

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC138063.5 + phase: 0 /pseudo
(840 letters)
Database: MTGI
36,976 sequences; 27,044,181 total letters
Searching..................................................done
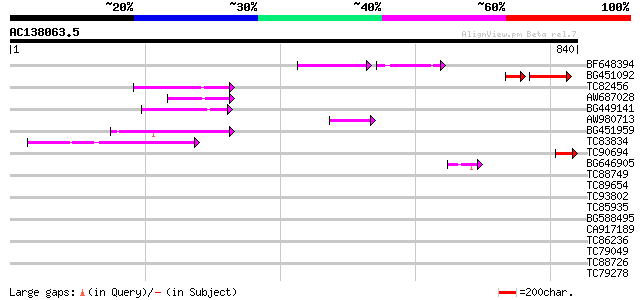
Score E
Sequences producing significant alignments: (bits) Value
BF648394 79 7e-30
BG451092 68 7e-14
TC82456 weakly similar to GP|18057159|gb|AAL58182.1 putative far... 66 5e-11
AW687028 weakly similar to GP|9909176|dbj| hypothetical protein~... 62 1e-09
BG449141 similar to GP|15983442|gb At1g10240/F14N23_12 {Arabidop... 54 3e-07
AW980713 PIR|H72173|H721 D5L protein - variola minor virus (stra... 51 2e-06
BG451959 homologue to GP|6175165|gb| Mutator-like transposase {A... 50 5e-06
TC83834 similar to PIR|G96565|G96565 F6D8.26 [imported] - Arabid... 48 1e-05
TC90694 47 2e-05
BG646905 weakly similar to GP|20804419|dbj contains EST C73272(E... 45 1e-04
TC88749 41 0.002
TC89654 weakly similar to PIR|T05644|T05644 hypothetical protein... 36 0.054
TC93802 similar to GP|2194139|gb|AAB61114.1| EST gb|ATTS0887 com... 33 0.35
TC85935 homologue to PIR|S20941|S20941 protochlorophyllide reduc... 33 0.59
BG588495 32 1.0
CA917189 similar to PIR|T05645|T05 hypothetical protein F20D10.3... 32 1.3
TC86236 similar to PIR|T05311|T05311 sulfolipid biosynthesis pro... 30 2.9
TC79049 weakly similar to PIR|T10798|T10798 pherophorin-S - Volv... 29 6.6
TC88726 similar to GP|11994239|dbj|BAB01361. gb|AAD27719.1~gene_... 29 8.6
TC79278 similar to GP|21537229|gb|AAM61570.1 unknown {Arabidopsi... 29 8.6
>BF648394
Length = 650
Score = 79.0 bits (193), Expect(2) = 7e-30
Identities = 47/111 (42%), Positives = 66/111 (59%), Gaps = 1/111 (0%)
Frame = -3
Query: 427 VESAHGTLKGYLEDSKGDLAKGWKAINAMLTVQFTEIQATFGRNSTVLEHKFKGNKMYKS 486
VESA LK YL++S GDL W+ I+ ML +QF IQ TFG++ +VLEH+FK +Y
Sbjct: 648 VESAXDLLKKYLDNSVGDLGTCWEKIHDMLLLQFIAIQTTFGQSVSVLEHRFKYVTLYSG 469
Query: 487 LEYNVSRTAMNYIYKQASRV-KDCGSDSKKCGCVIRWTHALPCTCLIYKKL 536
+ +VSR A++ I + + K + GCV R ++ LPC C I KL
Sbjct: 468 IGGHVSRYALDNIALEETHCRKTLCMYNDI*GCVQRTSYGLPCPCEIATKL 316
Score = 70.9 bits (172), Expect(2) = 7e-30
Identities = 44/103 (42%), Positives = 60/103 (57%), Gaps = 1/103 (0%)
Frame = -2
Query: 544 LDEINIHWKKLCFEVDDDVEESNR-HVSIQNELNAIQERFNKADYNMKLIIKEQMREMTF 602
+DEI HW KL EESN ++ EL AI E K + MK+ +KE +R++TF
