

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC137996.5 + phase: 0
(713 letters)
Database: MTGI
36,976 sequences; 27,044,181 total letters
Searching..................................................done
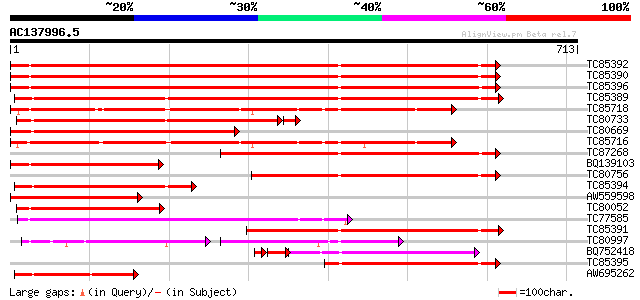
Score E
Sequences producing significant alignments: (bits) Value
TC85392 homologue to PIR|JC4786|JC4786 dnaK-type molecular chape... 725 0.0
TC85390 homologue to SP|P27322|HS72_LYCES Heat shock cognate 70 ... 724 0.0
TC85396 homologue to GP|6911551|emb|CAB72129.1 heat shock protei... 720 0.0
TC85389 homologue to SP|Q03684|BIP4_TOBAC Luminal binding protei... 561 e-160
TC85718 homologue to SP|Q01899|HS7M_PHAVU Heat shock 70 kDa prot... 435 e-122
TC80733 homologue to SP|Q01233|HS70_NEUCR Heat shock 70 kDa prot... 417 e-120
TC80669 homologue to PIR|S53498|S53498 dnaK-type molecular chape... 420 e-118
TC85716 homologue to PIR|S19140|S19140 dnaK-type molecular chape... 419 e-117
TC87268 homologue to PIR|S53500|S44168 dnaK-type molecular chape... 333 2e-91
BQ139103 homologue to GP|2655420|gb| heat shock cognate protein ... 283 1e-76
TC80756 homologue to PIR|S53498|S53498 dnaK-type molecular chape... 283 2e-76
TC85394 homologue to GP|2642238|gb|AAB86942.1| endoplasmic retic... 271 5e-73
AW559598 weakly similar to SP|P09189|HS7C Heat shock cognate 70 ... 261 9e-70
TC80052 similar to PIR|JC4786|JC4786 dnaK-type molecular chapero... 252 3e-67
TC77585 similar to GP|17473863|gb|AAL38353.1 putative heat-shock... 239 3e-63
TC85391 homologue to PIR|T06598|T06598 dnaK-type molecular chape... 236 2e-62
TC80997 similar to PIR|B84729|B84729 70kD heat shock protein [im... 134 7e-61
BQ752418 homologue to SP|P78695|GR78 78 kDa glucose-regulated pr... 168 3e-48
TC85395 homologue to GP|6911553|emb|CAB72130.1 heat shock protei... 182 3e-46
AW695262 homologue to GP|170386|gb|A glucose-regulated protein 7... 165 5e-41
>TC85392 homologue to PIR|JC4786|JC4786 dnaK-type molecular chaperone
hsc70-3 - tomato, complete
Length = 2295
Score = 725 bits (1871), Expect = 0.0
Identities = 386/621 (62%), Positives = 475/621 (76%), Gaps = 4/621 (0%)
Frame = +1
Query: 1 MARKYEGRAVGIDLGTTYSCVAVWLDDHNRVEIIHNDQGNRTTPSVVAFTDYQRLIGNGA 60
MA K EG A+GIDLGTTYSCV VW H+RVEII NDQGNRTTPS VAFTD +RLIG+ A
Sbjct: 130 MAGKGEGPAIGIDLGTTYSCVGVW--QHDRVEIIANDQGNRTTPSYVAFTDTERLIGDAA 303
Query: 61 ENQSATNPENTVFDAKRLIGRKYSDPIVQKDIMLWPFMVTSGVNDKPMITVKYKGQEKQF 120
+NQ A NP NTVFDAKRLIGR+ SD VQ D+ LWPF V G DKPMI V YK +EKQF
Sbjct: 304 KNQVAMNPTNTVFDAKRLIGRRISDVSVQSDMKLWPFKVIPGPADKPMIVVNYKAEEKQF 483
Query: 121 CAEEISSAVLTKMREIAEAYLGSPVKNVVVTVPAYFNDSQRKATIDAGAIAGLNVIRILN 180
AEEISS VL KM+EIAEAYLG+ +KN VVTVPAYFNDSQR+AT DAG I+GLNV+RI+N
Sbjct: 484 SAEEISSMVLMKMKEIAEAYLGTTIKNAVVTVPAYFNDSQRQATKDAGVISGLNVMRIIN 663
Query: 181 EPTAAAIAYGLDKRSNYDGKRNILVFDLGGGTFDVSILTIKGDVFDVKATAGNTHLGGED 240
EPTAAAIAYGLDK++ G++N+L+FDLGGGTFDVS+LTI+ +F+VKATAG+THLGGED
Sbjct: 664 EPTAAAIAYGLDKKATSTGEKNVLIFDLGGGTFDVSLLTIEEGIFEVKATAGDTHLGGED 843
Query: 241 FDNRMVYYFVEEFKKKNKVEISGNPKSLRRLRTACERAKRILSFTFVTTVEVDALFMGID 300
FDNRMV +FV+EFK+KNK +ISGNP++LRR+RTACERAKR LS T TT+E+D+LF GID
Sbjct: 844 FDNRMVNHFVQEFKRKNKKDISGNPRALRRVRTACERAKRTLSSTAQTTIEIDSLFEGID 1023
Query: 301 FSSSITRAKFEEINMDFFNECINIVDSCLRDSKIYKNDIDDVVLVGGSSRIPKVQDLLLE 360
F ++ITRA+FEE+NMD F +C+ V+ CLRD+K+ KN +DDVVLVGGS+RIPKVQ LL +
Sbjct: 1024FYTTITRARFEELNMDLFRKCMEPVEKCLRDAKMDKNSVDDVVLVGGSTRIPKVQQLLQD 1203
