

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
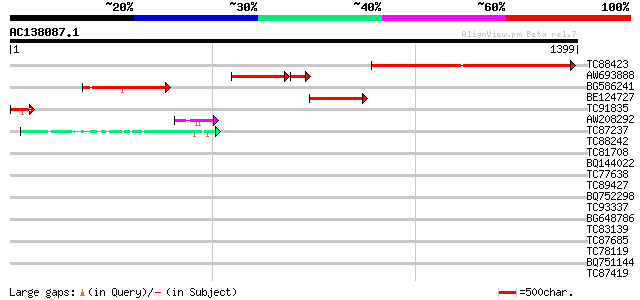
Query= AC138087.1 + phase: 0
(1399 letters)
Database: MTGI
36,976 sequences; 27,044,181 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
TC88423 similar to GP|9758777|dbj|BAB09075.1 contains similarity... 713 0.0
AW693888 weakly similar to GP|9758777|db contains similarity to ... 227 6e-77
BG586241 similar to GP|9758776|dbj dbj|BAA90625.1~gene_id:MNJ7.7... 271 2e-72
BE124727 weakly similar to GP|9758777|db contains similarity to ... 228 1e-59
TC91835 similar to GP|9758776|dbj|BAB09074.1 dbj|BAA90625.1~gene... 79 1e-14
AW208292 76 8e-14
TC87237 similar to GP|160409|gb|AAA29651.1|| mature-parasite-inf... 49 2e-05
TC88242 similar to GP|15100049|gb|AAK84220.1 transcription facto... 39 0.011
TC81708 similar to GP|12083536|gb|AAG48845.1 hypothetical protei... 37 0.041
BQ144022 homologue to GP|22831315|dbj hypothetical protein~predi... 37 0.054
TC77638 similar to GP|20161455|dbj|BAB90379. ankyrin-like protei... 37 0.070
TC89427 weakly similar to GP|4557063|gb|AAD22502.1| expressed pr... 36 0.092
BQ752298 similar to GP|11595639|em conserved hypothetical protei... 32 0.14
TC93337 weakly similar to GP|21240660|gb|AAM44371.1 hypothetical... 35 0.16
BG648786 weakly similar to SP|P10495|GRP1_ Glycine-rich cell wal... 34 0.45
TC83139 33 1.0
TC87685 weakly similar to PIR|T02289|T02289 probable polygalactu... 33 1.0
TC78119 similar to GP|13605807|gb|AAK32889.1 AT3g13930/MDC16_5 {... 33 1.0
BQ751144 weakly similar to PIR|PQ0479|PQ04 pistil extensin-like ... 33 1.0
TC87419 similar to GP|12006165|gb|AAG44765.1 biotin carboxyl car... 32 1.7
>TC88423 similar to GP|9758777|dbj|BAB09075.1 contains similarity to unknown
protein~dbj|BAA90625.1~gene_id:MNJ7.8 {Arabidopsis
thaliana}, partial (15%)
Length = 1640
Score = 713 bits (1840), Expect = 0.0
Identities = 374/508 (73%), Positives = 409/508 (79%), Gaps = 6/508 (1%)
Frame = +1
Query: 894 AIQRTELYEYSKVLGNSQFILLPFQPYKLIYAYMLAEVGKVSDSLKYCQAVLKSLKTGRA 953
AIQRTELYEYSK+LGNSQF+L FQPYKLIYA+MLAEVGKVSDSLKYCQAVLKSLKTGRA
Sbjct: 1 AIQRTELYEYSKLLGNSQFVLHSFQPYKLIYAHMLAEVGKVSDSLKYCQAVLKSLKTGRA 180
Query: 954 PEVETWKQRLSSLEERIRTHQQGGYAANLAPGKLVGKLLNFFDSTAHRVVGGLPPPAP-S 1012
PEVETWKQ + +LEERIRTHQQGGYAANLAP KLVGKLLNFFDSTAHRVVGGLPPPAP S
Sbjct: 181 PEVETWKQLVLALEERIRTHQQGGYAANLAPAKLVGKLLNFFDSTAHRVVGGLPPPAPTS 360
Query: 1013 SQGNVHGNEQNYQSGAHRVSNSQSTMAMSSLVPSGSMEPNGEWTADNNRMT-KSNRSVSE 1071
SQ VHG+EQ+YQ A RVS SQSTMAMSSLVPS S+EP EWTADNNRM K NRSVSE