Sbjct: 295 IDEIYHHWLKLRMG-----EESNEVGFCVEVELKAIVEHLKKLPFQMKVEVKEGLRQLTF 131
Query: 603 PHTTSLTPPIQKLKTKGAQKGAQKRKRSTLDDSSTTREPSFWE 645
P TT ++PP +K+ TKGA+K RS +ST+R PS WE
Sbjct: 130 PETTMMSPPPRKVPTKGAKKKVD-IARSKGKITSTSRIPSSWE 5
>BG451092
Length = 663
Score = 68.2 bits (165), Expect(2) = 7e-14
Identities = 33/66 (50%), Positives = 44/66 (66%), Gaps = 3/66 (4%)
Frame = -1
Query: 770 IAPEEKW--LTVPDMGHVIASLYNKVVV-VLKYGNNFSETCFPLRGRPPTNPSARIMCLG 826
IAP KW LT+PDM H++A+ YN++VV + S+T + +RGRPP NP +RIMCLG
Sbjct: 492 IAPSLKW*WLTLPDMSHIVATYYNRLVVEITSVEIRISDTFYLIRGRPPINPKSRIMCLG 313
Query: 827 LIPNHF 832
P F
Sbjct: 312 FNPKSF 295
Score = 27.7 bits (60), Expect(2) = 7e-14
Identities = 10/30 (33%), Positives = 20/30 (66%)
Frame = -2
Query: 735 KELTGKKNDYSGIFGGEDRLKFLIDSLYPP 764
+E+ +N Y I+GGE+ ++++ L+PP
Sbjct: 569 REVKEHRNHYMRIYGGENCYNYILN*LHPP 480
>TC82456 weakly similar to GP|18057159|gb|AAL58182.1 putative far-red
impaired response protein {Oryza sativa}, partial (7%)
Length = 1244
Score = 66.2 bits (160), Expect = 5e-11
Identities = 43/151 (28%), Positives = 69/151 (45%), Gaps = 2/151 (1%)
Frame = +2
Query: 184 RGSR*EMQELFQKFEENNYVFNYRT-VSGSRTVQDLFFAHPESVKLFNTFPTVLVMDSTY 242
RG + F + ++ N F + R +Q+LF+A + F V+ +D+TY
Sbjct: 710 RGDVDAIHSFFHRMQKQNSQFYCAMDMDDKRNIQNLFWADARCRAAYEYFGEVITLDTTY 889
Query: 243 KTNQYKMPLFEIVGLTSTEMTYNVGFAFLTMEKEDNFTWGLQESANLLKCKK-QMPKVIV 301
T++Y +PL VG+ T +G A L+ TW L+C P I+
Sbjct: 890 LTSKYDLPLVPFVGVNHHGQTILLGCAILSNLDAKTLTWLF---TRWLECMHGHAPNGII 1060
Query: 302 TDRDTGLMNAVANIFPKSTALVCEFHILKNV 332
T+ D + NA+ FPK+ C +HI+K V
Sbjct: 1061TEEDKAMKNAIEVAFPKARHRWCLWHIMKKV 1153
>AW687028 weakly similar to GP|9909176|dbj| hypothetical protein~similar to
Arabidopsis thaliana F20D10.300, partial (11%)
Length = 662
Score = 61.6 bits (148), Expect = 1e-09
Identities = 33/100 (33%), Positives = 51/100 (51%), Gaps = 1/100 (1%)
Frame = +1
Query: 235 VLVMDSTYKTNQYKMPLFEIVGLTSTEMTYNVGFAFLTMEKEDNFTWGLQESANLLKC-K 293
VL D+TYK N+Y PL G T G A L E +++ W L L+C +
Sbjct: 4 VLAFDTTYKKNKYNYPLCIFSGCNHHSQTIIFGVALLEDETIESYKWVLNR---FLECME 174
Query: 294 KQMPKVIVTDRDTGLMNAVANIFPKSTALVCEFHILKNVR 333
+ PK +VTD D + A+ +FP ++ +C +H+ KN +
Sbjct: 175 NKFPKAVVTDGDGSMREAIKQVFPDASHRLCAWHLHKNAQ 294
>BG449141 similar to GP|15983442|gb At1g10240/F14N23_12 {Arabidopsis
thaliana}, partial (33%)
Length = 690