Query: 361 FFKGKALFMNINPDEAIAYGAAVQAAILS-EDFKNVPNLVLRDVTPLSLGIATATYWENI 419
FF GK L +INPDEA+AYGAAVQAAILS E + V +L+L DVTPLSLG+ TA +
Sbjct: 1204FFNGKELCKSINPDEAVAYGAAVQAAILSGEGNEKVQDLLLLDVTPLSLGLETA---GGV 1374
Query: 420 MDVVIPRNTSIPVKNTKGYCTAIDN-CGASIIVYEGERPRASDNNLLGLFTLSCL-PGPR 477
M V+IPRNT+IP K + + T DN G I VYEGER R DNNLLG F LS + P PR
Sbjct: 1375MTVLIPRNTTIPTKKEQVFSTYSDNQPGVLIQVYEGERTRTRDNNLLGKFELSGIPPAPR 1554
Query: 478 GQP-LEVCFSIDENGILTVSGKDITTGILNEITITNEKERLSKFEIEKKIEEAEKYRVED 536
G P + VCF ID NGIL VS +D TTG N+ITITN+K RLSK +IEK ++EAEKY+ ED
Sbjct: 1555GVPQITVCFDIDANGILNVSAEDKTTGQKNKITITNDKGRLSKEDIEKMVQEAEKYKSED 1734
Query: 537 MKFLRKAKVISALDSCVYNIKSALNMKDVDLILSPQESEQINNAIIVAMNLLEMNKNNQH 596
+ RK + +AL++ YN+++ + + LS + ++I +AI A+ L+ NQ
Sbjct: 1735EEHKRKVEAKNALENYAYNMRNTIKDDKIASKLSADDKKKIEDAIEGAIQWLD---GNQL 1905
Query: 597 IEIDVLEHHLKEMNSMLNMLL 617
E D E +KE+ + N ++
Sbjct: 1906GEADEFEDKMKELEGICNPII 1968
>TC85390 homologue to SP|P27322|HS72_LYCES Heat shock cognate 70 kDa protein
2. [Tomato] {Lycopersicon esculentum}, complete
Length = 2278
Score = 724 bits (1868), Expect = 0.0
Identities = 385/621 (61%), Positives = 474/621 (75%), Gaps = 4/621 (0%)
Frame = +1
Query: 1 MARKYEGRAVGIDLGTTYSCVAVWLDDHNRVEIIHNDQGNRTTPSVVAFTDYQRLIGNGA 60
MA K EG A+GIDLGTTYSCV VW H+RVEII NDQGNRTTPS VAFTD +RLIG+ A
Sbjct: 136 MAGKGEGPAIGIDLGTTYSCVGVW--QHDRVEIIANDQGNRTTPSYVAFTDTERLIGDAA 309
Query: 61 ENQSATNPENTVFDAKRLIGRKYSDPIVQKDIMLWPFMVTSGVNDKPMITVKYKGQEKQF 120
+NQ A NP NTVFDAKRLIGR++ D VQ D+ LWPF V G DKPMI V YK +EKQF
Sbjct: 310 KNQVAMNPTNTVFDAKRLIGRRFGDVSVQSDMKLWPFKVIPGPADKPMIVVNYKAEEKQF 489
Query: 121 CAEEISSAVLTKMREIAEAYLGSPVKNVVVTVPAYFNDSQRKATIDAGAIAGLNVIRILN 180
AEEISS VL KM+EIAEAYLG+ +KN VVTVPAYFNDSQR+AT DAG I+GLNV+RI+N
Sbjct: 490 SAEEISSMVLMKMKEIAEAYLGTTIKNAVVTVPAYFNDSQRQATKDAGVISGLNVLRIIN 669
Query: 181 EPTAAAIAYGLDKRSNYDGKRNILVFDLGGGTFDVSILTIKGDVFDVKATAGNTHLGGED 240
EPTAAAIAYGLDK++ G++N+L+FDLGGGTFDVS+LTI+ +F+VKATAG+THLGGED
Sbjct: 670 EPTAAAIAYGLDKKATSTGEKNVLIFDLGGGTFDVSLLTIEEGIFEVKATAGDTHLGGED 849
Query: 241 FDNRMVYYFVEEFKKKNKVEISGNPKSLRRLRTACERAKRILSFTFVTTVEVDALFMGID 300
FDNRMV +FV+EFK+KNK +ISGNP++LRRLRTACERAKR LS T TT+E+D+LF GID
Sbjct: 850 FDNRMVNHFVQEFKRKNKKDISGNPRALRRLRTACERAKRTLSSTAQTTIEIDSLFEGID 1029
Query: 301 FSSSITRAKFEEINMDFFNECINIVDSCLRDSKIYKNDIDDVVLVGGSSRIPKVQDLLLE 360
F ++ITRA+FEE+NMD F +C+ V+ CLRD+K+ KN + DVVLVGGS+RIPKVQ LL +
Sbjct: 1030FYTTITRARFEELNMDLFRKCMEPVEKCLRDAKMDKNSVHDVVLVGGSTRIPKVQQLLQD 1209
Query: 361 FFKGKALFMNINPDEAIAYGAAVQAAILS-EDFKNVPNLVLRDVTPLSLGIATATYWENI 419
FF GK L +INPDEA+AYGAAVQAAILS E + V +L+L DVTPLSLG+ TA +
Sbjct: 1210FFNGKELCKSINPDEAVAYGAAVQAAILSGEGNEKVQDLLLLDVTPLSLGLETA---GGV 1380
Query: 420 MDVVIPRNTSIPVKNTKGYCTAIDN-CGASIIVYEGERPRASDNNLLGLFTLSCL-PGPR 477
M V+IPRNT+IP K + + T DN G I VYEGER R DNNLLG F LS + P PR
Sbjct: 1381MTVLIPRNTTIPTKKEQVFSTYSDNQPGVLIQVYEGERTRTRDNNLLGKFELSGIPPAPR 1560
Query: 478 GQP-LEVCFSIDENGILTVSGKDITTGILNEITITNEKERLSKFEIEKKIEEAEKYRVED 536
G P + VCF ID NGIL VS +D TTG N+ITITN+K RLSK EIEK ++EAEKY+ ED
Sbjct: 1561GVPQITVCFDIDANGILNVSAEDKTTGQKNKITITNDKGRLSKEEIEKMVQEAEKYKSED 1740
Query: 537 MKFLRKAKVISALDSCVYNIKSALNMKDVDLILSPQESEQINNAIIVAMNLLEMNKNNQH 596
+ +K + +AL++ YN+++ + + LS + ++I +AI A+ L+ NQ
Sbjct: 1741EEHKKKVEAKNALENYAYNMRNTIKDDKIASKLSADDKKKIEDAIEGAIQWLD---GNQL 1911
Query: 597 IEIDVLEHHLKEMNSMLNMLL 617
E D E +KE+ + N ++
Sbjct: 1912GEADEFEDKMKELEGICNPII 1974
>TC85396 homologue to GP|6911551|emb|CAB72129.1 heat shock protein 70
