|AAM61483.1 unknown {Arabidopsis
thaliana}, partial (65%)
Length = 1594
Score = 82.0 bits (201), Expect = 6e-16
Identities = 56/221 (25%), Positives = 110/221 (49%), Gaps = 15/221 (6%)
Frame = +1
Query: 415 KHHHNKGKIENVNHTKR-----KHYHEMHKEALLNARNTIVLVAVLIATVTFAAGISPPG 469
KH I+N +R K ++H+EA+ N N++ +VAVL A++ F A + PG
Sbjct: 460 KHEVQSQLIQNEKTRRRVSGIAKELKKLHREAVQNTINSVTVVAVLFASIAFLAIFNLPG 639
Query: 470 GVYQEGPKKGISMAGETSAFKVFAISNIIALFTSLSVVIVLVSIIPFRRKPQTILLTIAH 529
+G G S + F++F + N +LF SL+VV+V ++++ + + Q ++++ +
Sbjct: 640 QYIMKGSHIGESNIADHVGFQIFCLLNSTSLFISLAVVVVQITLVAWDTRAQKQIVSVVN 819
Query: 530 KVMWVAVAFMGTGYVAATWVILPHNQEMQWLSVVLLALGGGFL-GTIFISLSVMLVEHW- 587
K+MW A A ++A + ++ + +W+++ + LG L GT+ + +H+
Sbjct: 820 KLMWAACACTCGAFLAIAFEVV---GKKKWMAITITGLGIPILVGTLASMCYFVFRQHFG 990
Query: 588 -LRKSSWRKKRKESGDGTAE-------SDKESEDSDFQSSY 620
+ S R+ ++ SG + SD + SD + Y
Sbjct: 991 IFQSDSQRRIKRASGSKSFSWSYSANISDIDEYTSDIEKIY 1113
>TC81984 homologue to GP|12321945|gb|AAG51002.1 ankyrin-like protein;
93648-91299 {Arabidopsis thaliana}, partial (22%)
Length = 649
Score = 67.8 bits (164), Expect = 1e-11
Identities = 34/113 (30%), Positives = 68/113 (60%)
Frame = +3
Query: 487 SAFKVFAISNIIALFTSLSVVIVLVSIIPFRRKPQTILLTIAHKVMWVAVAFMGTGYVAA 546
SAFK+F I N IALFTSL+VV+V ++++ K + ++ + +K+MW+A ++A+
Sbjct: 15 SAFKIFFIFNAIALFTSLAVVVVQITLVRGETKAEKRVVEVINKLMWLASVCTSVAFIAS 194
Query: 547 TWVILPHNQEMQWLSVVLLALGGGFLGTIFISLSVMLVEHWLRKSSWRKKRKE 599
+++++ E W ++++ +GG + + +++ +V R S RKK K+
Sbjct: 195 SYIVVGRRNE--WAAILVTVVGGVIISGVIGTMTYYVVRS-KRTRSMRKKEKQ 344
>TC85019 weakly similar to GP|9293890|dbj|BAB01793.1
emb|CAB70981.1~gene_id:MVE11.3~similar to unknown
protein {Arabidopsis thaliana}, partial (10%)
Length = 663
Score = 60.5 bits (145), Expect = 2e-09
Identities = 43/138 (31%), Positives = 71/138 (51%), Gaps = 8/138 (5%)
Frame = +2
Query: 413 KQKHHHNKGKIENVNHTKRKHYHEMHKEALLNARN-------TIVLVAVLIATVTFAAGI 465
K H +K N T + + E HK L +N + +LVA LIAT+TFAA I
Sbjct: 221 KLDHPLHKDVKNNDGKTAWQVFKEEHKTLLEEGKNWMKDTSNSCMLVATLIATITFAAAI 400
Query: 466 SPPGGVYQEGPKKGISMAGETSAFKVFAISNIIALFTSLSVVIVLVSIIPFRRKPQTILL 525
+ PGG Q+ KGI + F +F +S+ +ALF+S+ +++ +SII R + ++
Sbjct: 401 TVPGGNNQD---KGIPIFLSDKTFMLFIVSDALALFSSMVSLLMFLSIIHGRYAKEDFVV 571
Query: 526 TIAHK-VMWVAVAFMGTG 542
+ + ++ +A F G
Sbjct: 572 ALPKRLILGMAALFFAVG 625
>CA921489 weakly similar to PIR|T47566|T475 hypothetical protein F24B22.30 -
Arabidopsis thaliana, partial (10%)