Sbjct: 361 SQATVHGSEQHYQHMAPRVSTSQSTMAMSSLVPSASLEPISEWTADNNRMAAKPNRSVSE 540
Query: 1072 PDFGRSPRQET-SHDAQGKA--SEGTSRFSRFSFGSQLLQKTMGLVLKPRPGKQAKLGEK 1128
PD GRSPRQE+ S DAQGK S G SRFSRF FGSQLLQKT+GLVL R GKQAKLGEK
Sbjct: 541 PDIGRSPRQESPSPDAQGKVQVSGGASRFSRFGFGSQLLQKTVGLVL--RSGKQAKLGEK 714
Query: 1129 NKFYYDENLKRWVEEGAAPPAEETALPPPP-TTAAFQNGLTEYNLQSALKTEGPPSKEGS 1187
NKFYYDE LKRWVEEGA PAEE ALPPPP TTAAFQNG +YNL+SALKTEG E S
Sbjct: 715 NKFYYDEKLKRWVEEGAEVPAEEAALPPPPPTTAAFQNGSADYNLKSALKTEGLTPNEFS 894
Query: 1188 DLKTSNPELTPGIPPIPPGTNHFSARGRVGIRSRYVDTFNQGGGNSANLFQSPSVPSAKP 1247
+TS+PEL+PG+PPIPP +N FSAR R+G+RSRYVDTFNQ GG+SANLFQSPSV S KP
Sbjct: 895 STRTSSPELSPGMPPIPPSSNQFSARSRLGVRSRYVDTFNQNGGSSANLFQSPSVQSVKP 1074
Query: 1248 VVAANAKFFIPTPAPSSNEQTMEAIEENNQEDDLAYENPSTSYRNDWSFQSPKHASASTW 1307
+ ANAKFFIP P PSS+EQ MEAI E+N ED A ENPSTS NDWS+ PKHA T
Sbjct: 1075 ALPANAKFFIPAPVPSSSEQNMEAIAESNLEDSAANENPSTSSTNDWSYHPPKHAQTMTM 1254
Query: 1308 QRCPSMGNFANHEAVVSGSNSRSPHSRRTVSWGGSTDVTYSPTKMREIMPLGEALGMPPS 1367
QR PS GN + + G+ S HSRRT SW GS + ++SP KM EI P G ALGMPPS
Sbjct: 1255 QRFPSAGNISK-QGQTDGNESHFSHSRRTASWSGSFNDSFSPPKMGEIKPSGAALGMPPS 1431
Query: 1368 TYMSDDISSMRTSMKSGNFGEDLHEVDL 1395
+M D S M+ +SG+FGEDL EV+L
Sbjct: 1432 AFMPDPSSLMQGPTRSGSFGEDLQEVEL 1515
>AW693888 weakly similar to GP|9758777|db contains similarity to unknown
protein~dbj|BAA90625.1~gene_id:MNJ7.8 {Arabidopsis
thaliana}, partial (7%)
Length = 611
Score = 227 bits (579), Expect(2) = 6e-77
Identities = 113/144 (78%), Positives = 126/144 (87%)
Frame = +1
Query: 547 GKLIILKDSSLSSSTYGSQGAAQGSVSVLNLMEAVSGSIGSSSIGNGAGDYFRALGQQSI 606
GKLII+KD S +++YGSQ + QGS+SVLNLMEAV+GS S +IGN GDYFRAL QQS
Sbjct: 1 GKLIIMKDPSALTASYGSQDSVQGSISVLNLMEAVTGSNNSLTIGNATGDYFRALSQQSF 180
Query: 607 PGPLVGGSVGSKELNKWIDERIAHCGSPDMDYKKSERMRLLLSLLKIACQYYGKLRSPFG 666
PGPLVGGSVGSKEL KW+DERIA C SPDMDYKK ER+RLLLSLLKIACQ+YGKLRSPFG
Sbjct: 181 PGPLVGGSVGSKELYKWLDERIARCESPDMDYKKGERLRLLLSLLKIACQHYGKLRSPFG 360
Query: 667 TDTILKDNDTPGSAVAKLFASAKM 690
TDTILK+ND P SAVAKLFASAK+
Sbjct: 361 TDTILKENDAPESAVAKLFASAKV 432
Score = 80.5 bits (197), Expect(2) = 6e-77
Identities = 38/48 (79%), Positives = 42/48 (87%)
Frame = +2
Query: 694 EYGVLSHCLQNLPSEAQMRATASEVQNLLVSGKKKEALQYAQEGQLWG 741
+YG+ SHCLQN PSE QM+A ASE+QNLLVSGKK EALQ AQEGQLWG
Sbjct: 452 QYGMPSHCLQNFPSEEQMKAIASEMQNLLVSGKKMEALQRAQEGQLWG 595
>BG586241 similar to GP|9758776|dbj dbj|BAA90625.1~gene_id:MNJ7.7~similar to
unknown protein {Arabidopsis thaliana}, partial (6%)
Length = 852
Score = 271 bits (692), Expect = 2e-72
Identities = 136/230 (59%), Positives = 167/230 (72%), Gaps = 13/230 (5%)