Score = 53.5 bits (127), Expect = 3e-07
Identities = 32/134 (23%), Positives = 69/134 (50%)
Frame = +3
Query: 196 KFEENNYVFNYRTVSGSRTVQDLFFAHPESVKLFNTFPTVLVMDSTYKTNQYKMPLFEIV 255
K ++ N+ F Y T+ + ++++ +++ S++L++ F +V D+T++ + MPL V
Sbjct: 267 KDKDPNFKFEY-TLDANNRLENIAWSYASSIQLYDIFGDAVVFDTTHRLTAFDMPLGLWV 443
Query: 256 GLTSTEMTYNVGFAFLTMEKEDNFTWGLQESANLLKCKKQMPKVIVTDRDTGLMNAVANI 315
G+ + M G L E +F+W ++ + K P+ I+TD++ L A++
Sbjct: 444 GINNYGMPCFFGCVLLRDETVRSFSWAIKAFLGFMNGK--APQTILTDQNICLKEALSAE 617
Query: 316 FPKSTALVCEFHIL 329
P + C + I+
Sbjct: 618 MPMTKHAFCIWMIV 659
>AW980713 PIR|H72173|H721 D5L protein - variola minor virus (strain
Garcia-1966), partial (13%)
Length = 560
Score = 50.8 bits (120), Expect = 2e-06
Identities = 25/69 (36%), Positives = 39/69 (56%), Gaps = 1/69 (1%)
Frame = +1
Query: 475 EHKFKGNKMYKSLEYNVSRTAMNYIYKQAS-RVKDCGSDSKKCGCVIRWTHALPCTCLIY 533
+H+FKG + L N+S ++ +A V CG+ KCGC+I T+ L TC+I+
Sbjct: 310 KHEFKGKLLCSKLVRNISSNIFYFLADKAHWAVAGCGTYKSKCGCLIWITYGLSYTCIIF 489
Query: 534 KKLNLGSCI 542
KK+ L + I
Sbjct: 490 KKIKLNTFI 516
>BG451959 homologue to GP|6175165|gb| Mutator-like transposase {Arabidopsis
thaliana}, partial (28%)
Length = 658
Score = 49.7 bits (117), Expect = 5e-06
Identities = 48/193 (24%), Positives = 83/193 (42%), Gaps = 9/193 (4%)
Frame = +2
Query: 150 KPKNILLDLKGRWKDNRTVAKQIYNARHRYKLSIRGSR*EMQELFQKFEENNYVFNYRTV 209
KPK IL ++ R KQ + + ++RGS E L ++ + N ++
Sbjct: 77 KPKEILEEIH-RVHGITLSYKQAWRGKEHIMAAMRGSFEEGYRLLPQYCAHVKRTNPGSI 253
Query: 210 SG------SRTVQDLFFAHPESVK-LFNTFPTVLVMDSTYKTNQYKMPLFEIVGLTSTEM 262
+ Q LF + S+ L N +L +D Y ++Y L G
Sbjct: 254 ASVYGNPSDNCFQRLFISFQASIYGLLNACRPLLGLDRIYLKSKYLGTLLLATGFDGDGA 433
Query: 263 TYNVGFAFLTMEKEDNFTWGLQESANLLKCK-KQMPKV-IVTDRDTGLMNAVANIFPKST 320
+ + F + E +DN+ W L + NLL+ + MP++ I++DR G+++ V FP +
Sbjct: 434 LFPLAFGVVDEENDDNWMWFLSKLHNLLEINTENMPRLTILSDRQQGIVDGVEANFPTAF 613
Query: 321 ALVCEFHILKNVR 333
C H+ N R
Sbjct: 614 HGFCMRHLSDNFR 652
>TC83834 similar to PIR|G96565|G96565 F6D8.26 [imported] - Arabidopsis
thaliana, partial (24%)
Length = 1032
Score = 48.1 bits (113), Expect = 1e-05
Identities = 56/259 (21%), Positives = 106/259 (40%), Gaps = 4/259 (1%)
Frame = +3
Query: 27 KYEDREEMINWARGQAIEAGFTLIIDKS-YSGSGRRKPKLVLAFERSGEYKGTKKSKREG 85
++E ++ ++ A E GF + + S + + + K VL G +K TK
Sbjct: 228 EFESYDDAYSYYICYAKEVGFCVRVKNSWFKRNSKEKYGAVLCCSSQG-FKRTKDVNNLR 404