Length = 809
Score = 58.9 bits (141), Expect = 6e-09
Identities = 39/137 (28%), Positives = 73/137 (52%), Gaps = 6/137 (4%)
Frame = -3
Query: 436 EMHKEALLNARNTIVLVAVLIATVTFAAGISPPGGVYQEGPKKGISMAGETSAFKVFAIS 495
E K + + N+ +LVA LIAT+ FAA I+ PGG Q+ KGI + + F VF +S
Sbjct: 591 EEGKNWMKDTSNSCMLVATLIATIAFAAAITVPGGNNQD---KGIPIFLSDNTFMVFVVS 421
Query: 496 NIIALFTSLSVVIVLVSIIPFRRKPQTILLTIAHKV------MWVAVAFMGTGYVAATWV 549
+ +ALF+S++ +++ ++I+ R + ++ + ++ ++ AV + AA +
Sbjct: 420 DALALFSSMASLLMFLAILNARYTEEDFMMALPERLILGMASLFFAVVTTMVAFGAALSM 241
Query: 550 ILPHNQEMQWLSVVLLA 566
+L + + LLA
Sbjct: 240 LLKERLTWAPIPIALLA 190
>TC88966 similar to GP|21305941|gb|AAM45816.1 NADH dehydrogenase subunit 2
{Neoheterandria umbratilis}, partial (4%)
Length = 915
Score = 53.9 bits (128), Expect = 2e-07
Identities = 51/164 (31%), Positives = 79/164 (48%), Gaps = 20/164 (12%)
Frame = -2
Query: 440 EALLNARNTIVLVAVLIATVTFAAGISPPGGVYQ-------EGPKKGISMAGET------ 486
E L + +++I L A LIAT+TF+ +PPGGV Q E K IS T
Sbjct: 821 EWLKDMKSSISLTASLIATLTFSLATNPPGGVVQASVGDSNECGKILISTINTTICVGEA 642
Query: 487 -------SAFKVFAISNIIALFTSLSVVIVLVSIIPFRRKPQTILLTIAHKVMWVAVAFM 539
+ F I N I SLSV++VLVS IP K LL++ M + ++ +
Sbjct: 641 ILATRSHDKYLAFLICNTICFIASLSVILVLVSGIPIDNKFSMWLLSMG---MSIILSSL 471
Query: 540 GTGYVAATWVILPHNQEMQWLSVVLLALGGGFLGTIFISLSVML 583
Y+ A ++ P++ W S+ G GF ++F+ ++V+L
Sbjct: 470 ALTYLFAANMVTPNS---IWNSL----SGNGFGISLFVWIAVVL 360
>TC84356
Length = 670
Score = 45.8 bits (107), Expect = 5e-05
Identities = 38/130 (29%), Positives = 56/130 (42%), Gaps = 21/130 (16%)
Frame = +3
Query: 446 RNTIVLVAVLIATVTFAAGISPPGGVYQEGPK---------------------KGISMAG 484
+ +I L A LIAT+TF+ +PPGGV Q + I
Sbjct: 114 KGSISLTASLIATLTFSLATNPPGGVVQASLDDSNYCSTILNTRMVNTTICLGEAILATR 293
Query: 485 ETSAFKVFAISNIIALFTSLSVVIVLVSIIPFRRKPQTILLTIAHKVMWVAVAFMGTGYV 544
+ F I N SLSV+ VLVS IP K LL+I VM + ++ + Y+
Sbjct: 294 LKDDYLAFLICNTFCFIASLSVIFVLVSGIPINNKFLIWLLSI---VMSIILSGLALTYL 464
Query: 545 AATWVILPHN 554
A ++ P++
Sbjct: 465 FAAGMVTPNS 494
>TC82204 similar to GP|21537142|gb|AAM61483.1 unknown {Arabidopsis
thaliana}, partial (15%)
Length = 522
Score = 38.5 bits (88), Expect = 0.008
Identities = 25/93 (26%), Positives = 48/93 (50%), Gaps = 1/93 (1%)
Frame = +1
Query: 61 DMVSAENKNMETPIHEACRQENVKVLMLLLEVNPTAACKL-NPTCKSAFLVACSHGHLDL 119
D++S +N + ETP++ A +V L+++ K+ + + +AF VA GHLD+
Sbjct: 163 DVMSLQNDHGETPLYIAAHNNLKEVFTFLIKLCDFEVLKIRSKSDMNAFHVAAKRGHLDI 342
