

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC138012.9 - phase: 0
(672 letters)
Database: MTGI
36,976 sequences; 27,044,181 total letters
Searching..................................................done
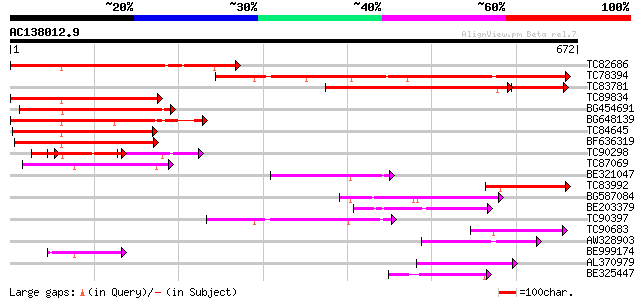
Score E
Sequences producing significant alignments: (bits) Value
TC82686 weakly similar to GP|10177020|dbj|BAB10258. leucine zipp... 335 4e-92
TC78394 similar to GP|10177020|dbj|BAB10258. leucine zipper prot... 300 2e-81
TC83781 homologue to PIR|S21495|S21495 tomato leucine zipper-con... 238 4e-81
TC89834 285 5e-77
BG454691 similar to GP|8347078|emb| NADH dehydrogenase subunit 4... 278 4e-75
BG648139 274 7e-74
TC84645 213 1e-55
BF636319 weakly similar to PIR|G71155|G711 hypothetical protein ... 166 3e-41
TC90298 130 1e-30
TC87069 weakly similar to GP|9758877|dbj|BAB09431.1 contains sim... 114 9e-26
BE321047 similar to GP|10177020|db leucine zipper protein {Arabi... 110 2e-24
TC83992 weakly similar to GP|10177020|dbj|BAB10258. leucine zipp... 88 9e-18
BG587084 homologue to GP|6002696|gb| histone acetyltransferase M... 87 3e-17
BE203379 similar to GP|22136732|gb unknown protein {Arabidopsis ... 74 1e-13
TC90397 similar to GP|15010740|gb|AAK74029.1 AT5g59730/mth12_130... 73 3e-13
TC90683 similar to GP|22136732|gb|AAM91685.1 unknown protein {Ar... 69 6e-12
AW328903 weakly similar to GP|8439895|gb|A It contains a interfe... 68 1e-11
BE999174 similar to GP|11546040|emb bA218B22.1.1 (novel protein ... 57 2e-08
AL370979 weakly similar to GP|15010740|gb| AT5g59730/mth12_130 {... 57 3e-08
BE325447 weakly similar to GP|18700111|gb| At1g07000/F10K1_20 {A... 52 6e-07
>TC82686 weakly similar to GP|10177020|dbj|BAB10258. leucine zipper protein
{Arabidopsis thaliana}, partial (5%)
Length = 858
Score = 335 bits (858), Expect = 4e-92
Identities = 172/282 (60%), Positives = 222/282 (77%), Gaps = 9/282 (3%)
Frame = +1
Query: 1 MIPVIIRIRMCLLQTKVWRFVGFASAAVGLLCYALSTSFNHLFGNWNLLKIFLYTLFSFI 60
MIP+II+IR LL+TKVWRFVGF SAAVGL+CYALS+SFNHLFGNWNLLK+ LYTL SFI
Sbjct: 19 MIPLIIQIRCWLLETKVWRFVGFVSAAVGLICYALSSSFNHLFGNWNLLKVILYTLSSFI 198
Query: 61 ---MILYANIWKNSRSLRFKAHTAFLVLTITSVYSFFFDKVVNGKPDAYSLISCAAFAIM 117
+ILYAN W+NSRSLRFKAHTAFLVLTIT+VYSFF DK++NGKPDAY+LISC +F++M
Sbjct: 199 ICLLILYANTWQNSRSLRFKAHTAFLVLTITAVYSFFSDKIMNGKPDAYTLISCTSFSMM 378
Query: 118 SMSLSRQSQCGIEVDLLYFFLGCLIMQLMKIKLQLVIVGAGFSYSLIIIRSSLSSTNSVP 177
S+SLSRQ QCG EVDL+YF+LGCLI+QLMKIKL L IVG +SY +II+RSS SS N
Sbjct: 379 SLSLSRQIQCGFEVDLMYFYLGCLIVQLMKIKLSLAIVGVCYSYCVIILRSSFSSLNVTQ 558
Query: 178 ENEYFGVGLRGENSVVIEMDSLLRPSNDIYSGMMQQLTTCINALQQDSLNIIDRLMEEY- 236
E + G+ E V+I++DS + +N + +MQ+ TC+N L+Q++ NI ++ +E+