Query: 86 TGSRKCGCPFRLRGYFSATKLWSLNIVSGLHNHKMEPKLEGHMLAGRLTAKESKIVGDMT 145
+R GCP +R ++ W + V+ HNH + K+ + K S D
Sbjct: 405 KETRT-GCPAMIRMKLVESQRWRICEVTLEHNHVLGAKIH------KSIKKNSLPSSDAE 563
Query: 146 RNLIKPKNIL-LDLKGRWKDNRTVAKQIYNARHRYKLSIR-GSR*EMQELFQKFEENNYV 203
IK + L +D +G N +++ KL++R G + + + N
Sbjct: 564 GKTIKVYHALVIDTEGNDNLNSNARDDRAFSKYSNKLNLRKGDTQAIYNFLCRMQLTNPN 743
Query: 204 FNY-RTVSGSRTVQDLFFAHPESVKLFNTFPTVLVMDSTYKTNQYKMPLFEIVGLTSTEM 262
F Y + +++ + +S F V+ D+TY N+Y++PL +VG+
Sbjct: 744 FFYLMDFNDEGHLRNALWVDAKSRAACGYFSDVIYFDNTYLVNKYEIPLVALVGINHHGQ 923
Query: 263 TYNVGFAFLTMEKEDNFTW 281
+ +G L E +++ W
Sbjct: 924 SVLLGCGLLAGEIIESYKW 980
>TC90694
Length = 588
Score = 47.4 bits (111), Expect = 2e-05
Identities = 19/32 (59%), Positives = 25/32 (77%)
Frame = +2
Query: 809 PLRGRPPTNPSARIMCLGLIPNHFVHVFLRPG 840
P+RG+PP N + ++CLGLI HFVHVFL+ G
Sbjct: 188 PVRGQPPLNSNTYVICLGLI*EHFVHVFLKDG 283
>BG646905 weakly similar to GP|20804419|dbj contains EST
C73272(E3606)~similar to Arabidopsis thaliana chromosome
5 MZF18.9~unknown protein, partial (10%)
Length = 735
Score = 45.1 bits (105), Expect = 1e-04
Identities = 28/64 (43%), Positives = 32/64 (49%), Gaps = 12/64 (18%)
Frame = +2
Query: 649 AQFPDSQASQTKPSLQKKKSVCVVNPSSTPTPTQ------------YIPCIDQMPNVMKP 696
+QFPD + K S K SVC+ NPS TP P YIP +D MP M P
Sbjct: 479 SQFPDGLILEKKLSFPK--SVCIGNPSPTPVPRPTPVPKPTMVLMLYIPHVDWMPLFMCP 652
Query: 697 YIEK 700
YIEK
Sbjct: 653 YIEK 664
>TC88749
Length = 1221
Score = 40.8 bits (94), Expect = 0.002
Identities = 27/63 (42%), Positives = 39/63 (61%), Gaps = 1/63 (1%)
Frame = +2
Query: 28 YEDREEMINWARGQAIEAGFTLIIDKSYSGSGRRKPKLVLAFERSGEYKGTKKS-KREGT 86
++D+E+M++ R QA AGFT+ I + S + P L L ERSG++K +KK K E T
Sbjct: 551 WKDKEDMLSSIRRQANRAGFTVGIKR----SSIQNPMLELMCERSGDHKVSKKRLKHEAT 718
Query: 87 GSR 89
SR
Sbjct: 719 *SR 727
Score = 37.4 bits (85), Expect = 0.024
Identities = 18/34 (52%), Positives = 22/34 (63%), Gaps = 1/34 (2%)
Frame = +2
Query: 776 WLTVPDMGHVIASLYNKVVVVL-KYGNNFSETCF 808
WLT+PDMGH+I + N VVV L + SE CF
Sbjct: 764 WLTLPDMGHIIVTYCNMVVVELTNHEIGISEICF 865
>TC89654 weakly similar to PIR|T05644|T05644 hypothetical protein F20D10.290
- Arabidopsis thaliana, partial (65%)
Length = 1358
Score = 36.2 bits (82), Expect = 0.054
Identities = 21/90 (23%), Positives = 39/90 (43%), Gaps = 1/90 (1%)
Frame = +2