Frame = +3
Query: 181 QYAQTYNRDSNTEVKLGNEIPSDGMNASVDYVQYQEGQ-SYDASARNSTSGEDVNSSQYW 239
QY D++ + G + DG+N SV+YVQYQEG +YDAS+ +G+D++SSQ W
Sbjct: 3 QYQGVQGYDTSFDNHTGKQ--GDGLNTSVNYVQYQEGGGAYDASSNLHNNGQDLSSSQNW 176
Query: 240 ESLYPGWKYDYNTGQWYQVDEHNATAATQGSSEVNTA-----------EVSYMQQTAQSA 288
E LYPGWKYD+ TGQWYQ++++NAT +Q +SE NTA E+SY+QQ AQS
Sbjct: 177 EDLYPGWKYDHITGQWYQIEDYNATTTSQQTSEANTAVDWAAASDGKTEISYLQQAAQS- 353
Query: 289 VAGTLAESAATETVPSWNQVSQGNNGYPEHMIFDPQYPGWYYDTIAQEWRSLETYHSSIQ 348
VAGTLAE+ TE+V SWNQVSQGNNGY EHM+FDPQYP WYYDTIAQEWRSL TY+SS+Q
Sbjct: 354 VAGTLAETGTTESVSSWNQVSQGNNGYLEHMVFDPQYPDWYYDTIAQEWRSLATYNSSVQ 533
Query: 349 YAVQGHGNGHASSGTFSHN-DNSLYRDYGQVGYYESQGVGSQAANNNWSG 397
+V G NGH S+ T S N DNSLY +Y Q G + SQGVGSQA N +WSG
Sbjct: 534 SSVHGLQNGHTSTSTSSFNDDNSLYSEYSQAGNHVSQGVGSQAVNGSWSG 683
Score = 40.8 bits (94), Expect = 0.004
Identities = 25/62 (40%), Positives = 33/62 (52%), Gaps = 1/62 (1%)
Frame = +2
Query: 377 QVGYYE-SQGVGSQAANNNWSGSYGINHQQDLDRHTTDTATKSGGSAYGGNQQFDHSFGS 435
Q G Y+ S GVGSQA N +WSGS+G+N A GGS G+Q + S+
Sbjct: 698 QAGNYDGSHGVGSQAVNGSWSGSHGVNQ-----------AGNYGGSQGVGSQAVNGSWSG 844
Query: 436 SN 437
S+
Sbjct: 845 SH 850
>BE124727 weakly similar to GP|9758777|db contains similarity to unknown
protein~dbj|BAA90625.1~gene_id:MNJ7.8 {Arabidopsis
thaliana}, partial (10%)
Length = 439
Score = 228 bits (581), Expect = 1e-59
Identities = 111/144 (77%), Positives = 126/144 (87%), Gaps = 1/144 (0%)
Frame = +3
Query: 741 GPALVLASQLGEKFYVDTVKQMALRQLVAGSPLRTLCLLIAGQPAEVF-SSDSSNSGDPS 799
GPALVLASQLGE+FYVDTV+QMALRQ+VAGSPLRTLCLLIAG+P +VF + ++S SG P
Sbjct: 3 GPALVLASQLGEQFYVDTVRQMALRQVVAGSPLRTLCLLIAGRPNDVFPTEETSISGHPG 182
Query: 800 AFNMPQNPAQFGSSGMLDDWEENLAVITSNRTKDDELVIIHLGDCLWKERSEITAAHICY 859
A MPQ Q GS+ ML+DWEENLAVIT+NRTK DELV++HLGDCLWKE+ EITAAHICY
Sbjct: 183 AVRMPQQSEQAGSNDMLEDWEENLAVITANRTKGDELVMMHLGDCLWKEKREITAAHICY 362
Query: 860 LIAEANFESYSDSARLCLIGADHW 883
LIAE NF SYSD+ RLCLIGADHW
Sbjct: 363 LIAEVNFSSYSDATRLCLIGADHW 434
>TC91835 similar to GP|9758776|dbj|BAB09074.1
dbj|BAA90625.1~gene_id:MNJ7.7~similar to unknown protein
{Arabidopsis thaliana}, partial (2%)
Length = 618
Score = 79.0 bits (193), Expect = 1e-14
Identities = 45/71 (63%), Positives = 50/71 (70%), Gaps = 10/71 (14%)
Frame = +2
Query: 1 MASNPPFHVEDQDDEDFFDKLVE--DDVGNV--------NDEANDSDDVKAFSNLSIGGD 50
MASNPPF +EDQ DEDFFDKLVE DDVG V NDE NDSDD AF+NLSI
Sbjct: 404 MASNPPFQMEDQTDEDFFDKLVEDDDDVGPVKPVVIKSGNDEGNDSDDANAFANLSI--- 574
Query: 51 DADVNASAFEN 61
+DV+A AF+N
Sbjct: 575 -SDVDAGAFDN 604