Sbjct: 559 ETQCLGL---EEQHVIIQVDSQHKNTNS-HDNIMQEFMTCMNELKQNNSNIANKFLEKVK 726
Query: 237 NQGKC-----DFEMDTLPSEKLHNLHEIVKLMLCAGYEKECS 273
GK +F +D LP+E +++LH+ VK+M+ AG+EKECS
Sbjct: 727 GNGKLVVTDHNFIIDALPNETINHLHKTVKMMVDAGFEKECS 852
>TC78394 similar to GP|10177020|dbj|BAB10258. leucine zipper protein
{Arabidopsis thaliana}, partial (83%)
Length = 2109
Score = 300 bits (767), Expect = 2e-81
Identities = 175/440 (39%), Positives = 268/440 (60%), Gaps = 20/440 (4%)
Frame = +1
Query: 245 MDTLPSEKLHNLHEIVKLMLCAGYEKECSAVYISWRKVLLQKGL----LNKIFVLPEAKI 300
+D LP +++L EI K M+ AG+ KECS VY R+ L++ L L K+ + K+
Sbjct: 745 IDALPPATVNDLREIAKRMIAAGFGKECSHVYGGCRREFLEESLSRLGLQKLSISEVHKM 924
Query: 301 NTERERERYLDTMFQRWMTASDIATTVLFPIEQKFCDLVFSGFSSATSHC---FIEICQE 357
+ L+ +RW+ AS++A +LFP E++ CD VFSG SS+++ F+E+C+
Sbjct: 925 QWQD-----LEDEIERWIKASNVALKILFPSERRLCDRVFSGLSSSSAAADLSFMEVCRG 1089
Query: 358 ATFQLLNFADVIAYGSLSKWRLFKMVDIFVKLNNLVPKFESLFPNS----LVNEAIAVRN 413
+ QLLNF+D +A GS S RLF+++D+F + +L+P+FESLF + LVN+AI
Sbjct: 1090 SAIQLLNFSDAVAIGSRSPERLFRVLDVFETMRDLIPEFESLFSDQYCSFLVNQAITNWK 1269
Query: 414 RLGDASRVLFMKMHNFIFRVPAAKQVVSSYGQHHQMTIQVMSYVSSACRKRRKLEQI--- 470
RLG+A FM++ N I R P K VV G H +T VM+Y+ +ACR + LE +
Sbjct: 1270 RLGEAITGTFMELENLISRDPV-KAVVPGGGLH-PITRYVMNYLRAACRSSKTLELVFKD 1443
Query: 471 ----LEEYPEVHNEVEASSFFLKQMEQIMRMLQRKLIVKSENCKDRALRHIFMLNNRSHI 526
L++Y + ++++S F Q+ IM +L+R L KS KD AL +FM+NN +I
Sbjct: 1444 NALSLKDYHKHDESLQSNSSFSVQISWIMDLLERNLEAKSRIYKDPALCSVFMMNNGRYI 1623
Query: 527 EAMNKFSRLETIFGNDWFQNNKAKIQQNLDLYKRSAWDEVMDFLKLDNNESITKELLKEK 586
K S L T+ G+DW + + K++Q Y+RS+W++++ FLK+ E++ + +KEK
Sbjct: 1624 VQKTKDSELGTLMGDDWIRKHSTKVRQCHTNYQRSSWNKLLGFLKV---ETLAAKPMKEK 1794
Query: 587 IHLFNNRFEAICRVQSAWFIYGSQLRGEIISSVGNILLPAYGIFVGRLHGI--LGNQAYK 644
+ +FN FE ICRVQS WF++ QL+ EI S+ +LLPAYG F+GR + L + K
Sbjct: 1795 LKMFNLHFEEICRVQSQWFVFDEQLKEEIRISIEKLLLPAYGSFIGRFQILPELAKNSDK 1974
Query: 645 YIKYGMIEIQDLLNHLFLGN 664
YIK+GM +I+ LNHLF G+
Sbjct: 1975 YIKFGMEDIEARLNHLFQGS 2034
>TC83781 homologue to PIR|S21495|S21495 tomato leucine zipper-containing
protein - tomato, partial (5%)
Length = 1047
Score = 238 bits (607), Expect(2) = 4e-81
Identities = 118/227 (51%), Positives = 166/227 (72%), Gaps = 6/227 (2%)
Frame = +2
Query: 375 SKWRLFKMVDIFVKLNNLVPKFESLFPNSLVNEAIAVRNRLGDASRVLFMKMHNFIFRVP 434
S W L KM+ IF L++L+ KF +S+ A+ V+NRLG+A R LF+K++ FRVP
Sbjct: 44 SIWCLSKMLAIFETLHHLISKFHLGLDSSVKEAAVTVQNRLGEAIRDLFLKLNYLTFRVP 223
Query: 435 AAKQVVSSYGQHHQMTIQVMSYVSSACRKRRKLEQILEEYPEVHNEVEASSFFLKQMEQI 494
AAK+V S G+HH +Q++SYV+SACR R LEQ+L+EYP+V+N V F++QME I