Query: 120 VNLLLNLSEIVGQEVAGFDQACFHVAAVRGHTE 152
V +L+ V + + + + AAV+ H +
Sbjct: 343 VREILSAWPAVCKLCDSTNTSPLYAAAVQDHLD 441
Score = 37.7 bits (86), Expect = 0.013
Identities = 26/96 (27%), Positives = 50/96 (52%), Gaps = 2/96 (2%)
Frame = +1
Query: 33 DDTFSAPLHLASKYGCIEMVSEIVKLCP-DMVSAENKNMETPIHEACRQENVKVLMLLLE 91
+D PL++A+ E+ + ++KLC +++ +K+ H A ++ ++ ++ +L
Sbjct: 181 NDHGETPLYIAAHNNLKEVFTFLIKLCDFEVLKIRSKSDMNAFHVAAKRGHLDIVREILS 360
Query: 92 VNPTAACKL-NPTCKSAFLVACSHGHLDLVNLLLNL 126
P A CKL + T S A HLD VN +L++
Sbjct: 361 AWP-AVCKLCDSTNTSPLYAAAVQDHLDGVNAILDV 465
>BG644888 weakly similar to PIR|F85043|F850 hypothetical protein AT4g03440
[imported] - Arabidopsis thaliana, partial (2%)
Length = 727
Score = 38.5 bits (88), Expect = 0.008
Identities = 17/56 (30%), Positives = 34/56 (60%)
Frame = +1
Query: 35 TFSAPLHLASKYGCIEMVSEIVKLCPDMVSAENKNMETPIHEACRQENVKVLMLLL 90
T + LH+A+ YG ++V+ +++ P ++ NKN ++P+H A R ++ + LL
Sbjct: 292 TKNTVLHIAASYGNNDIVNLVIEHSPKLLFTFNKNNDSPLHVAARGGHISTVKTLL 459
>TC82160
Length = 839
Score = 38.1 bits (87), Expect = 0.010
Identities = 29/93 (31%), Positives = 44/93 (47%)
Frame = +3
Query: 492 FAISNIIALFTSLSVVIVLVSIIPFRRKPQTILLTIAHKVMWVAVAFMGTGYVAATWVIL 551
F I N I SLSV+++LVS IP K LL+I M + + + Y+ A ++
Sbjct: 246 FLICNTICFIASLSVILLLVSGIPINNKFSMWLLSIG---MSIVITSLALTYLFAAKMVN 416
Query: 552 PHNQEMQWLSVVLLALGGGFLGTIFISLSVMLV 584
PH+ W S+ G + I I L V ++
Sbjct: 417 PHS---IWSSLPGTGFGVALIVWIAIVLLVYII 506
>BG587066 similar to PIR|E84725|E84 ankyrin-like protein [imported] -
Arabidopsis thaliana, partial (32%)
Length = 656
Score = 35.0 bits (79), Expect(2) = 0.017
Identities = 25/81 (30%), Positives = 36/81 (43%)
Frame = +2
Query: 72 TPIHEACRQENVKVLMLLLEVNPTAACKLNPTCKSAFLVACSHGHLDLVNLLLNLSEIVG 131
T +H A Q ++ V+ LLLE + A K+ A GHL++V LL+ G
Sbjct: 32 TALHTAATQGHIDVVNLLLESDSNLAKIARNNGKTVLHSAARMGHLEVVKALLSKDPSTG 211
Query: 132 QEVAGFDQACFHVAAVRGHTE 152
Q H+ AV+G E
Sbjct: 212 FRTDKKGQTALHM-AVKGQNE 271
Score = 34.3 bits (77), Expect = 0.15
Identities = 29/96 (30%), Positives = 46/96 (47%), Gaps = 3/96 (3%)
Frame = +2
Query: 31 KTDDTFSAPLHLASKYGCIEMVSEIVKLCPDMVSAENKNMETPIHEAC---RQENVKVLM 87
+TD LH+A K E++ E+VK P +++ E+ T +H A R +NV+ L+
Sbjct: 215 RTDKKGQTALHMAVKGQNEEILLELVKPDPAVLNLEDNKGNTALHIAAKKGRNQNVRRLL 394
Query: 88 LLLEVNPTAACKLNPTCKSAFLVACSHGHLDLVNLL 123
+N A K T VA G DLV+++
Sbjct: 395 STEGININATNKAGET---PLDVAEKFGSPDLVSIM 493
Score = 21.2 bits (43), Expect(2) = 0.017