Query: 182 SIRGSR*EMQELFQKFEENNYVFNYRTVSG-SRTVQDLFFAHPESVKLFNTFPTVLVMDS 240
++ G ++ + ++ N+ F Y S ++F+A ++ F ++ D+
Sbjct: 107 TLGGGGHQVLDYLKRMRAENHAFFYAVQSDVDNAGGNIFWADETCRTNYSYFGDTVIFDT 286
Query: 241 TYKTNQYKMPLFEIVGLTSTEMTYNVGFAF 270
TYKTNQY++P G G AF
Sbjct: 287 TYKTNQYRVPFASFTGFNHHGQPVLFGCAF 376
>TC93802 similar to GP|2194139|gb|AAB61114.1| EST gb|ATTS0887 comes from
this gene. {Arabidopsis thaliana}, partial (25%)
Length = 603
Score = 33.5 bits (75), Expect = 0.35
Identities = 22/73 (30%), Positives = 35/73 (47%), Gaps = 4/73 (5%)
Frame = +1
Query: 614 KLKTKGAQKGAQKRKRSTLDDSSTTREPSFWEHVDAQFPDSQASQT----KPSLQKKKSV 669
K TK A+K ++ RSTL+ S+++ +P EHV + SQ+ +T + K
Sbjct: 307 KDSTKDAKKTKPQKNRSTLEISNSSAKPHIVEHVYGELITSQSQRTLLDDSSASDNKSQA 486
Query: 670 CVVNPSSTPTPTQ 682
V S P T+
Sbjct: 487 YVTYRSCIPADTE 525
>TC85935 homologue to PIR|S20941|S20941 protochlorophyllide reductase (EC
1.3.1.33) precursor - garden pea, complete
Length = 1481
Score = 32.7 bits (73), Expect = 0.59
Identities = 24/75 (32%), Positives = 36/75 (48%), Gaps = 4/75 (5%)
Frame = +2
Query: 603 PHTTSLTPPIQKLKTKG--AQKGAQKRKRSTLDDSSTTREPSF--WEHVDAQFPDSQASQ 658
P +L PP QK TKG ++ A KR + D S T+ + W A F ++Q SQ
Sbjct: 1022 PLFRTLFPPFQKYITKGYVSEDEAGKRLAQVVSDPSLTKSGVYWSWNKASASF-ENQLSQ 1198
Query: 659 TKPSLQKKKSVCVVN 673
++K + V V+
Sbjct: 1199 EASDVEKARKVWEVS 1243
>BG588495
Length = 540
Score = 32.0 bits (71), Expect = 1.0
Identities = 19/53 (35%), Positives = 28/53 (51%)
Frame = +3
Query: 27 KYEDREEMINWARGQAIEAGFTLIIDKSYSGSGRRKPKLVLAFERSGEYKGTK 79
K +DR+E++ W R QA +AGFT+ K +L FE GE + T+
Sbjct: 15 KSKDRDELLEWTRHQANKAGFTI---------DDLKAQLNTYFEHLGENQYTR 146
>CA917189 similar to PIR|T05645|T05 hypothetical protein F20D10.300 -
Arabidopsis thaliana, partial (12%)
Length = 312
Score = 31.6 bits (70), Expect = 1.3
Identities = 12/39 (30%), Positives = 22/39 (55%)
Frame = +1
Query: 297 PKVIVTDRDTGLMNAVANIFPKSTALVCEFHILKNVRAR 335
P I TD D+ + +A+ +FP + C++HI K + +
Sbjct: 25 PLSITTDHDSVIQSAIMQVFPDTRHRFCKWHIFKQCQEK 141
>TC86236 similar to PIR|T05311|T05311 sulfolipid biosynthesis protein SQD1
chloroplast [validated] - Arabidopsis thaliana, partial
(83%)
Length = 1918
Score = 30.4 bits (67), Expect = 2.9
Identities = 24/79 (30%), Positives = 40/79 (50%)
Frame = +1
Query: 666 KKSVCVVNPSSTPTPTQYIPCIDQMPNVMKPYIEKIVNVDGDGYCGYRVVAENLGLGSDS 725