Sbjct: 224 AAKKVSRSDGRHHPTAVQIISYVTSACRSRHTLEQVLQEYPKVNNGVVVKDSFIEQMEWI 403
Query: 495 MRMLQRKLIVKSENCKDRALRHIFMLNNRSHIEAMNKFSRLETIFGNDWFQNNKAKIQQN 554
M ML+++L KS+ ++ ALR++FM+NNR HIEAM K LETIFGNDWFQ N+AK QQ+
Sbjct: 404 MDMLEKRLTYKSKEYRELALRYLFMMNNRRHIEAMIKSLDLETIFGNDWFQRNQAKFQQD 583
Query: 555 LDLYKRSAWDEVMDFLKLDNNE------SITKELLKEKIHLFNNRFE 595
LDLY+R +W++V++FLKLDNN+ + +++LKEK+ LFN +FE
Sbjct: 584 LDLYQRYSWNKVLEFLKLDNNDCAALNGDVAEDILKEKLKLFNKQFE 724
Score = 82.4 bits (202), Expect(2) = 4e-81
Identities = 41/68 (60%), Positives = 49/68 (71%)
Frame = +1
Query: 595 EAICRVQSAWFIYGSQLRGEIISSVGNILLPAYGIFVGRLHGILGNQAYKYIKYGMIEIQ 654
+AI RVQS WF+Y +L+ EII SV N LLP YGIF+GR LG A + I+YGM EIQ
Sbjct: 712 QAI*RVQSNWFVYDKKLKEEIIISVANTLLPVYGIFIGRFRDCLGIHANQCIRYGMFEIQ 891
Query: 655 DLLNHLFL 662
D LN+LFL
Sbjct: 892 DRLNNLFL 915
>TC89834
Length = 771
Score = 285 bits (728), Expect = 5e-77
Identities = 142/183 (77%), Positives = 161/183 (87%), Gaps = 3/183 (1%)
Frame = +2
Query: 2 IPVIIRIRMCLLQTKVWRFVGFASAAVGLLCYALSTSFNHLFGNWNLLKIFLYTLFSFI- 60
+ ++I+++ LLQTKVWRFVGFASAAVGL+CYALS+SFN+LFGNWNLLKI LYT+FSFI
Sbjct: 206 VHMLIKVQRWLLQTKVWRFVGFASAAVGLVCYALSSSFNYLFGNWNLLKIILYTVFSFII 385
Query: 61 --MILYANIWKNSRSLRFKAHTAFLVLTITSVYSFFFDKVVNGKPDAYSLISCAAFAIMS 118
MILY NIWK SRSLRFKAHTAFLVLTITSVYSF+FDKVVNGKPDAYSLISCAAFAIMS
Sbjct: 386 CIMILYENIWKQSRSLRFKAHTAFLVLTITSVYSFYFDKVVNGKPDAYSLISCAAFAIMS 565
Query: 119 MSLSRQSQCGIEVDLLYFFLGCLIMQLMKIKLQLVIVGAGFSYSLIIIRSSLSSTNSVPE 178
+S SRQSQCG EVDLLYFFLGCL + LMKIKL+L I+GAG SYSL+I RSS SS +
Sbjct: 566 LSFSRQSQCGFEVDLLYFFLGCLTVLLMKIKLELFIIGAGLSYSLLIFRSSFSSVXXIGL 745
Query: 179 NEY 181
+E+
Sbjct: 746 SEF 754
>BG454691 similar to GP|8347078|emb| NADH dehydrogenase subunit 4 {Sciurus
vulgaris}, partial (3%)
Length = 589
Score = 278 bits (712), Expect = 4e-75
Identities = 139/188 (73%), Positives = 166/188 (87%), Gaps = 3/188 (1%)
Frame = +2
Query: 12 LLQTKVWRFVGFASAAVGLLCYALSTSFNHLFGNWNLLKIFLYTLFSFIM---ILYANIW 68
++Q KVWRFVGFAS+A+GLLCYALS+SFN+LFG+WNLLKI LY++FSFI+ +L+A +W
Sbjct: 26 MMQPKVWRFVGFASSAIGLLCYALSSSFNYLFGDWNLLKIILYSIFSFIISLVVLFARMW 205
Query: 69 KNSRSLRFKAHTAFLVLTITSVYSFFFDKVVNGKPDAYSLISCAAFAIMSMSLSRQSQCG 128
+NSRSLRFKAH AFLVLTITS+YSFFFDKVVNGKPDAYSLISCA+F+IM MSLSRQ+QCG
Sbjct: 206 QNSRSLRFKAHAAFLVLTITSLYSFFFDKVVNGKPDAYSLISCASFSIMLMSLSRQTQCG 385
Query: 129 IEVDLLYFFLGCLIMQLMKIKLQLVIVGAGFSYSLIIIRSSLSSTNSVPENEYFGVGLRG 188
EVD LYFFLGCLI+QLMKIKLQL IVGAGFSY +II+RSS SS + V +NE+ GL
Sbjct: 386 FEVDFLYFFLGCLIVQLMKIKLQLFIVGAGFSYFVIILRSSFSSIDFVIDNEH-PTGLXN 562
Query: 189 ENSVVIEM 196
E+ +VIE+
Sbjct: 563 ESRIVIEV 586
>BG648139
Length = 753
Score = 274 bits (701), Expect = 7e-74