Identities = 8/17 (47%), Positives = 12/17 (70%)
Frame = +3
Query: 168 KGIQHCIMLVIKDTLRL 184
K IQHCI+L + +R+
Sbjct: 330 KEIQHCILLQRRAVIRM 380
>BQ138717 similar to GP|12321945|gb ankyrin-like protein; 93648-91299
{Arabidopsis thaliana}, partial (36%)
Length = 676
Score = 37.4 bits (85), Expect = 0.017
Identities = 34/149 (22%), Positives = 64/149 (42%)
Frame = +2
Query: 6 FDAIKKNDMITFSSIVKEREGILNQKTDDTFSAPLHLASKYGCIEMVSEIVKLCPDMVSA 65
F A +K + ++K K + + PLH+A+ G +V ++ P +
Sbjct: 191 FTAAEKGHLDVVKELLKHSTLQTVSKKNRSGFDPLHIAASQGHHAIVQVLLDYDPSLSKT 370
Query: 66 ENKNMETPIHEACRQENVKVLMLLLEVNPTAACKLNPTCKSAFLVACSHGHLDLVNLLLN 125
+ TP+ A + +V+V+ LL + + K+A +A GH ++V LL+
Sbjct: 371 IGPSNATPLITAATRGHVEVVNELLSKDGSLLEIARSNGKNALHLAARPGHTEIVKALLS 550
Query: 126 LSEIVGQEVAGFDQACFHVAAVRGHTESC 154
+ + Q H+ AV+G +SC
Sbjct: 551 KDPQLARRTDKKGQTALHM-AVKG--QSC 628
Score = 35.8 bits (81), Expect = 0.051
Identities = 26/105 (24%), Positives = 51/105 (47%), Gaps = 1/105 (0%)
Frame = +2
Query: 49 IEMVSEIVKLCPDMVSAENKNMETPIHEACRQENVKVLMLLLEVNPT-AACKLNPTCKSA 107
+++ +EI ++ +V+ EN+ ET + A + ++ V+ LL+ + K N +
Sbjct: 113 VDLNAEIAEVRALVVNEENELGETALFTAAEKGHLDVVKELLKHSTLQTVSKKNRSGFDP 292
Query: 108 FLVACSHGHLDLVNLLLNLSEIVGQEVAGFDQACFHVAAVRGHTE 152
+A S GH +V +LL+ + + + + AA RGH E
Sbjct: 293 LHIAASQGHHAIVQVLLDYDPSLSKTIGPSNATPLITAATRGHVE 427
>TC85563 similar to GP|14279688|gb|AAK58690.1 receptor-like kinase
Xa21-binding protein 3 {Oryza sativa}, partial (71%)
Length = 2214
Score = 37.0 bits (84), Expect = 0.023
Identities = 27/121 (22%), Positives = 55/121 (45%)
Frame = +3
Query: 5 FFDAIKKNDMITFSSIVKEREGILNQKTDDTFSAPLHLASKYGCIEMVSEIVKLCPDMVS 64
F A+ +M S+V++ G+L+ +PLHLA+ G IE++S ++ V
Sbjct: 195 FASAVVNGEMEIVESMVEDDVGVLDSTVGRAKLSPLHLAAANGRIEVLSMLLNR-NVKVD 371
Query: 65 AENKNMETPIHEACRQENVKVLMLLLEVNPTAACKLNPTCKSAFLVACSHGHLDLVNLLL 124
N++ +TP+ A + L++ + ++ A +GH+D + +L
Sbjct: 372 VLNRHKQTPLMLAVMHGKTGCMEKLIQAGANILMFDSIRRRTCLHYAAYYGHVDSLKAIL 551
Query: 125 N 125
+
Sbjct: 552 S 554
>CB894501 weakly similar to PIR|F86279|F86 hypothetical protein AAF43949.1
[imported] - Arabidopsis thaliana, partial (26%)
Length = 812
Score = 36.6 bits (83), Expect = 0.030
Identities = 21/70 (30%), Positives = 35/70 (50%)
Frame = +3
Query: 21 VKEREGILNQKTDDTFSAPLHLASKYGCIEMVSEIVKLCPDMVSAENKNMETPIHEACRQ 80
VK REG+ PLH+A++ G ++V + + +CP + ET +H A +
Sbjct: 231 VKGREGL----------TPLHIATQTGNFDLVVKFLFVCPGCIEDVTVRSETALHIAVKY 380
Query: 81 ENVKVLMLLL 90