{Cucumis sativus}, complete
Length = 2320
Score = 720 bits (1858), Expect = 0.0
Identities = 383/621 (61%), Positives = 475/621 (75%), Gaps = 4/621 (0%)
Frame = +2
Query: 1 MARKYEGRAVGIDLGTTYSCVAVWLDDHNRVEIIHNDQGNRTTPSVVAFTDYQRLIGNGA 60
MA K EG A+GIDLGTTYSCV VW H+RVEII NDQGNRTTPS VAFTD +RLIG+ A
Sbjct: 107 MAGKGEGPAIGIDLGTTYSCVGVW--QHDRVEIIANDQGNRTTPSYVAFTDSERLIGDAA 280
Query: 61 ENQSATNPENTVFDAKRLIGRKYSDPIVQKDIMLWPFMVTSGVNDKPMITVKYKGQEKQF 120
+NQ A NP NTVFDAKRLIGR+ SD VQ D+ LWPF V +G +KPMI V YKG+EK F
Sbjct: 281 KNQVAMNPTNTVFDAKRLIGRRISDASVQSDMKLWPFKVIAGPGEKPMIGVNYKGEEKLF 460
Query: 121 CAEEISSAVLTKMREIAEAYLGSPVKNVVVTVPAYFNDSQRKATIDAGAIAGLNVIRILN 180
+EEISS VL KMREIAEAYLG +KN VVTVPAYFNDSQR+AT DAG IAGLNV+RI+N
Sbjct: 461 ASEEISSMVLIKMREIAEAYLGVTIKNAVVTVPAYFNDSQRQATKDAGVIAGLNVLRIIN 640
Query: 181 EPTAAAIAYGLDKRSNYDGKRNILVFDLGGGTFDVSILTIKGDVFDVKATAGNTHLGGED 240
EPTAAAIAYGLDK++ G++N+L+FDLGGGTFDVS+LTI+ +F+VKATAG+THLGGED
Sbjct: 641 EPTAAAIAYGLDKKATSVGEKNVLIFDLGGGTFDVSLLTIEEGIFEVKATAGDTHLGGED 820
Query: 241 FDNRMVYYFVEEFKKKNKVEISGNPKSLRRLRTACERAKRILSFTFVTTVEVDALFMGID 300
FDNRMV +FV+EFK+KNK +ISGNP++LRRLRTACERAKR LS T TT+E+D+LF G+D
Sbjct: 821 FDNRMVTHFVQEFKRKNKKDISGNPRALRRLRTACERAKRTLSSTAQTTIEIDSLFEGVD 1000
Query: 301 FSSSITRAKFEEINMDFFNECINIVDSCLRDSKIYKNDIDDVVLVGGSSRIPKVQDLLLE 360
F ++ITRA+FEE+NMD F +C+ V+ CLRD+K+ K+ + DVVLVGGS+RIPKVQ LL +
Sbjct: 1001FYTTITRARFEELNMDLFRKCMEPVEKCLRDAKMDKSTVHDVVLVGGSTRIPKVQQLLQD 1180
Query: 361 FFKGKALFMNINPDEAIAYGAAVQAAILS-EDFKNVPNLVLRDVTPLSLGIATATYWENI 419
FF GK L +INPDEA+AYGAAVQAAILS E + V +L+L DVTPLS G+ TA +
Sbjct: 1181FFNGKELCKSINPDEAVAYGAAVQAAILSGEGNEKVQDLLLLDVTPLSQGLETA---GGV 1351
Query: 420 MDVVIPRNTSIPVKNTKGYCTAIDN-CGASIIVYEGERPRASDNNLLGLFTLSCL-PGPR 477
M V+IPRNT+IP K + + T DN G I VYEGER R DNNLLG F LS + P PR
Sbjct: 1352MTVLIPRNTTIPTKKEQVFSTYSDNQPGVLIQVYEGERTRTKDNNLLGKFELSGIPPAPR 1531
Query: 478 GQP-LEVCFSIDENGILTVSGKDITTGILNEITITNEKERLSKFEIEKKIEEAEKYRVED 536
G P + VCF ID NGIL VS +D TTG N+ITITN+K RLSK EIEK ++EAEKY+ ED
Sbjct: 1532GVPQITVCFDIDANGILNVSAEDKTTGQKNKITITNDKGRLSKEEIEKMVQEAEKYKSED 1711
Query: 537 MKFLRKAKVISALDSCVYNIKSALNMKDVDLILSPQESEQINNAIIVAMNLLEMNKNNQH 596
+ +K + ++L++ YN+++ + + + LS + +QI +AI A+ L+ NQ
Sbjct: 1712EEHKKKVEAKNSLENYAYNMRNTIKDEKISSKLSGGDKKQIEDAIEGAIQWLDA---NQL 1882
Query: 597 IEIDVLEHHLKEMNSMLNMLL 617
E D E +KE+ ++ N ++
Sbjct: 1883AEADEFEDKMKELETICNPII 1945
>TC85389 homologue to SP|Q03684|BIP4_TOBAC Luminal binding protein 4
precursor (BiP 4) (78 kDa glucose-regulated protein
homolog 4) (GRP 78-4)., partial (94%)
Length = 2265
Score = 561 bits (1446), Expect = e-160
Identities = 317/620 (51%), Positives = 426/620 (68%), Gaps = 6/620 (0%)
Frame = +2
Query: 7 GRAVGIDLGTTYSCVAVWLDDHNRVEIIHNDQGNRTTPSVVAFTDYQRLIGNGAENQSAT 66
G +GIDLGTTYSCV V+ + H VEII NDQGNR TPS V+FTD +RLIG A+N +A
Sbjct: 155 GTVIGIDLGTTYSCVGVYKNGH--VEIIANDQGNRITPSWVSFTDDERLIGEAAKNLAAV 328
Query: 67 NPENTVFDAKRLIGRKYSDPIVQKDIMLWPFMVTSGVNDKPMITVKYK-GQEKQFCAEEI 125
NPE T+FD KRLIGRK++D VQ+D+ L P+ + + + KP I V+ K G+ K F EE+
Sbjct: 329 NPERTIFDVKRLIGRKFADKEVQRDMKLVPYKIVNK-DGKPYIQVRVKDGETKVFSPEEV 505
Query: 126 SSAVLTKMREIAEAYLGSPVKNVVVTVPAYFNDSQRKATIDAGAIAGLNVIRILNEPTAA 185
S+ +LTKM+E AEA+LG +++ VVTVPAYFND+QR+AT DAG IAGLNV RI+NEPTAA
Sbjct: 506 SAMILTKMKETAEAFLGKTIRDAVVTVPAYFNDAQRQATKDAGVIAGLNVARIINEPTAA 685
Query: 186 AIAYGLDKRSNYDGKRNILVFDLGGGTFDVSILTIKGDVFDVKATAGNTHLGGEDFDNRM 245
AIAYGLDK+ G++NILVFDLGGGTFDVSILTI VF+V +T G+THLGGEDFD R+