Identities = 148/241 (61%), Positives = 179/241 (73%), Gaps = 7/241 (2%)
Frame = -1
Query: 1 MIPVIIRIRMCLLQTKVWRFVGFASAAVGLLCYALSTSFNHLFGNWNLLKIFLYTLFSFI 60
M+P++I+I L+Q KVWRFVGFASA VGL+CYALS+SFN+LFG WNL KI LY++ SFI
Sbjct: 687 MMPILIQILRWLMQEKVWRFVGFASAIVGLVCYALSSSFNYLFGEWNLSKIILYSVISFI 508
Query: 61 ---MILYANIWKNSRSLRFKAHTAFLVLTITSVYSFFFDKVVNGKPDAYSLISCAAFAIM 117
MILY +W++SR L+FKAHTAF VLT+TSVYSF +DKVVN KPDAYSLISCAAFAIM
Sbjct: 507 ICLMILYPKVWQSSRGLKFKAHTAFFVLTLTSVYSFIYDKVVNEKPDAYSLISCAAFAIM 328
Query: 118 SMSLS----RQSQCGIEVDLLYFFLGCLIMQLMKIKLQLVIVGAGFSYSLIIIRSSLSST 173
S+SLS RQ+QCG EVDLLYFFLGCLI+QLMK+KLQL +VG GFSY +II RS+ SS
Sbjct: 327 SLSLSRQTERQTQCGFEVDLLYFFLGCLIVQLMKVKLQLAVVGVGFSYFVIIFRSAFSS- 151
Query: 174 NSVPENEYFGVGLRGENSVVIEMDSLLRPSNDIYSGMMQQLTTCINALQQDSLNIIDRLM 233
S N Y GL ENSVV+E+DSLL LQ ++ N++D ++
Sbjct: 150 -SAARNGY--AGLPSENSVVVEVDSLL-----------------TRTLQLENPNLMDMIL 31
Query: 234 E 234
E
Sbjct: 30 E 28
>TC84645
Length = 672
Score = 213 bits (543), Expect = 1e-55
Identities = 105/176 (59%), Positives = 140/176 (78%), Gaps = 4/176 (2%)
Frame = +2
Query: 4 VIIRIRMCLLQTKVWRFVGFASAAVGLLCYALSTSFNHLFGNWNLLKIFLYTLFSF---I 60
V+ +I C++Q KVWRFVGFA+ GLLC ALS+SFN+LFG W +L I LYT+FSF +
Sbjct: 98 VLFQIWRCMMQPKVWRFVGFAAVVAGLLCNALSSSFNYLFGGWTMLVIALYTVFSFALCV 277
Query: 61 MILYANIWKNSRSLRFKAHTAFLVLTITSVYSFFFDKVVNGKPDAYSLISCAAFAIMSMS 120
++L+ IW++SRS F A+T F+VL +TS+YS+FFDK+++ PDAY+LISCA+FA+ ++S
Sbjct: 278 LVLFPRIWQHSRSHLFIANTTFVVLAMTSLYSYFFDKLMHNTPDAYTLISCASFAVSALS 457
Query: 121 LSR-QSQCGIEVDLLYFFLGCLIMQLMKIKLQLVIVGAGFSYSLIIIRSSLSSTNS 175
LSR ++QCG E+DLLYF LGCL+MQLMKI L+L I GA F Y LIIIRSS SS ++
Sbjct: 458 LSRNKTQCGFEIDLLYFLLGCLMMQLMKITLKLFIFGAVFXYFLIIIRSSFSSIDA 625
>BF636319 weakly similar to PIR|G71155|G711 hypothetical protein PH0446 -
Pyrococcus horikoshii, partial (5%)
Length = 595
Score = 166 bits (420), Expect = 3e-41
Identities = 90/174 (51%), Positives = 111/174 (63%), Gaps = 3/174 (1%)
Frame = +2
Query: 6 IRIRMCLLQTKVWRFVGFASAAVGLLCYALSTSFNHLFGNWNLLKIFLYTLFSFI---MI 62
IRI L+Q WR +GF S+ +G + YA S+SFNHLFG WN +KI +YT+ SF M+
Sbjct: 68 IRISNRLIQKNQWRILGFFSSIMGFISYASSSSFNHLFGKWNFIKIIIYTVVSFSISSMM 247
Query: 63 LYANIWKNSRSLRFKAHTAFLVLTITSVYSFFFDKVVNGKPDAYSLISCAAFAIMSMSLS 122
L+ WK S+ +AH LVL +TS+YSF DK N KPD S+ISC AFA+MS LS
Sbjct: 248 LFLKKWKLSKRFLLRAHLGVLVLLLTSLYSFVSDKASNEKPDLLSMISCVAFALMSFCLS 427
Query: 123 RQSQCGIEVDLLYFFLGCLIMQLMKIKLQLVIVGAGFSYSLIIIRSSLSSTNSV 176
RQ G E DLL FFLG L +QLMKI L L I A YSLI +RS L S + +
Sbjct: 428 RQIDLGFESDLLNFFLGILTVQLMKINLILSIXAAILCYSLIGLRSKLGSRHEI 589
>TC90298
Length = 1593
Score = 130 bits (328), Expect = 1e-30