+KS+ VVN S+ T Q P + KP ++++ + GDGYCG+ A L L +
Sbjct: 322 RKSLAVVNASTISTG-QEAPVQTSSGDPFKP--KRVMVIGGDGYCGW---ATALHLSNKG 483
Query: 726 YGLVPLALIKELTGKKNDY 744
Y +A++ L + D+
Sbjct: 484 Y---EVAIVDNLVRRLFDH 531
>TC79049 weakly similar to PIR|T10798|T10798 pherophorin-S - Volvox carteri,
partial (6%)
Length = 957
Score = 29.3 bits (64), Expect = 6.6
Identities = 18/52 (34%), Positives = 22/52 (41%), Gaps = 5/52 (9%)
Frame = +1
Query: 650 QFPDSQASQ-----TKPSLQKKKSVCVVNPSSTPTPTQYIPCIDQMPNVMKP 696
QFP SQ Q PS S P +PTP Y PC + P+ +P
Sbjct: 760 QFPSSQVHQFFSPNPSPSPHSPLSPSSTLPLPSPTPRPYPPCPSKPPSNPQP 915
>TC88726 similar to GP|11994239|dbj|BAB01361.
gb|AAD27719.1~gene_id:MUJ8.10~similar to unknown protein
{Arabidopsis thaliana}, partial (58%)
Length = 2001
Score = 28.9 bits (63), Expect = 8.6
Identities = 23/86 (26%), Positives = 36/86 (41%), Gaps = 5/86 (5%)
Frame = +2
Query: 644 WEHVDAQFPDSQASQTKPSLQKKKSVCVVNPSSTPTPTQY----IPCIDQMPNVMKPYIE 699
W +D ++ + P + S +V P S Y + I + + P
Sbjct: 1283 WLKMDGTLSANEPFEIPPKAIRLASERMVFPLSLRHANSYATKRVVLIGDAAHTIHPLAG 1462
Query: 700 KIVNVD-GDGYCGYRVVAENLGLGSD 724
+ VN+ GD Y RV+AE + LGSD
Sbjct: 1463 QGVNLGFGDAYSLSRVIAEGIALGSD 1540
>TC79278 similar to GP|21537229|gb|AAM61570.1 unknown {Arabidopsis
thaliana}, partial (87%)
Length = 1134
Score = 28.9 bits (63), Expect = 8.6
Identities = 13/43 (30%), Positives = 23/43 (53%)
Frame = +3
Query: 702 VNVDGDGYCGYRVVAENLGLGSDSYGLVPLALIKELTGKKNDY 744
+ ++GDG C +R +A+ L D + V ++K+L K Y
Sbjct: 477 LQMEGDGNCQFRAIADQLFQNPDYHKYVRRQVVKQLKHHKKLY 605
Database: MTGI
Posted date: Oct 22, 2004 3:39 PM
Number of letters in database: 27,044,181
Number of sequences in database: 36,976
Lambda K H
0.319 0.136 0.406
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 26,632,294
Number of Sequences: 36976
Number of extensions: 388219
Number of successful extensions: 2013
Number of sequences better than 10.0: 41
Number of HSP's better than 10.0 without gapping: 1992
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 2004
length of query: 840
length of database: 9,014,727
effective HSP length: 104
effective length of query: 736
effective length of database: 5,169,223
effective search space: 3804548128
effective search space used: 3804548128
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.8 bits)
S2: 63 (28.9 bits)
Medicago: description of AC138063.5