+ VL +LL
Sbjct: 381 KQFHVLEILL 410
Score = 35.0 bits (79), Expect = 0.086
Identities = 16/57 (28%), Positives = 33/57 (57%)
Frame = +3
Query: 270 INMATLSYISQFQAVAIRTKVDINTRNNEGLTALDILDQAMDNAENRQLQAIFIRDG 326
++M+ L+ Q + I + +DIN +N + TALDI++Q + +++ + I+ G
Sbjct: 498 LHMSVLNSFPQAVGLLIDSNIDINAKNLDEQTALDIVEQIQSQVYSAEMKDMLIKAG 668
Score = 34.3 bits (77), Expect = 0.15
Identities = 24/107 (22%), Positives = 47/107 (43%)
Frame = +3
Query: 39 PLHLASKYGCIEMVSEIVKLCPDMVSAENKNMETPIHEACRQENVKVLMLLLEVNPTAAC 98
PLH+A+ G +EI++L P N+ +PIH A + + +++ +++N
Sbjct: 51 PLHIAAASGHTSFATEIMRLKPSFAWKLNEYGLSPIHLALQNKYHRMVCRFVDINKDLVR 230
Query: 99 KLNPTCKSAFLVACSHGHLDLVNLLLNLSEIVGQEVAGFDQACFHVA 145
+ +A G+ DLV L + ++V + H+A
Sbjct: 231 VKGREGLTPLHIATQTGNFDLVVKFLFVCPGCIEDVTVRSETALHIA 371
Score = 33.9 bits (76), Expect = 0.19
Identities = 21/87 (24%), Positives = 41/87 (46%)
Frame = +3
Query: 38 APLHLASKYGCIEMVSEIVKLCPDMVSAENKNMETPIHEACRQENVKVLMLLLEVNPTAA 97
+P+HLA + MV V + D+V + + TP+H A + N +++ L V P
Sbjct: 150 SPIHLALQNKYHRMVCRFVDINKDLVRVKGREGLTPLHIATQTGNFDLVVKFLFVCPGCI 329
Query: 98 CKLNPTCKSAFLVACSHGHLDLVNLLL 124
+ ++A +A + ++ +LL
Sbjct: 330 EDVTVRSETALHIAVKYKQFHVLEILL 410
>BQ146254 weakly similar to GP|11994366|dbj gene_id:MDC16.7~unknown protein
{Arabidopsis thaliana}, partial (19%)
Length = 408
Score = 35.8 bits (81), Expect = 0.051
Identities = 18/44 (40%), Positives = 26/44 (58%)
Frame = +3
Query: 428 HTKRKHYHEMHKEALLNARNTIVLVAVLIATVTFAAGISPPGGV 471
HTK ++ + L R +VA+ IAT+TF G++PPGGV
Sbjct: 147 HTKTRN------DWLEKMRGNFSVVAIFIATITFQMGLNPPGGV 260
>AW693918 similar to GP|9759271|dbj contains similarity to
ankyrin~gene_id:MRI1.10 {Arabidopsis thaliana}, partial
(21%)
Length = 375
Score = 35.4 bits (80), Expect = 0.066
Identities = 21/80 (26%), Positives = 39/80 (48%)
Frame = +3
Query: 72 TPIHEACRQENVKVLMLLLEVNPTAACKLNPTCKSAFLVACSHGHLDLVNLLLNLSEIVG 131
+P+H + + +++ LLLE + N ++A + AC HGH ++V L+ +
Sbjct: 90 SPLHYSAAHGHHEIVYLLLESGVDINLR-NYRGQTALMQACQHGHWEVVQTLIIFKANIH 266
Query: 132 QEVAGFDQACFHVAAVRGHT 151
+ H+AA+ GHT
Sbjct: 267 KTDYLNGGTALHLAALNGHT 326
>AW736367 weakly similar to PIR|T01879|T01 hypothetical protein F8M12.17 -
Arabidopsis thaliana, partial (4%)
Length = 463
Score = 35.4 bits (80), Expect = 0.066
Identities = 23/86 (26%), Positives = 42/86 (48%)
Frame = +3
Query: 38 APLHLASKYGCIEMVSEIVKLCPDMVSAENKNMETPIHEACRQENVKVLMLLLEVNPTAA 97
+P+HLA + MV +V + D+V + K TP+H A + ++L LL P +