Sbjct: 686 AIAYGLDKKG---GEKNILVFDLGGGTFDVSILTIDNGVFEVLSTNGDTHLGGEDFDQRI 856
Query: 246 VYYFVEEFKKKNKVEISGNPKSLRRLRTACERAKRILSFTFVTTVEVDALFMGIDFSSSI 305
+ YF++ KKK+ +IS + ++L +LR ERAKR LS VE+++LF G+DFS +
Sbjct: 857 MEYFIKLIKKKHSKDISKDNRALGKLRRESERAKRALSSQHQVRVEIESLFDGVDFSEPL 1036
Query: 306 TRAKFEEINMDFFNECINIVDSCLRDSKIYKNDIDDVVLVGGSSRIPKVQDLLLEFFKGK 365
TRA+FEE+N D F + + V + D+ + KN ID++VLVGGS+RIPKVQ LL ++F GK
Sbjct: 1037TRARFEELNNDLFRKTMGPVKKAMDDAGLQKNQIDEIVLVGGSTRIPKVQQLLKDYFDGK 1216
Query: 366 ALFMNINPDEAIAYGAAVQAAILS-EDFKNVPNLVLRDVTPLSLGIATATYWENIMDVVI 424
+NPDEA+A+GAAVQ +ILS E + +++L DV PL+LGI T +M +I
Sbjct: 1217EPNKGVNPDEAVAFGAAVQGSILSGEGGEETKDILLLDVAPLTLGIETV---GGVMTKLI 1387
Query: 425 PRNTSIPVKNTKGYCTAID-NCGASIIVYEGERPRASDNNLLGLFTLSCL-PGPRGQP-L 481
PRNT IP K ++ + T D SI V+EGER D LLG F LS + P PRG P +
Sbjct: 1388PRNTVIPTKKSQVFTTYQDQQTTVSIQVFEGERSLTKDCRLLGNFDLSGIPPAPRGTPQI 1567
Query: 482 EVCFSIDENGILTVSGKDITTGILNEITITNEKERLSKFEIEKKIEEAEKYRVEDMKFLR 541
EV F +D NGIL V +D TG +ITITNEK RLS+ EI++ + EAE++ ED K
Sbjct: 1568EVTFEVDANGILNVRAEDKGTGKSEKITITNEKGRLSQEEIDRMVREAEEFAEEDKKVKE 1747
Query: 542 KAKVISALDSCVYNIKSALNMKD-VDLILSPQESEQINNAIIVAMNLLEMNKNNQHIEID 600
+ +AL++ VYN+K+ ++ KD + L E E+I A+ A+ L+ +NQ +E +
Sbjct: 1748RIDARNALETYVYNMKNQISDKDKLADKLESDEKEKIEAAVKEALEWLD---DNQTVEKE 1918
Query: 601 VLEHHLKEMNSMLNMLLKIV 620
E LKE+ ++ N ++ V
Sbjct: 1919EFEEKLKEVEAVCNPIITAV 1978
>TC85718 homologue to SP|Q01899|HS7M_PHAVU Heat shock 70 kDa protein
mitochondrial precursor. [Kidney bean French bean]
{Phaseolus vulgaris}, complete
Length = 2371
Score = 435 bits (1118), Expect = e-122
Identities = 267/576 (46%), Positives = 364/576 (62%), Gaps = 15/576 (2%)
Frame = +2
Query: 1 MARKYEGRA-----VGIDLGTTYSCVAVWLDDHNRVEIIHNDQGNRTTPSVVAFTDY-QR 54
++R + RA +GIDLGTT SCV+V + +++ N +G RTTPSVVAFT +
Sbjct: 221 LSRPFSSRAAGNDVIGIDLGTTNSCVSVM--EGKNPKVVENSEGARTTPSVVAFTQKGEL 394
Query: 55 LIGNGAENQSATNPENTVFDAKRLIGRKYSDPIVQKDIMLWPFMVTSGVNDKPMITVKYK 114
L+G A+ Q+ TNPENT+ AKRLIGR++ DP QK++ + P+ + N V+ K
Sbjct: 395 LVGTPAKRQAVTNPENTISGAKRLIGRRFDDPQTQKEMKMVPYKIVKAPNGDAW--VEAK 568
Query: 115 GQEKQFCAEEISSAVLTKMREIAEAYLGSPVKNVVVTVPAYFNDSQRKATIDAGAIAGLN 174
GQ Q+ +I + VLTKM+E AEAYLG V V+TVPAYFND+QR+AT DAG IAGL
Sbjct: 569 GQ--QYSPSQIGAFVLTKMKETAEAYLGKTVPKAVITVPAYFNDAQRQATKDAGRIAGLE 742
Query: 175 VIRILNEPTAAAIAYGLDKRSNYDGKRNILVFDLGGGTFDVSILTIKGDVFDVKATAGNT 234
V+RI+NEPTAAA++YG++K I VFDLGGGTFDVSIL I VF+VKAT G+T
Sbjct: 743 VLRIINEPTAAALSYGMNKEG------LIAVFDLGGGTFDVSILEISNGVFEVKATNGDT 904
Query: 235 HLGGEDFDNRMVYYFVEEFKKKNKVEISGNPKSLRRLRTACERAKRILSFTFVTTVEVDA 294
LGGEDFDN ++ + V EFK+ +++S + +L+RLR A E+AK LS T T E++
Sbjct: 905 FLGGEDFDNALLDFLVNEFKRSESIDLSKDKLALQRLREAAEKAKIELSSTSQT--EINL 1078
Query: 295 LFMGIDFSS------SITRAKFEEINMDFFNECINIVDSCLRDSKIYKNDIDDVVLVGGS 348
F+ D S ++TR+KFE + + SCL+D+ I DID+V+LVGG
Sbjct: 1079PFITADASGAKHLNITLTRSKFEALVNNLIERTKAPCQSCLKDANISTKDIDEVLLVGGM 1258
Query: 349 SRIPKVQDLLLEFFKGKALFMNINPDEAIAYGAAVQAAILSEDFKNVPNLVLRDVTPLSL 408
+R+PKVQ+++ F GK+ +NPDEA+A GAA+Q IL D K L+L DVTPLSL
Sbjct: 1259TRVPKVQEVVSTIF-GKSPCKGVNPDEAVAMGAALQGGILRGDVK---ELLLLDVTPLSL 1426
Query: 409 GIATATYWENIMDVVIPRNTSIPVKNTKGYCTAIDN-CGASIIVYEGERPRASDNNLLGL 467
GI T I +I RNT+IP K ++ + TA DN I V +GER ASDN +LG
Sbjct: 1427GIETL---GGIFTRLINRNTTIPTKKSQVFSTAADNQTQVGIKVLQGEREMASDNKMLGE 1597
Query: 468 FTLSCL-PGPRGQP-LEVCFSIDENGILTVSGKDITTGILNEITITNEKERLSKFEIEKK 525
F L L P PRG P +EV F ID NGI+TVS KD +TG +ITI LS EI+