Identities = 68/96 (70%), Positives = 77/96 (79%), Gaps = 3/96 (3%)
Frame = +1
Query: 46 WNLLKIFLYTLFSFIM---ILYANIWKNSRSLRFKAHTAFLVLTITSVYSFFFDKVVNGK 102
WNLLKI LYT FSFI+ IL+AN W+NS SLR +AH F V TIT+VYS+FFDKV NGK
Sbjct: 64 WNLLKIILYTFFSFIICLAILFANKWQNSPSLRMRAHLVFSVFTITTVYSYFFDKV-NGK 240
Query: 103 PDAYSLISCAAFAIMSMSLSRQSQCGIEVDLLYFFL 138
PD YSL+S AAFAIMS+ LS+QS G EVDLLYFFL
Sbjct: 241 PDVYSLVSGAAFAIMSLGLSKQSHFGFEVDLLYFFL 348
Score = 47.8 bits (112), Expect = 1e-05
Identities = 36/107 (33%), Positives = 51/107 (47%), Gaps = 5/107 (4%)
Frame = +2
Query: 128 GIEVDLLYFFLGCLIMQLMKIKLQLVIVGAGFSYSLIIIRSSLSSTNSVPEN-----EYF 182
G ++ FF G L +Q MKIKL LVI GA FSY LII+R L E +Y
Sbjct: 317 GSKLIFCIFFCGYLTLQFMKIKLFLVIAGAIFSYPLIILRFYLHVPTEYSEELTNTADYH 496
Query: 183 GVGLRGENSVVIEMDSLLRPSNDIYSGMMQQLTTCINALQQDSLNII 229
V + G + DSL+ P D + + ++ + QQ ++ I
Sbjct: 497 VVQIDGSQ---VNFDSLVAPDFDAFPDSLTLVSQINSPSQQVNIGHI 628
Score = 43.9 bits (102), Expect = 2e-04
Identities = 19/34 (55%), Positives = 24/34 (69%)
Frame = +3
Query: 26 AAVGLLCYALSTSFNHLFGNWNLLKIFLYTLFSF 59
A GLLCYALS+SFNHLFGN ++ + +F F
Sbjct: 3 AVAGLLCYALSSSFNHLFGNLEFVEDYSLHIFQF 104
>TC87069 weakly similar to GP|9758877|dbj|BAB09431.1 contains similarity to
disease resistance protein~gene_id:K24G6.11 {Arabidopsis
thaliana}, partial (4%)
Length = 1310
Score = 114 bits (286), Expect = 9e-26
Identities = 69/187 (36%), Positives = 98/187 (51%), Gaps = 8/187 (4%)
Frame = +3
Query: 16 KVWRFVGFASAAVGLLCYALSTSFNHLFGNWNLLKIFLYTLFSFIMILYANIWKNSRSLR 75
K WR VG S+ VGLLCYALS SFN L G W K FLY + S ++ K S +
Sbjct: 672 KTWRVVGLMSSVVGLLCYALSPSFNRLIGRWKPFKFFLYGVSSLVIFTTVLFAKQSSLPK 851
Query: 76 -----FKAHTAFLVLTITSVYSFFFDKVVNGKPDAYSLISCAAFAIMSMSLSRQSQCGIE 130
K T F VL I VYSF +D+ VNGKP+ S++S A+FA++S+SL + + E
Sbjct: 852 QHAQFIKTCTIFAVLVII*VYSFVYDRAVNGKPNILSVVSNASFALVSLSLHKLFKLKSE 1031
Query: 131 VDLLYFFLGCLIMQLMKIKLQLVIVGAGFSYSLIIIRSSLSS---TNSVPENEYFGVGLR 187
+ + +FL C +QL+ I L+ V + + L ++ S S NS V
Sbjct: 1032MGIFSYFLCCFTVQLLTINWILIFVAIIYCFLLFVMHSQPSGGQVINSSTSYSEIAVDSG 1211
Query: 188 GENSVVI 194
G++ + I
Sbjct: 1212GQHVIAI 1232
>BE321047 similar to GP|10177020|db leucine zipper protein {Arabidopsis
thaliana}, partial (25%)
Length = 495
Score = 110 bits (275), Expect = 2e-24
Identities = 61/151 (40%), Positives = 91/151 (59%), Gaps = 4/151 (2%)
Frame = +1
Query: 310 LDTMFQRWMTASDIATTVLFPIEQKFCDLVFSGFSSATSHCFIEICQEATFQLLNFADVI 369
L+ +RW+ S +A +LF E++ CD VF G S+ F +IC+E+ QLLNFA+ I
Sbjct: 46 LEDEIKRWIKVSKVALRILFRSERRLCDQVFFGLSTTADLSFTDICRESMLQLLNFAEAI 225
Query: 370 AYGSLSKWRLFKMVDIFVKLNNLVPKFESLFPN----SLVNEAIAVRNRLGDASRVLFMK 425
A GS S RLF+++D+F + +L+P+FESLF + S+ NEA + RLG+A +FM+
