

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
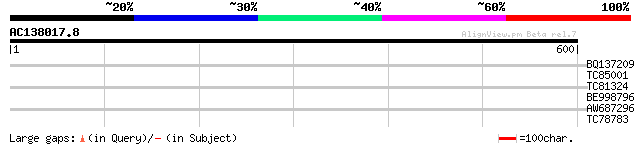
Query= AC138017.8 - phase: 2 /pseudo
(600 letters)
Database: MTGI
36,976 sequences; 27,044,181 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
BQ137209 32 0.91
TC85001 similar to GP|17529298|gb|AAL38876.1 unknown protein {Ar... 31 1.5
TC81324 similar to GP|21553871|gb|AAM62964.1 unknown {Arabidopsi... 30 2.6
BE998796 similar to GP|5441893|dbj| ESTs C99174(E10437) D22295(C... 30 2.6
AW687296 weakly similar to GP|20521419|db putative receptor-like... 30 3.5
TC78783 similar to GP|21537171|gb|AAM61512.1 unknown {Arabidopsi... 29 4.5
>BQ137209
Length = 1136
Score = 31.6 bits (70), Expect = 0.91
Identities = 12/20 (60%), Positives = 14/20 (70%)
Frame = -2
Query: 396 CCVSWTVKINLLWVFFTEIC 415
CCVS V LLW+ FTE+C
Sbjct: 1129 CCVSVYVVTELLWICFTEVC 1070
>TC85001 similar to GP|17529298|gb|AAL38876.1 unknown protein {Arabidopsis
thaliana}, partial (15%)
Length = 588
Score = 30.8 bits (68), Expect = 1.5
Identities = 17/56 (30%), Positives = 33/56 (58%)
Frame = -2
Query: 39 KRQGSRKEKLNMQ*VMKSKHKEDNLGQWLLHLKRERTMEALIITFCLEQLPDHNLL 94
+R+ +R + L + S+ K+D+L +LL LK+++ E L+ ++ L + NLL
Sbjct: 494 RRKSTRSKPLKLTRQKNSQRKDDHLVSFLLKLKKKKEKENLL*KPYVKSLKEINLL 327
>TC81324 similar to GP|21553871|gb|AAM62964.1 unknown {Arabidopsis
thaliana}, partial (75%)
Length = 1156
Score = 30.0 bits (66), Expect = 2.6
Identities = 16/56 (28%), Positives = 28/56 (49%)
Frame = +2
Query: 97 VCCKPRKL*KSVILHLQNGSLLHLFPSMQQIHHIFSLRSMLFVAWEPDIKLLLYMI 152
+ K L + I H +NG+L+ L +HH+ +M V W+P + +L +I
Sbjct: 422 ILLKAANLVTNAISHTENGNLVSLL-----LHHLMITVTMTTVTWDPPMLAVLAVI 574
>BE998796 similar to GP|5441893|dbj| ESTs C99174(E10437) D22295(C10709)
correspond to a region of the predicted gene.~Similar
to, partial (28%)
Length = 609
Score = 30.0 bits (66), Expect = 2.6
Identities = 16/56 (28%), Positives = 28/56 (49%)
Frame = +2
Query: 97 VCCKPRKL*KSVILHLQNGSLLHLFPSMQQIHHIFSLRSMLFVAWEPDIKLLLYMI 152
+ K L + I H +NG+L+ L +HH+ +M V W+P + +L +I
Sbjct: 365 ILLKAANLVTNAISHTENGNLVSLL-----LHHLMITVTMTTVTWDPPMLAVLAVI 517
>AW687296 weakly similar to GP|20521419|db putative receptor-like kinase
{Oryza sativa (japonica cultivar-group)}, partial (7%)
Length = 458
Score = 29.6 bits (65), Expect = 3.5
Identities = 14/30 (46%), Positives = 20/30 (66%), Gaps = 1/30 (3%)
Frame = +3
Query: 114 NGSLLHLFPSMQQIHHIFSLRSMLF-VAWE 142
N L+ LF S QQIH +FS+ M+F + W+
Sbjct: 339 NDMLIPLFFSYQQIHSLFSILCMIFSIPWQ 428
>TC78783 similar to GP|21537171|gb|AAM61512.1 unknown {Arabidopsis
thaliana}, partial (21%)
Length = 1431
Score = 29.3 bits (64), Expect = 4.5
Identities = 17/44 (38%), Positives = 29/44 (65%)
Frame = +3
Query: 290 KRQFHMPQMLPSMYIIIVIHCI**GNLLMEERYFVLLQLALPLI 333
K+ +H+ Q+ P + + IV H I * LL++ + ++LQL +PLI
Sbjct: 102 KK*WHL-QLEPVVLLTIVSHKIS*CPLLLQNNWSLILQLVIPLI 230
Database: MTGI
Posted date: Oct 22, 2004 3:39 PM
Number of letters in database: 27,044,181
Number of sequences in database: 36,976
Lambda K H
0.360 0.161 0.561
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 21,144,706
Number of Sequences: 36976
Number of extensions: 339568
Number of successful extensions: 4126
Number of sequences better than 10.0: 12
Number of HSP's better than 10.0 without gapping: 1729
Number of HSP's successfully gapped in prelim test: 203
Number of HSP's that attempted gapping in prelim test: 2356
Number of HSP's gapped (non-prelim): 2031
length of query: 600
length of database: 9,014,727
effective HSP length: 102
effective length of query: 498
effective length of database: 5,243,175
effective search space: 2611101150
effective search space used: 2611101150
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 14 ( 7.3 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 37 (21.9 bits)
S2: 61 (28.1 bits)
Medicago: description of AC138017.8