>AW208292
Length = 373
Score = 76.3 bits (186), Expect = 8e-14
Identities = 49/126 (38%), Positives = 62/126 (48%), Gaps = 19/126 (15%)
Frame = +1
Query: 408 DRHTTDTATKSGGS-AYGGNQQFDHSFGSSNSVNKNQQNASSSFGSVPLYNK-------- 458
D + T+ +TK G + A GNQQ HS+G+ Q N SSSFGSV L NK
Sbjct: 13 DMYATEASTKIGNNTASSGNQQVHHSYGNQ------QVNTSSSFGSVALNNKGSFEPKAF 174
Query: 459 -----VNHGHGLV-----NGTVEVQRFAPSGNFGQHYNYSNTQFDEQKNISNDYAESHQP 508
+ H NGT + F P G+ Q +NY NT+FDEQK SN +AE+
Sbjct: 175 VPHRDIAHQFNYQDTEFDNGTFAPKTFVPHGDIAQQFNYPNTKFDEQKQFSNVFAENQNS 354
Query: 509 FGYSNQ 514
YS Q
Sbjct: 355 HSYSQQ 372
>TC87237 similar to GP|160409|gb|AAA29651.1|| mature-parasite-infected
erythrocyte surface antigen {Plasmodium falciparum},
partial (2%)
Length = 2007
Score = 48.5 bits (114), Expect = 2e-05
Identities = 101/515 (19%), Positives = 189/515 (36%), Gaps = 22/515 (4%)
Frame = +1
Query: 28 NVNDEANDSDDVKAFSNLSIGGDDADVNASAFENSSGGGSGGEGKERKEEGDVKLDGGNV 87
N D+ N+ + G++ + ++ + G GGE +E + + V
Sbjct: 379 NSKDKENEESNQNGSDAKEQVGENHEQDSKQGTEETNGTEGGEKEEHDKIKEDTSSDNQV 558
Query: 88 QEGSSSGCDGMMDRSDHGMESRNSSGSSADKSNRRSSLDVKE----KDWNAFNVDS--NG 141
Q+G + + + N+S + D ++ SS +E K+ N F ++S N
Sbjct: 559 QDGEKNN-----EAREENYSGDNASSAVVDNKSQESSNKTEEQFDKKEKNEFELESQKNS 723
Query: 142 GAGSESYSDFFSEFGDQNGKGYDHDLNTEVKHANEIPGDQYAQTYNRDSNTEVKLGNEIP 201
+ES + QN +G NE DQ AQT N+ P
Sbjct: 724 NETTESTDSTIT----QNSQG------------NESEKDQ-AQT-----------ENDTP 819
Query: 202 SDGMNASVDYVQYQEGQSYDASARNSTSGEDVNSSQYWESLYPGWKYDYNTGQWYQVDEH 261
+ S + Q QE +N+T+ +DV ++ + +T +
Sbjct: 820 KGSASESDEQKQEQE--------QNNTTKDDVQTTD------TSSQNGNDTTEKQNETSE 957
Query: 262 NATAATQGSSEVNTAEVSYMQQTAQSAVAGTLAESAATETVPSWNQVSQGNNGYPEHMIF 321
+A + + SS +NT + ++S VAG A+S T S ++ GN + E+
Sbjct: 958 DANSKKEDSSALNTTP---NNEDSKSGVAGDQADST---TTTSSSETQDGNTNHGEYKDT 1119
Query: 322 DPQYPGWYYDTIAQEWRSLETYHSSIQYAVQGHGNGHASSGTFSHNDNSLYRDYGQVGYY 381
+ P + + + + E+ SS + + + T S + +D
Sbjct: 1120TNENP----EKNSGQEGTQESGSSSNTFDNKDAASNKVQLTTTSDTSSEQKKDESSSAES 1287
Query: 382 ESQGVGSQAAN---NNWSGSYGINHQQDLDRHTTDTATKSGGSAYGGNQQFDHSFGSSNS 438
+S+ + AN +N + N +D + TT + + G++ N ++ S N+
Sbjct: 1288KSESSQNDNANSGQSNTTSDESANDNKDSSQVTTSSENSAEGNSNTENNSDENQNDSKNN 1467
Query: 439 VNKNQQNASSSFGSV-------PLYNKVNHGHG-LVNGTVEVQRFAPSGNFGQHYNY--- 487
N N +S+ +V K + G N +VE ++ N H +
Sbjct: 1468ENTNDSGNTSNDANVNENQNENAAQTKTSENEGDAQNESVESKK---ENNESAHKDVDNN 1638
Query: 488 --SNTQFDEQKNISNDYAESHQPFGYSNQSYQSGH 520
SN Q +++ D ES G +S H
Sbjct: 1639SNSNDQGSSDTSVTQDDKESRVDLGTLPESNGESH 1743
>TC88242 similar to GP|15100049|gb|AAK84220.1 transcription factor bZIP29