Sbjct: 226 AIGSRSPERLFRVLDMFETMRDLIPEFESLFRDQYNGSMQNEATTIWKRLGEAIIGIFME 405
Query: 426 MHNFIFRVPAAKQVVSSYGQHHQMTIQVMSY 456
+ N I P + V G H +T VM+Y
Sbjct: 406 LENLICHDPMNLEAVPG-GGIHPITHYVMNY 495
>TC83992 weakly similar to GP|10177020|dbj|BAB10258. leucine zipper protein
{Arabidopsis thaliana}, partial (14%)
Length = 615
Score = 88.2 bits (217), Expect = 9e-18
Identities = 46/107 (42%), Positives = 68/107 (62%), Gaps = 6/107 (5%)
Frame = +3
Query: 564 DEVMDFLKLDNNESITK----ELLKEKIHLFNNRFEAICRVQSAWFIYGSQLRGEIISSV 619
++V FLK++NN S+ + + +KEK+ FN F+ +CRVQS WFI+ QL+ EI S+
Sbjct: 3 NKVFGFLKVENNGSMQQNGVAKSMKEKLKSFNMMFDDLCRVQSTWFIFDEQLKEEIRISI 182
Query: 620 GNILLPAYGIFVGRLHGI--LGNQAYKYIKYGMIEIQDLLNHLFLGN 664
+LLPAY F+ R + +G A KY+KYG +I+ LN LF G+
Sbjct: 183 EKLLLPAYANFIARFQNVAEVGKHADKYVKYGTEDIEAKLNDLFQGS 323
>BG587084 homologue to GP|6002696|gb| histone acetyltransferase MORF beta
{Homo sapiens}, partial (1%)
Length = 685
Score = 86.7 bits (213), Expect = 3e-17
Identities = 63/215 (29%), Positives = 105/215 (48%), Gaps = 20/215 (9%)
Frame = +1
Query: 391 NLVPKFESLFPN----SLVNEAIAVRNRLGDASRVLFMKMHNFIFRVPAAKQVVSSYGQH 446
+L+P FESLF + SL NE V +LG+ + N I R GQ
Sbjct: 1 DLIPNFESLFCDQYSVSLRNELNTVLKKLGETIVGTLREFENTI-RSKGPGNAPFFGGQL 177
Query: 447 HQMTIQVMSYVSSACRKRRKLEQILEEYPEV------HNEV----------EASSFFLKQ 490
H + VM++++ C R LEQ+ E++ V H++ +SS Q
Sbjct: 178 HPLVRFVMNFLTWICDYREILEQVFEDHGHVLLEYTKHDDTVPSSSSSSSSSSSSSSSLQ 357
Query: 491 MEQIMRMLQRKLIVKSENCKDRALRHIFMLNNRSHIEAMNKFSRLETIFGNDWFQNNKAK 550
ME+IM +L+ KL D L +++++N+ +I + L T+ G+ Q + AK
Sbjct: 358 MERIMEVLESKLEAMFNIFNDPTLGYVYLMNSSRYIIIKTMENELGTLLGDGMLQRHSAK 537
Query: 551 IQQNLDLYKRSAWDEVMDFLKLDNNESITKELLKE 585
++ N + Y RS+W +V++FL+LDNN + ++ E
Sbjct: 538 LRYNFEEYIRSSWGKVLEFLRLDNNLLVHPNMVGE 642
>BE203379 similar to GP|22136732|gb unknown protein {Arabidopsis thaliana},
partial (28%)
Length = 584
Score = 74.3 bits (181), Expect = 1e-13
Identities = 46/165 (27%), Positives = 88/165 (52%)
Frame = +1
Query: 408 AIAVRNRLGDASRVLFMKMHNFIFRVPAAKQVVSSYGQHHQMTIQVMSYVSSACRKRRKL 467
A+A+ +L ++ F + + A K V+ G H +T V++YV R L
Sbjct: 34 AMALTKKLAQTAQETFGDFEEAVEK-DATKTAVTD-GTVHPLTSYVINYVKFLFDYRSTL 207
Query: 468 EQILEEYPEVHNEVEASSFFLKQMEQIMRMLQRKLIVKSENCKDRALRHIFMLNNRSHIE 527
+Q+ +E+ ++ + ++ ++ IM+ LQ L KS+ KD AL H+F++NN +I
Sbjct: 208 KQLFQEFEGGNDSSQLATVTMR----IMQTLQINLDGKSKQYKDLALTHLFLMNNIHYIV 375
Query: 528 AMNKFSRLETIFGNDWFQNNKAKIQQNLDLYKRSAWDEVMDFLKL 572
+ S + + G+DW Q ++ +QQ+ + YKR+AW +++ L +
Sbjct: 376 RSVRRSEAKDLLGDDWVQRHRRIVQQHANQYKRNAWAKILQCLSI 510
>TC90397 similar to GP|15010740|gb|AAK74029.1 AT5g59730/mth12_130