Sbjct: 1598FELVGLPPAPRGMPQIEVTFDIDANGIVTVSAKDKSTGKEQQITI-KSSGGLSDDEIQNM 1774
Query: 526 IEEAEKYRVEDMKFLRKAKVISALDSCVYNIKSALN 561
++EAE + +D + + ++ D+ +Y+I+ +L+
Sbjct: 1775VKEAELHAQKDQERKSLIDIRNSADTSIYSIEKSLS 1882
>TC80733 homologue to SP|Q01233|HS70_NEUCR Heat shock 70 kDa protein
(HSP70). {Neurospora crassa}, partial (54%)
Length = 1258
Score = 417 bits (1073), Expect(2) = e-120
Identities = 214/335 (63%), Positives = 266/335 (78%)
Frame = +2
Query: 9 AVGIDLGTTYSCVAVWLDDHNRVEIIHNDQGNRTTPSVVAFTDYQRLIGNGAENQSATNP 68
A+GIDLGTTYSCV + +D ++EII NDQGNRTTPS VAF D +RLIG+ A+NQ A NP
Sbjct: 188 AIGIDLGTTYSCVGRYAND--KIEIIANDQGNRTTPSYVAFNDTERLIGDAAKNQVAMNP 361
Query: 69 ENTVFDAKRLIGRKYSDPIVQKDIMLWPFMVTSGVNDKPMITVKYKGQEKQFCAEEISSA 128
NTVFDAKRLIGRK+SD VQ D+ +PF V KP I V++KG+ K F EEIS+
Sbjct: 362 HNTVFDAKRLIGRKFSDSEVQADMKHFPFKVIDK-GGKPNIEVEFKGENKTFTPEEISAM 538
Query: 129 VLTKMREIAEAYLGSPVKNVVVTVPAYFNDSQRKATIDAGAIAGLNVIRILNEPTAAAIA 188
VL KMRE AEAYLG V N V+TVPAYFNDSQR+AT DAG IAGLNV+RI+NEPTAAAIA
Sbjct: 539 VLVKMRETAEAYLGGQVTNAVITVPAYFNDSQRQATKDAGLIAGLNVLRIINEPTAAAIA 718
Query: 189 YGLDKRSNYDGKRNILVFDLGGGTFDVSILTIKGDVFDVKATAGNTHLGGEDFDNRMVYY 248
YGLDK++ +G+RN+L+FDLGGGTFDVS+LTI+ +F+VK+TAG+THLGGEDFDNR+V +
Sbjct: 719 YGLDKKA--EGERNVLIFDLGGGTFDVSLLTIEEGIFEVKSTAGDTHLGGEDFDNRLVNH 892
Query: 249 FVEEFKKKNKVEISGNPKSLRRLRTACERAKRILSFTFVTTVEVDALFMGIDFSSSITRA 308
FV EFK+K+K ++S N ++LRRLRTACERAKR LS + T++E+D+LF GIDF +SITRA
Sbjct: 893 FVNEFKRKHKKDLSSNARALRRLRTACERAKRTLSSSAQTSIEIDSLFEGIDFYTSITRA 1072
Query: 309 KFEEINMDFFNECINIVDSCLRDSKIYKNDIDDVV 343
+FEE+ D F I VD L D+KI K+ + ++V
Sbjct: 1073RFEELCQDLFRSTIQPVDRVLSDAKIDKSQVHEIV 1177
Score = 32.3 bits (72), Expect(2) = e-120
Identities = 12/21 (57%), Positives = 18/21 (85%)
Frame = +3
Query: 345 VGGSSRIPKVQDLLLEFFKGK 365
VGGS+RIP++Q L+ ++F GK
Sbjct: 1182 VGGSTRIPRIQKLISDYFNGK 1244
>TC80669 homologue to PIR|S53498|S53498 dnaK-type molecular chaperone
HSP71.2 - garden pea, partial (44%)
Length = 955
Score = 420 bits (1080), Expect = e-118
Identities = 211/289 (73%), Positives = 248/289 (85%)
Frame = +2
Query: 1 MARKYEGRAVGIDLGTTYSCVAVWLDDHNRVEIIHNDQGNRTTPSVVAFTDYQRLIGNGA 60
MA K EG+A+GIDLGTTYSCV VW +D RVEII NDQGNRTTPS VAFTD +RLIG+ A
Sbjct: 98 MATK-EGKAIGIDLGTTYSCVGVWQND--RVEIIPNDQGNRTTPSYVAFTDTERLIGDAA 268
Query: 61 ENQSATNPENTVFDAKRLIGRKYSDPIVQKDIMLWPFMVTSGVNDKPMITVKYKGQEKQF 120
+NQ A NP+NTVFDAKRLIGR++SD VQ D+ LWPF V G +KPMI V YKG+EK+F
Sbjct: 269 KNQVAMNPQNTVFDAKRLIGRRFSDESVQNDMKLWPFKVVPGPAEKPMIVVNYKGEEKKF 448
Query: 121 CAEEISSAVLTKMREIAEAYLGSPVKNVVVTVPAYFNDSQRKATIDAGAIAGLNVIRILN 180
AEEISS VL KMRE+AEA+LG PVKN VVTVPAYFNDSQR+AT DAGAI+GLNV+RI+N
Sbjct: 449 AAEEISSMVLIKMREVAEAFLGHPVKNAVVTVPAYFNDSQRQATKDAGAISGLNVLRIIN 628
Query: 181 EPTAAAIAYGLDKRSNYDGKRNILVFDLGGGTFDVSILTIKGDVFDVKATAGNTHLGGED 240
EPTAAAIAYGLDK+++ G++N+L+FDLGGGTFDVS+LTI+ +F+VKATAG+THLGGED
Sbjct: 629 EPTAAAIAYGLDKKASRKGEQNVLIFDLGGGTFDVSLLTIEEGIFEVKATAGDTHLGGED 808
Query: 241 FDNRMVYYFVEEFKKKNKVEISGNPKSLRRLRTACERAKRILSFTFVTT 289
FDNRMV +FV EF++KNK +ISGN ++LRRLRTACERAKR LS T TT
Sbjct: 809 FDNRMVNHFVSEFRRKNKKDISGNARALRRLRTACERAKRTLSSTAQTT 955
>TC85716 homologue to PIR|S19140|S19140 dnaK-type molecular chaperone PHSP1
precursor mitochondrial - garden pea, partial (98%)
Length = 2422
Score = 419 bits (1076), Expect = e-117
Identities = 257/578 (44%), Positives = 356/578 (61%), Gaps = 17/578 (2%)
Frame = +1
Query: 1 MARKYEGR-----AVGIDLGTTYSCVAVWLDDHNRVEIIHNDQGNRTTPSVVAFTDY-QR 54