{Arabidopsis thaliana}, partial (39%)
Length = 1930
Score = 39.3 bits (90), Expect = 0.011
Identities = 32/121 (26%), Positives = 55/121 (45%), Gaps = 9/121 (7%)
Frame = +2
Query: 15 EDFFDKLVE-DDVGNVNDEANDS-DDVKAFSNLSIGGDDADVNASAFENSSGGG-SGGEG 71
+D F + D++ +ND+ N + DD +A + GGD +D A + N SG EG
Sbjct: 758 DDLFSAYMNLDNIDAINDDKNAATDDSRASGTKTNGGDSSDNEAESSVNESGDSMQRREG 937
Query: 72 KERKEEGDVKLDGGNVQEGSSSGCDGMMDRSDHGME------SRNSSGSSADKSNRRSSL 125
+R GD+ + + S G ++ +D ++ S G+S D +N SL
Sbjct: 938 NKRSAGGDIAPTTRHYRSVSMDSFIGKLNFNDESLKMPPSPGGLMSPGNSGDGNNAAFSL 1117
Query: 126 D 126
+
Sbjct: 1118E 1120
>TC81708 similar to GP|12083536|gb|AAG48845.1 hypothetical protein {Oryza
sativa}, partial (6%)
Length = 605
Score = 37.4 bits (85), Expect = 0.041
Identities = 36/147 (24%), Positives = 54/147 (36%), Gaps = 14/147 (9%)
Frame = +1
Query: 48 GGDDADVNASAFENSSGGGSGGEGK--ERKEEGDVKLDGG------------NVQEGSSS 93
GGDD + GGG G+ K +K+ G K GG + + G ++
Sbjct: 94 GGDDGGKKNKGEKQKGGGGDWGDEKNSSKKKNGKGKNGGGGFLVKFLGLGKKSKKGGGAA 273
Query: 94 GCDGMMDRSDHGMESRNSSGSSADKSNRRSSLDVKEKDWNAFNVDSNGGAGSESYSDFFS 153
+ G S NS G K + LD E D+ F++ +G +G
Sbjct: 274 DTTNKKKNNGGGGNSNNSKGKDGKKGEK---LDKVEFDFQDFDITPHGKSGKGGN----- 429
Query: 154 EFGDQNGKGYDHDLNTEVKHANEIPGD 180
G+ NGKG N H + G+
Sbjct: 430 --GNGNGKGAPAKGNANNGHGSNNNGN 504
>BQ144022 homologue to GP|22831315|dbj hypothetical protein~predicted by
GlimmerM etc. {Oryza sativa (japonica cultivar-group)},
partial (16%)
Length = 1286
Score = 37.0 bits (84), Expect = 0.054
Identities = 31/122 (25%), Positives = 44/122 (35%)
Frame = +1
Query: 64 GGGSGGEGKERKEEGDVKLDGGNVQEGSSSGCDGMMDRSDHGMESRNSSGSSADKSNRRS 123
GGG G GK RK EG K++ G SG G R + G E + S+ + R
Sbjct: 514 GGGGGRRGKSRK-EGRKKIERGPPAVDRKSG-GGQRARGEKGREGKGSAREAGGSGGRGE 687
Query: 124 SLDVKEKDWNAFNVDSNGGAGSESYSDFFSEFGDQNGKGYDHDLNTEVKHANEIPGDQYA 183
SL + GAG + + S + HD + D+ A
Sbjct: 688 SLPTPKP----------AGAGGRKHKERESARARERSASGGHDGRVDAAGTTRTDADEQA 837
Query: 184 QT 185
+T
Sbjct: 838 RT 843
>TC77638 similar to GP|20161455|dbj|BAB90379. ankyrin-like protein {Oryza
sativa (japonica cultivar-group)}, partial (62%)
Length = 2345
Score = 36.6 bits (83), Expect = 0.070
Identities = 32/155 (20%), Positives = 68/155 (43%)
Frame = +1
Query: 23 EDDVGNVNDEANDSDDVKAFSNLSIGGDDADVNASAFENSSGGGSGGEGKERKEEGDVKL 82
ED+ G++ ++A D ++ ++ S ++S + EGK+ ++EG
Sbjct: 538 EDNPGDLPEDATKGDS-------NVSSEEKSEENSTEKSSEDTKTEDEGKKTEDEGSNTE 696
Query: 83 DGGNVQEGSSSGCDGMMDRSDHGMESRNSSGSSADKSNRRSSLDVKEKDWNAFNVDSNGG 142
+ + +E S+ + D S+ ES ++ S +D+S ++SS + D N