{Arabidopsis thaliana}, partial (22%)
Length = 1162
Score = 73.2 bits (178), Expect = 3e-13
Identities = 57/234 (24%), Positives = 112/234 (47%), Gaps = 9/234 (3%)
Frame = +3
Query: 234 EEYNQGKCDFEMDTLPSEKLHNLHEIVKLMLCAGYEKECSAVYISWRKVLLQKGL----L 289
E++ G E + + + +L I M+ GY KEC VYI RK ++ + L +
Sbjct: 390 EKFVVGNQISETERVSMLAMADLKAIADCMINCGYGKECVKVYIVMRKSIVDEALYHLGI 569
Query: 290 NKIFVLPEAKINTERERERYLDTMFQRWMTASDIATTVLFPIEQKFCDLVFSGFSSATSH 349
++ K++ E ++ + W+ A +A LF E+ CD VF+ +
Sbjct: 570 ERLTFSQIQKMDWE-----VIELKIKTWLKAVKVAVRTLFHGERILCDDVFAAAGKRIAE 734
Query: 350 -CFIEICQEATFQLLNFADVIAYGSLSKWRLFKMVDIFVKLNNLVPKFESLF----PNSL 404
CF EI +E L F D++A + ++F+ +D++ +++ + +S+F +++
Sbjct: 735 SCFAEITKEGATSLFTFPDMVAKCKKTPEKMFRTLDLYEAISDHFQQIQSIFSFESTSNV 914
Query: 405 VNEAIAVRNRLGDASRVLFMKMHNFIFRVPAAKQVVSSYGQHHQMTIQVMSYVS 458
+AI +L +A + + + + I + + KQV S G H +T VM+Y++
Sbjct: 915 RLQAINSMEKLAEAVKTMLKEFESAIQKDSSKKQV--SGGGVHPLTRYVMNYLT 1070
>TC90683 similar to GP|22136732|gb|AAM91685.1 unknown protein {Arabidopsis
thaliana}, partial (19%)
Length = 554
Score = 68.9 bits (167), Expect = 6e-12
Identities = 37/126 (29%), Positives = 68/126 (53%), Gaps = 11/126 (8%)
Frame = +3
Query: 547 NKAKIQQNLDLYKRSAWDEVMDFLKL---------DNNESITKELLKEKIHLFNNRFEAI 597
++ +QQ+ + YKR +W +++ L + D+N +I++ +KE+ FN +FE +
Sbjct: 6 HRRTVQQHANQYKRISWAKILQRLTVQGGNSSGGGDSNRNISRSSVKERFKTFNTQFEEL 185
Query: 598 CRVQSAWFIYGSQLRGEIISSVGNILLPAYGIFVGRLHGIL--GNQAYKYIKYGMIEIQD 655
+ QS W + S+LR + +V +LLPAY F+ R ++ G KYI+Y ++
Sbjct: 186 HQRQSQWTVPDSELRESLRLAVAEVLLPAYRSFLKRFGPMIENGKNPQKYIRYSPENLEQ 365
Query: 656 LLNHLF 661
+L F
Sbjct: 366 MLGEFF 383
>AW328903 weakly similar to GP|8439895|gb|A It contains a interferon
alpha/beta domain PF|00143. EST gb|N96176 comes from
this gene., partial (15%)
Length = 485
Score = 68.2 bits (165), Expect = 1e-11
Identities = 41/142 (28%), Positives = 73/142 (50%)
Frame = +3
Query: 489 KQMEQIMRMLQRKLIVKSENCKDRALRHIFMLNNRSHIEAMNKFSRLETIFGNDWFQNNK 548
K++ ++ ++ KL K+E KD AL ++F+ NN ++ + S L I G DW ++
Sbjct: 15 KRLSWLILVVLCKLDGKAEFYKDVALSYLFLANNMQYVVVKVRKSNLRFILGEDWLLKHE 194
Query: 549 AKIQQNLDLYKRSAWDEVMDFLKLDNNESITKELLKEKIHLFNNRFEAICRVQSAWFIYG 608
K+++ + Y+R AW +V+ + E+ T E E FN F+ R+Q W +
Sbjct: 195 MKVKEYVTKYERMAWSKVLSSIP----ENPTVEKASENFQGFNVEFDEAFRMQYLWVVPD 362
Query: 609 SQLRGEIISSVGNILLPAYGIF 630
+LR EI S+ + ++ Y F
Sbjct: 363 LELRNEIKESLVSKIVFKYREF 428
>BE999174 similar to GP|11546040|emb bA218B22.1.1 (novel protein (presumed
ortholog of mouse diaphenous-related formin (DIA2)),
partial (4%)
Length = 416
Score = 57.0 bits (136), Expect = 2e-08
Identities = 35/99 (35%), Positives = 55/99 (55%), Gaps = 5/99 (5%)