Sbjct: 204 SPIHLAIQNNHRSMVFCLVDMNKDLVRLKGKEGMTPLHFASQNGEDEILAKLLFACPESI 383
Query: 98 CKLNPTCKSAFLVACSHGHLDLVNLL 123
+ K+A +A + +++ LL
Sbjct: 384 EDVTVRGKTALHIAVQNNQYEVLKLL 461
Score = 35.4 bits (80), Expect = 0.066
Identities = 24/107 (22%), Positives = 52/107 (48%)
Frame = +3
Query: 46 YGCIEMVSEIVKLCPDMVSAENKNMETPIHEACRQENVKVLMLLLEVNPTAACKLNPTCK 105
YG I++ S I+++ N+ ++TP+H A + ++ + ++ + P+ A KLN
Sbjct: 39 YGVIQVNSSILEIIDS-----NEFVKTPLHIAATRGHLPFAIEVMNLKPSFALKLNQEGL 203
Query: 106 SAFLVACSHGHLDLVNLLLNLSEIVGQEVAGFDQACFHVAAVRGHTE 152
S +A + H +V L+++++ + + H A+ G E
Sbjct: 204 SPIHLAIQNNHRSMVFCLVDMNKDLVRLKGKEGMTPLHFASQNGEDE 344
Score = 32.3 bits (72), Expect = 0.56
Identities = 15/62 (24%), Positives = 32/62 (51%)
Frame = +3
Query: 32 TDDTFSAPLHLASKYGCIEMVSEIVKLCPDMVSAENKNMETPIHEACRQENVKVLMLLLE 91
+++ PLH+A+ G + E++ L P N+ +PIH A + + ++ L++
Sbjct: 84 SNEFVKTPLHIAATRGHLPFAIEVMNLKPSFALKLNQEGLSPIHLAIQNNHRSMVFCLVD 263
Query: 92 VN 93
+N
Sbjct: 264 MN 269
>TC79077 similar to GP|17529252|gb|AAL38853.1 unknown protein {Arabidopsis
thaliana}, partial (38%)
Length = 2200
Score = 35.0 bits (79), Expect = 0.086
Identities = 28/108 (25%), Positives = 46/108 (41%), Gaps = 15/108 (13%)
Frame = +3
Query: 32 TDDTFSAPLHLASKYGCIEMVSEIVKLCPDMVSAENKNMETPIHEACR----------QE 81
TD + LH+A+ G + V+ ++ L P ++S N ET +H+A
Sbjct: 1365 TDHQGNTALHVAASRGQLSAVNALISLFPTLISHRNNAGETFLHKAVSGFQTHAFRRLDR 1544
Query: 82 NVKVLMLLLEVN-----PTAACKLNPTCKSAFLVACSHGHLDLVNLLL 124
V++L LL N K N + + + H+DLV LL+
Sbjct: 1545 QVELLKKLLSTNHFHVEEIINIKNNDGRTALHMAIIGNIHIDLVQLLM 1688
Database: MTGI
Posted date: Oct 22, 2004 3:39 PM
Number of letters in database: 27,044,181
Number of sequences in database: 36,976
Lambda K H
0.330 0.140 0.426
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 18,908,820
Number of Sequences: 36976
Number of extensions: 285761
Number of successful extensions: 2485
Number of sequences better than 10.0: 62
Number of HSP's better than 10.0 without gapping: 2392
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 2468
length of query: 626
length of database: 9,014,727
effective HSP length: 102
effective length of query: 524
effective length of database: 5,243,175
effective search space: 2747423700
effective search space used: 2747423700
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 15 ( 7.1 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 40 (21.9 bits)
S2: 61 (28.1 bits)
Medicago: description of AC138130.17