+AR + R +GIDLGTT SCV+ L + ++I N +G RTTPSVVAF +
Sbjct: 205 LARPFSSRPAGSDVIGIDLGTTNSCVS--LMEGKNPKVIENSEGARTTPSVVAFNQKGEL 378
Query: 55 LIGNGAENQSATNPENTVFDAKRLIGRKYSDPIVQKDIMLWPFMVTSGVNDKPMITVKYK 114
L+G A+ Q+ TNP NT+F KRLIGR++ DP QK++ + P+ + N + +
Sbjct: 379 LVGTPAKRQAVTNPTNTLFGTKRLIGRRFDDPQTQKEMKMVPYKIVKAPNGDAWVEIN-- 552
Query: 115 GQEKQFCAEEISSAVLTKMREIAEAYLGSPVKNVVVTVPAYFNDSQRKATIDAGAIAGLN 174
++Q+ +I + VLTKM+E AEAYLG + VVTVPAYFND+QR+AT DAG IAGL
Sbjct: 553 --KQQYSPSQIGAFVLTKMKETAEAYLGKTISKAVVTVPAYFNDAQRQATKDAGRIAGLE 726
Query: 175 VIRILNEPTAAAIAYGLDKRSNYDGKRNILVFDLGGGTFDVSILTIKGDVFDVKATAGNT 234
V RI+NEPTAAA++YG++ + I VFDLGGGTFDVSIL I VF+VKAT G+T
Sbjct: 727 VKRIINEPTAAALSYGMNNKEGL-----IAVFDLGGGTFDVSILEISNGVFEVKATNGDT 891
Query: 235 HLGGEDFDNRMVYYFVEEFKKKNKVEISGNPKSLRRLRTACERAKRILSFTFVTTVEVDA 294
LGGEDFDN ++ + V EFK+ + ++++ + +L+RLR A E+AK LS T T E++
Sbjct: 892 FLGGEDFDNALLDFLVSEFKRTDSIDLAKDKLALQRLREAAEKAKIELSSTSQT--EINL 1065
Query: 295 LFMGIDFSS------SITRAKFEEINMDFFNECINIVDSCLRDSKIYKNDIDDVVLVGGS 348
F+ D S ++TR+KFE + + SCL+D+ I D+D+V+LVGG
Sbjct: 1066PFITADASGAKHLNITLTRSKFEALVNNLIERTKAPCKSCLKDANISIKDVDEVLLVGGM 1245
Query: 349 SRIPKVQDLLLEFFKGKALFMNINPDEAIAYGAAVQAAILSEDFKNVPNLVLRDVTPLSL 408
+R+PKVQ+++ E F GK+ +NPDEA+A GAA+Q IL D K L+L DVTPLSL
Sbjct: 1246TRVPKVQEVVSEIF-GKSPSKGVNPDEAVAMGAALQGGILRGDVK---ELLLLDVTPLSL 1413
Query: 409 GIATATYWENIMDVVIPRNTSIPVKNTKGYCTAIDN---CGASIIVYEGERPRASDNNLL 465
GI T I +I RNT+IP K ++ + TA DN G EG R A+DN L
Sbjct: 1414GIETL---GGIFTRLISRNTTIPTKKSQVFSTAADNQTQRGYXRCSQEGXREMAADNKSL 1584
Query: 466 GLFTLSCL-PGPRGQP-LEVCFSIDENGILTVSGKDITTGILNEITITNEKERLSKFEIE 523
G F L + P PRG P +EV F ID NGI+TVS KD +TG +ITI LS EI
Sbjct: 1585GEFDLVGIPPAPRGLPQIEVTFDIDANGIVTVSAKDKSTGKEQQITI-RSSGGLSDDEIN 1761
Query: 524 KKIEEAEKYRVEDMKFLRKAKVISALDSCVYNIKSALN 561
++EAE + D + + ++ D+ +Y+I+ +L+
Sbjct: 1762NMVKEAELHAQRDQERKALIDIKNSADTSIYSIEKSLS 1875
>TC87268 homologue to PIR|S53500|S44168 dnaK-type molecular chaperone
HSC71.0 - garden pea, partial (59%)
Length = 1336
Score = 333 bits (853), Expect = 2e-91
Identities = 188/356 (52%), Positives = 247/356 (68%), Gaps = 4/356 (1%)
Frame = +2
Query: 266 KSLRRLRTACERAKRILSFTFVTTVEVDALFMGIDFSSSITRAKFEEINMDFFNECINIV 325
++LRRLRTACERAKR LS T TT+E+D+LF GIDF S ITRA+FEE+NMD F +C+ V
Sbjct: 2 RALRRLRTACERAKRTLSSTAQTTIEIDSLFEGIDFYSPITRARFEELNMDLFRKCMEPV 181
Query: 326 DSCLRDSKIYKNDIDDVVLVGGSSRIPKVQDLLLEFFKGKALFMNINPDEAIAYGAAVQA 385
+ CLRD+K+ K + DVVLVGGS+RIPKVQ LL +FF GK L +INPDEA+AYGAAVQA
Sbjct: 182 EKCLRDAKMDKKSVHDVVLVGGSTRIPKVQQLLQDFFNGKELCKSINPDEAVAYGAAVQA 361
Query: 386 AILS-EDFKNVPNLVLRDVTPLSLGIATATYWENIMDVVIPRNTSIPVKNTKGYCTAIDN 444
AILS E + V +L+L DVTPLSLG+ TA +M V+IPRNT+IP K + + T DN
Sbjct: 362 AILSGEGNEKVQDLLLLDVTPLSLGLETA---GGVMTVLIPRNTTIPTKKEQVFSTYSDN 532
Query: 445 -CGASIIVYEGERPRASDNNLLGLFTLSCL-PGPRGQP-LEVCFSIDENGILTVSGKDIT 501
G I V+EGER R DNNLLG F LS + P PRG P + VCF ID NGIL VS +D T
Sbjct: 533 QPGVLIQVFEGERTRTRDNNLLGKFELSGIPPAPRGVPQITVCFDIDANGILNVSAEDKT 712
Query: 502 TGILNEITITNEKERLSKFEIEKKIEEAEKYRVEDMKFLRKAKVISALDSCVYNIKSALN 561
TG N+ITITN+K RLSK +IEK ++EAEKY+ ED + +K + +AL++ YN+++ +
Sbjct: 713 TGQKNKITITNDKGRLSKEDIEKMVQEAEKYKSEDEEHKKKVEAKNALENYAYNMRNTIK 892
Query: 562 MKDVDLILSPQESEQINNAIIVAMNLLEMNKNNQHIEIDVLEHHLKEMNSMLNMLL 617
+ + L + ++I + I A+ L+ NQ E D E +KE+ + N ++
Sbjct: 893 DEKIAGKLDSDDKKKIEDTIEAAIQWLDA---NQLAEADEFEDKMKELEGVCNPII 1051