Frame = +1
Query: 45 NWNLLKIFLYTLFSFIMILYANIWKNSRSLR-----FKAHTAFLVLTITSVYSFFFDKVV 99
+W+ K FLY + S ++ K S + K T F VL I VYSF +D+ V
Sbjct: 121 HWDF-KFFLYGVSSLVIFTTVLFAKQSSLPKQHAQFIKTCTIFAVLVII*VYSFVYDRAV 297
Query: 100 NGKPDAYSLISCAAFAIMSMSLSRQSQCGIEVDLLYFFL 138
NGKP+ S++S A+FA++S+SL + + E+ + +FL
Sbjct: 298 NGKPNILSVVSNASFALVSLSLHKLFKLKSEMGIFSYFL 414
>AL370979 weakly similar to GP|15010740|gb| AT5g59730/mth12_130 {Arabidopsis
thaliana}, partial (14%)
Length = 455
Score = 56.6 bits (135), Expect = 3e-08
Identities = 32/122 (26%), Positives = 66/122 (53%), Gaps = 2/122 (1%)
Frame = +2
Query: 483 ASSFFLKQMEQIMRMLQRKLIVKSENCKDRALRHIFMLNNRSHIEAMNKFSRLETIFGND 542
+SS +++ ++ +L KL K+E KD AL ++F+ NN ++ + S L + G +
Sbjct: 89 SSSEISEKIAWLILVLLCKLDTKAEFYKDVALSYLFLANNMQYVVVKVRRSNLGFLLGEE 268
Query: 543 WFQNNKAKIQQNLDLYKRSAWDEVMDFLKLDNNESITK--ELLKEKIHLFNNRFEAICRV 600
W N++ K+++ ++ + + W++V+ L + N + K E +K FN F+ C+
Sbjct: 269 WLTNHELKVKEYVNKFVQIGWNKVLSTLPENENSTAEKTVEQVKAIFVKFNAAFDEECKK 448
Query: 601 QS 602
Q+
Sbjct: 449 QT 454
>BE325447 weakly similar to GP|18700111|gb| At1g07000/F10K1_20 {Arabidopsis
thaliana}, partial (7%)
Length = 422
Score = 52.4 bits (124), Expect = 6e-07
Identities = 35/127 (27%), Positives = 62/127 (48%), Gaps = 4/127 (3%)
Frame = +3
Query: 449 MTIQVMSYVSSACRKRRKLEQILEEYPEVHNEVEASSFFLKQMEQIMRMLQRKLIVKSEN 508
+T VM+Y+ CR ++ LEQ+ + S ++ +I+ L+ L KS+
Sbjct: 15 ITQHVMNYLRVVCRSQQTLEQVFYD-----------SSLSSKIHRIIDTLESNLEAKSKC 161
Query: 509 CKDRALRHIFMLNNRSHIEAMNKFSRLETIFGNDWFQNNKAKIQQNLDLYKRS----AWD 564
D +L +IF++NN ++I M K + L T+ G+ W Q K+ Y ++ A
Sbjct: 162 YVDPSLGYIFLINNHTYIVEMTKDNELGTLLGDYWLQKYTEKVWHYHRQYHKTKGAIASS 341
Query: 565 EVMDFLK 571
V+DF +
Sbjct: 342 VVLDFFR 362
Database: MTGI
Posted date: Oct 22, 2004 3:39 PM
Number of letters in database: 27,044,181
Number of sequences in database: 36,976
Lambda K H
0.327 0.139 0.412
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 21,513,257
Number of Sequences: 36976
Number of extensions: 312414
Number of successful extensions: 2304
Number of sequences better than 10.0: 53
Number of HSP's better than 10.0 without gapping: 2257
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 2278
length of query: 672
length of database: 9,014,727
effective HSP length: 103
effective length of query: 569
effective length of database: 5,206,199
effective search space: 2962327231
effective search space used: 2962327231
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 15 ( 7.1 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 40 (21.7 bits)
S2: 62 (28.5 bits)
Medicago: description of AC138012.9