Sbjct: 697 NNKDGEEASTK--ESESDESEKKDESEENNKSDSDESEKKSSDSNETTDSNV-------- 846
Query: 143 AGSESYSDFFSEFGDQNGKGYDHDLNTEVKHANEI 177
E ++ D+N + D N + + +NE+
Sbjct: 847 --EEKVEQSQNKESDENASEKNTDDNAKDQSSNEV 945
>TC89427 weakly similar to GP|4557063|gb|AAD22502.1| expressed protein
{Arabidopsis thaliana}, partial (50%)
Length = 753
Score = 36.2 bits (82), Expect = 0.092
Identities = 21/70 (30%), Positives = 36/70 (51%)
Frame = +1
Query: 10 EDQDDEDFFDKLVEDDVGNVNDEANDSDDVKAFSNLSIGGDDADVNASAFENSSGGGSGG 69
E +DD+D EDD +VNDE +D+D+ GD+ D +A ++ G+GG
Sbjct: 304 ETEDDDD------EDDDDDVNDEDDDNDE-------DFSGDEDDEDADPEDDPVPNGAGG 444
Query: 70 EGKERKEEGD 79
+ +++ D
Sbjct: 445 SDDDDEDDDD 474
>BQ752298 similar to GP|11595639|em conserved hypothetical protein {Neurospora
crassa}, partial (2%)
Length = 834
Score = 32.3 bits (72), Expect(2) = 0.14
Identities = 13/19 (68%), Positives = 14/19 (73%)
Frame = -2
Query: 1122 QAKLGEKNKFYYDENLKRW 1140
+AKLGE N F YD LKRW
Sbjct: 707 KAKLGEANNFVYDPELKRW 651
Score = 21.9 bits (45), Expect(2) = 0.14
Identities = 9/18 (50%), Positives = 10/18 (55%)
Frame = -1
Query: 1144 GAAPPAEETALPPPPTTA 1161
GA +TA PPPP A
Sbjct: 636 GAEATQAKTATPPPPRAA 583
>TC93337 weakly similar to GP|21240660|gb|AAM44371.1 hypothetical protein
{Dictyostelium discoideum}, partial (7%)
Length = 743
Score = 35.4 bits (80), Expect = 0.16
Identities = 48/231 (20%), Positives = 91/231 (38%), Gaps = 12/231 (5%)
Frame = +1
Query: 20 KLVEDDVGNVNDEANDSDDVKAFSNLSIGGDDADVNASAFENSSGGGSG----GEGKERK 75
K+ ED+ D D + GG+D +++ + E EGKE++
Sbjct: 10 KVKEDEESRTRDGGEKEDGER-------GGEDDEMDENDPEKPEVVDRSEEFVDEGKEKE 168
Query: 76 EEGDVKLDGGNVQEGSSSGCDGMMDRSDHGMESRNSSGSSA-DKSNRRSSLDVKEKDWNA 134
EEGD K + + EG+ + + + SSA R +S + +
Sbjct: 169 EEGDEKENENSKDEGNQGLVESNNIHEAREEQYKGDDASSAVTHDTRTTSTETETVSMEN 348
Query: 135 FNVDSNGGAGSESYSDFFSEFGDQNGKGYDHDLNTEVKHANEIPGDQYAQTYNRDSNTEV 194
+V++ G ++E G ++ G + + TEVK + Y+ + + +
Sbjct: 349 SDVNAEMGITKPENKPNYTEDGIRSQPGSNFSI-TEVK----LVVGAYSNSSSSNETGNN 513
Query: 195 KLGNEIPSDGMN--ASVDYVQYQEGQSYD-----ASARNSTSGEDVNSSQY 238
L N + N A + V++ E S + +A+N+ SG D +S +
Sbjct: 514 SLSNPVAGSHQNNTALIYSVRHSEETSSNLTIVVPAAKNNMSGTDASSEHH 666
>BG648786 weakly similar to SP|P10495|GRP1_ Glycine-rich cell wall structural
protein 1.0 precursor (GRP 1.0). [Kidney bean French
bean], partial (17%)
Length = 668
Score = 33.9 bits (76), Expect = 0.45
Identities = 41/170 (24%), Positives = 64/170 (37%), Gaps = 22/170 (12%)
Frame = +1
Query: 343 YHSSIQY----AVQGHGNGHAS---SGTFSHNDNSLYRDYG---------QVGYYESQGV 386