>BQ139103 homologue to GP|2655420|gb| heat shock cognate protein HSC70
{Brassica napus}, partial (29%)
Length = 628
Score = 283 bits (725), Expect = 1e-76
Identities = 144/193 (74%), Positives = 162/193 (83%)
Frame = +2
Query: 1 MARKYEGRAVGIDLGTTYSCVAVWLDDHNRVEIIHNDQGNRTTPSVVAFTDYQRLIGNGA 60
MA K EG A+GIDLGTTYSCV VW H+RVEII NDQGNRTTPS VAFTD +RLIG+ A
Sbjct: 56 MAGKGEGPAIGIDLGTTYSCVGVW--QHDRVEIIANDQGNRTTPSYVAFTDSERLIGDAA 229
Query: 61 ENQSATNPENTVFDAKRLIGRKYSDPIVQKDIMLWPFMVTSGVNDKPMITVKYKGQEKQF 120
+NQ A NP NTVFDAKRLIGR++SD VQ D+ LWPF + SG +KP+I V YKG++K+F
Sbjct: 230 KNQVAMNPINTVFDAKRLIGRRFSDASVQSDMKLWPFKIISGPAEKPLIGVNYKGEDKEF 409
Query: 121 CAEEISSAVLTKMREIAEAYLGSPVKNVVVTVPAYFNDSQRKATIDAGAIAGLNVIRILN 180
AEEISS VL KMREIAEAYLGS +KN VVTVPAYFNDSQR+AT DAG IAGLNV+RI+N
Sbjct: 410 AAEEISSMVLMKMREIAEAYLGSAIKNAVVTVPAYFNDSQRQATKDAGVIAGLNVMRIIN 589
Query: 181 EPTAAAIAYGLDK 193
EPTAAAIAYGLDK
Sbjct: 590 EPTAAAIAYGLDK 628
>TC80756 homologue to PIR|S53498|S53498 dnaK-type molecular chaperone
HSP71.2 - garden pea, partial (53%)
Length = 1189
Score = 283 bits (724), Expect = 2e-76
Identities = 159/317 (50%), Positives = 217/317 (68%), Gaps = 4/317 (1%)
Frame = +3
Query: 305 ITRAKFEEINMDFFNECINIVDSCLRDSKIYKNDIDDVVLVGGSSRIPKVQDLLLEFFKG 364
ITRA+FEE+NMD F +C+ V+ CLRD+KI K+ + +VVLVGGS+RIPKVQ LL +FF G
Sbjct: 3 ITRARFEELNMDLFRKCMEPVEKCLRDAKIDKSHVHEVVLVGGSTRIPKVQQLLQDFFNG 182
Query: 365 KALFMNINPDEAIAYGAAVQAAILS-EDFKNVPNLVLRDVTPLSLGIATATYWENIMDVV 423
K L +INPDEA+AYGAAVQAAILS E + V +L+L DVTPLSLG+ TA +M +
Sbjct: 183 KELCKSINPDEAVAYGAAVQAAILSGEGDEKVQDLLLLDVTPLSLGLETA---GGVMTTL 353
Query: 424 IPRNTSIPVKNTKGYCTAIDN-CGASIIVYEGERPRASDNNLLGLFTLSCL-PGPRGQP- 480
IPRNT+IP K + + T DN G I V+EGER R DNNLLG F L+ + P PRG P
Sbjct: 354 IPRNTTIPTKKEQIFSTYSDNQPGVLIQVFEGERARTKDNNLLGKFELTGIPPAPRGVPQ 533
Query: 481 LEVCFSIDENGILTVSGKDITTGILNEITITNEKERLSKFEIEKKIEEAEKYRVEDMKFL 540
+ VCF ID NGIL VS +D T G+ N+ITITN+K RLSK EIEK +++AEKY+ ED +
Sbjct: 534 INVCFDIDANGILNVSAEDKTAGVKNKITITNDKGRLSKEEIEKMVKDAEKYKAEDEEVK 713
Query: 541 RKAKVISALDSCVYNIKSALNMKDVDLILSPQESEQINNAIIVAMNLLEMNKNNQHIEID 600
+K + +++++ YN+++ + + + LS ++ E+I A+ A+ LE NQ E+D
Sbjct: 714 KKVEAKNSIENYAYNMRNTIKDEKIGGKLSHEDKEKIEKAVEDAIQWLE---GNQMAEVD 884
Query: 601 VLEHHLKEMNSMLNMLL 617
E KE+ + N ++
Sbjct: 885 EFEDKQKELEGICNPII 935
>TC85394 homologue to GP|2642238|gb|AAB86942.1| endoplasmic reticulum
HSC70-cognate binding protein precursor {Glycine max},
partial (38%)
Length = 962
Score = 271 bits (694), Expect = 5e-73
Identities = 144/230 (62%), Positives = 176/230 (75%), Gaps = 1/230 (0%)
Frame = +1
Query: 7 GRAVGIDLGTTYSCVAVWLDDHNRVEIIHNDQGNRTTPSVVAFTDYQRLIGNGAENQSAT 66
G +GIDLGTTYSCV V+ + H VEII NDQGNR TPS V+FTD +RLIG A+N +A
Sbjct: 289 GTVIGIDLGTTYSCVGVYKNGH--VEIIANDQGNRITPSWVSFTDDERLIGEAAKNLAAV 462
Query: 67 NPENTVFDAKRLIGRKYSDPIVQKDIMLWPFMVTSGVNDKPMITVKYK-GQEKQFCAEEI 125
NPE T+FD KRLIGRK+ D VQ+D+ L P+ + + + KP I V+ K G+ K F EEI
Sbjct: 463 NPERTIFDVKRLIGRKFEDKEVQRDMKLVPYKIVNR-DGKPYIQVRVKDGETKVFSPEEI 639
Query: 126 SSAVLTKMREIAEAYLGSPVKNVVVTVPAYFNDSQRKATIDAGAIAGLNVIRILNEPTAA 185
S+ +L KM+E AE +LG +++ VVTVPAYFND+QR+AT DAG IAGLNV RI+NEPTAA
Sbjct: 640 SAMILGKMKETAEGFLGKTIRDAVVTVPAYFNDAQRQATKDAGVIAGLNVARIINEPTAA 819
Query: 186 AIAYGLDKRSNYDGKRNILVFDLGGGTFDVSILTIKGDVFDVKATAGNTH 235
AIAYGLDK+ G++NILVFDLGGGTFDVSILTI VF+V AT G+TH
Sbjct: 820 AIAYGLDKKG---GEKNILVFDLGGGTFDVSILTIDNGVFEVLATNGDTH 960
>AW559598 weakly similar to SP|P09189|HS7C Heat shock cognate 70 kDa protein.