Y +++ Y AV G+G + S +GT ++ YG VG ES V
Sbjct: 55 YGAAVNYGSAPAVGGYGGNNVSYGAAGTANYGSAPDVGGYGGGNSANYGADVGGGESNSV 234
Query: 387 --GSQAANNNWSGSYGINHQQDLD-RHTTDTATKSGGSAYGGNQQFDHSFGSSNSVNKNQ 443
GS + + G+ N+ + R + S YGG ++S SNSVN
Sbjct: 235 NYGSAPVDGGYGGANNANYGAAVGARESNSVNYGSAPGGYGGGNSANYSDRESNSVNYGS 414
Query: 444 QNASSSFGSVPLYNKVNHGHGLVNG---TVEVQRFAPSGNFGQHYNYSNT 490
+GS N N+G + G +V G +G + N+
Sbjct: 415 APGVGGYGS---GNNANYGAAVGGGESNSVNYGTAPVDGGYGAAAGFGNS 555
>TC83139
Length = 848
Score = 32.7 bits (73), Expect = 1.0
Identities = 22/63 (34%), Positives = 28/63 (43%), Gaps = 5/63 (7%)
Frame = +2
Query: 1293 DWSFQSPKHASASTWQRCPSMGNFANHEAVV-----SGSNSRSPHSRRTVSWGGSTDVTY 1347
DW PK + S+ R + + NHE VV SGS SR +V+ G Y
Sbjct: 173 DWEVSKPKDTNISSDNRLNA--DLGNHEVVVIQMSSSGSESRLDGDDPSVNQSGGGHTDY 346
Query: 1348 SPT 1350
SPT
Sbjct: 347 SPT 355
>TC87685 weakly similar to PIR|T02289|T02289 probable polygalacturonase (EC
3.2.1.15) 1 beta chain T13D8.26 - Arabidopsis thaliana,
partial (42%)
Length = 1572
Score = 32.7 bits (73), Expect = 1.0
Identities = 47/207 (22%), Positives = 76/207 (36%), Gaps = 2/207 (0%)
Frame = +1
Query: 41 AFSNLSIGGDDADVNASAFENSSGGGSGGEGKERKEEGDVKLDGGNVQEGSSSGCDGMMD 100
+F+N ++ + AD N + + + + GGSG K G NV + +
Sbjct: 553 SFTNYALETNVADQNFNTYGSGAAGGSGDFKAYAK--------GTNVPNLRFTTYSVGVA 708
Query: 101 RSDHGMESRNSSGSSADKS--NRRSSLDVKEKDWNAFNVDSNGGAGSESYSDFFSEFGDQ 158
S + +G++ D+S N E + A+ DSN S FS + DQ
Sbjct: 709 GRQQEFTSYSEAGNAGDQSFGNYGKDSAGAENKFTAYGTDSNVA------SSGFSSYADQ 870
Query: 159 NGKGYDHDLNTEVKHANEIPGDQYAQTYNRDSNTEVKLGNEIPSDGMNASVDYVQYQEGQ 218
D +N V N P + + + K N D N D
Sbjct: 871 GTGNKDTFVNYGVNMNN--PTENFKNYASGSLGAAEKFSNY--RDQANVGAD-------- 1014
Query: 219 SYDASARNSTSGEDVNSSQYWESLYPG 245
S+ + A++ST G V+ Y +S G
Sbjct: 1015SFTSYAKDSTGGTHVDFDNYGKSFNEG 1095
>TC78119 similar to GP|13605807|gb|AAK32889.1 AT3g13930/MDC16_5 {Arabidopsis
thaliana}, partial (35%)
Length = 1147
Score = 32.7 bits (73), Expect = 1.0
Identities = 31/129 (24%), Positives = 52/129 (40%)
Frame = +1
Query: 1061 RMTKSNRSVSEPDFGRSPRQETSHDAQGKASEGTSRFSRFSFGSQLLQKTMGLVLKPRPG 1120
RM S+ PD ++ QE A+ SE + F++ SFG Q ++ M + +
Sbjct: 196 RMFCSDVQPRNPDVWKTQLQERESSARNHVSESSPNFTKLSFGVQ--KRNMSTMKRGYMR 369
Query: 1121 KQAKLGEKNKFYYDENLKRWVEEGAAPPAEETALPPPPTTAAFQNGLTEYNLQSALKTEG 1180
+ E ++ + + + PP +E +P T +TE N+ LK EG
Sbjct: 370 ESLLNREISQNSQVLSRRSYSSASDLPPHQEIGMPSLSPT------MTEGNIARWLKKEG 531
Query: 1181 PPSKEGSDL 1189
G L
Sbjct: 532 DKVSPGEVL 558
Score = 30.4 bits (67), Expect = 5.0
Identities = 16/41 (39%), Positives = 19/41 (46%)
Frame = +1
Query: 1143 EGAAPPAEETALPPPPTTAAFQNGLTEYNLQSALKTEGPPS 1183
E +APPA+ET PPPP + E K PPS
Sbjct: 736 ESSAPPAKETPAPPPPKKEVAEEPAREPE-PKVSKPSAPPS 855
>BQ751144 weakly similar to PIR|PQ0479|PQ04 pistil extensin-like protein
(clone pMG14) - common tobacco (fragment), partial (11%)
Length = 632
Score = 32.7 bits (73), Expect = 1.0
Identities = 27/111 (24%), Positives = 39/111 (34%)
Frame = -1
Query: 50 DDADVNASAFENSSGGGSGGEGKERKEEGDVKLDGGNVQEGSSSGCDGMMDRSDHGMESR 109
D D A + GG+ +G ++G DGGN G DG+ G++
Sbjct: 596 DRGDDGGDAGRDGLDGGARADGGRLGDQGGGLADGGNSSTGGDR--DGLDAGLGGGLDGG 423
Query: 110 NSSGSSADKSNRRSSLDVKEKDWNAFNVDSNGGAGSESYSDFFSEFGDQNG 160
N GS A + S + D +GG G + GD G
Sbjct: 422 NGDGSVASGRSGSGSRGDGVDGGGGRDRDGDGGGGENRELGRVGDDGDDAG 270
>TC87419 similar to GP|12006165|gb|AAG44765.1 biotin carboxyl carrier protein
subunit precursor {Glycine max}, partial (70%)
Length = 1412
Score = 32.0 bits (71), Expect = 1.7
Identities = 45/195 (23%), Positives = 76/195 (38%), Gaps = 1/195 (0%)
Frame = +2
Query: 1009 PAPSSQGNVHGNEQNYQSGAHRVSNSQSTMAMSSLVPSGSMEPNGEWTADNNRMTKSNRS 1068
P PSS + N+ +QSG + NSQS ++S+ S ++ + + N +
Sbjct: 119 PVPSSLLGLKSNKILFQSGLS-LKNSQSFGSLSAESASFGIQCLNKKQFPVKIQAQLNEA 295
Query: 1069 VSEPDFGRSPRQETSHDAQGK-ASEGTSRFSRFSFGSQLLQKTMGLVLKPRPGKQAKLGE 1127
+ +P +S + + + S S S S S + + LV+ +L
Sbjct: 296 AVVENLNSAPTVASSEEKENQNGSVPASTVSDESAISAFMSQVADLVILVDSRDIVELQL 475
Query: 1128 KNKFYYDENLKRWVEEGAAPPAEETALPPPPTTAAFQNGLTEYNLQSALKTEGPPSKEGS 1187
K Y E + R ++ AL PPP TA Q+ Y L PP+ +
Sbjct: 476 KQSDY--ELVIR----------KKEALQPPPATAMPQSAPPLYYPTLPLPPPPPPTAYSA 619
Query: 1188 DLKTSNPELTPGIPP 1202
+ + TP +PP
Sbjct: 620 TASSPPSKATPALPP 664
Database: MTGI
Posted date: Oct 22, 2004 3:39 PM
Number of letters in database: 27,044,181
Number of sequences in database: 36,976
Lambda K H
0.309 0.128 0.371
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 40,246,490
Number of Sequences: 36976
Number of extensions: 569032
Number of successful extensions: 2804
Number of sequences better than 10.0: 76
Number of HSP's better than 10.0 without gapping: 2644
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 2765
length of query: 1399
length of database: 9,014,727
effective HSP length: 108
effective length of query: 1291
effective length of database: 5,021,319
effective search space: 6482522829
effective search space used: 6482522829
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.1 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 42 (21.7 bits)
S2: 65 (29.6 bits)
Medicago: description of AC138087.1