[Petunia] {Petunia hybrida}, partial (24%)
Length = 532
Score = 261 bits (666), Expect = 9e-70
Identities = 127/167 (76%), Positives = 145/167 (86%)
Frame = +1
Query: 1 MARKYEGRAVGIDLGTTYSCVAVWLDDHNRVEIIHNDQGNRTTPSVVAFTDYQRLIGNGA 60
MA E AVGIDLGTTYSCVAVWL++ NRVEIIHND+G++ TPS VAFTD QRL+G A
Sbjct: 31 MANNCECYAVGIDLGTTYSCVAVWLEEKNRVEIIHNDEGSKITPSFVAFTDGQRLVGAAA 210
Query: 61 ENQSATNPENTVFDAKRLIGRKYSDPIVQKDIMLWPFMVTSGVNDKPMITVKYKGQEKQF 120
++Q+A NP+NTVFDAKRLIGRK+SDP+VQKDI+LWPF V SGVNDKPMI++K+KGQEK
Sbjct: 211 KDQAAINPQNTVFDAKRLIGRKFSDPVVQKDILLWPFKVVSGVNDKPMISLKFKGQEKLL 390
Query: 121 CAEEISSAVLTKMREIAEAYLGSPVKNVVVTVPAYFNDSQRKATIDA 167
CAEEISS VLT MREIAE YL SPVKN +TVPAYFND+QRKATIDA
Sbjct: 391 CAEEISSIVLTNMREIAEMYLESPVKNAGITVPAYFNDAQRKATIDA 531
>TC80052 similar to PIR|JC4786|JC4786 dnaK-type molecular chaperone hsc70-3
- tomato, partial (27%)
Length = 667
Score = 252 bits (644), Expect = 3e-67
Identities = 127/186 (68%), Positives = 151/186 (80%)
Frame = +3
Query: 9 AVGIDLGTTYSCVAVWLDDHNRVEIIHNDQGNRTTPSVVAFTDYQRLIGNGAENQSATNP 68
A+GIDLGTTYSCVAV + ++II ND G RTTPS VAF D +R+IG+ A N +A+NP
Sbjct: 105 AIGIDLGTTYSCVAVC--KNGEIDIIVNDLGKRTTPSFVAFKDSERMIGDAAFNIAASNP 278
Query: 69 ENTVFDAKRLIGRKYSDPIVQKDIMLWPFMVTSGVNDKPMITVKYKGQEKQFCAEEISSA 128
NT+FDAKRLIGRK+SDPIVQ D+ LWPF V +NDKPMI V Y +EK F AEEISS
Sbjct: 279 TNTIFDAKRLIGRKFSDPIVQSDVKLWPFKVIGDLNDKPMIVVNYNDEEKHFAAEEISSM 458
Query: 129 VLTKMREIAEAYLGSPVKNVVVTVPAYFNDSQRKATIDAGAIAGLNVIRILNEPTAAAIA 188
VL KMREIAE +LGS V++VV+TVPAYFNDSQR++T DAGAIAGLNV+RI+NEPTAAAIA
Sbjct: 459 VLVKMREIAETFLGSIVEDVVITVPAYFNDSQRQSTRDAGAIAGLNVMRIINEPTAAAIA 638
Query: 189 YGLDKR 194
YG + +
Sbjct: 639 YGFNTK 656
>TC77585 similar to GP|17473863|gb|AAL38353.1 putative heat-shock protein
{Arabidopsis thaliana}, partial (92%)
Length = 3027
Score = 239 bits (609), Expect = 3e-63
Identities = 137/432 (31%), Positives = 231/432 (52%), Gaps = 10/432 (2%)
Frame = +1
Query: 10 VGIDLGTTYSCVAVWLDDHNRVEIIHNDQGNRTTPSVVAFTDYQRLIGNGAENQSATNPE 69
VG D G VAV ++++ ND+ R TP++V F D QR IG + NP+
Sbjct: 247 VGFDFGNESCIVAV--ARQRGIDVVLNDESKRETPAIVCFGDKQRFIGTAGAASTMMNPK 420
Query: 70 NTVFDAKRLIGRKYSDPIVQKDIMLWPFMVTSGVNDKPMITVKYKGQEKQFCAEEISSAV 129
N++ KRLIG+K++DP +Q+D+ PF VT G + P+I +Y G+ ++F A ++ +
Sbjct: 421 NSISQIKRLIGKKFADPELQRDLKSLPFNVTEGPDGYPLIHARYLGESREFTATQVFGMM 600
Query: 130 LTKMREIAEAYLGSPVKNVVVTVPAYFNDSQRKATIDAGAIAGLNVIRILNEPTAAAIAY 189
L+ ++EIA+ L + V + + +P YF D QR++ +DA IAGL+ + +++E TA A+AY
Sbjct: 601 LSNLKEIAQKNLNAAVVDCCIGIPVYFTDLQRRSVLDAATIAGLHPLHLIHETTATALAY 780
Query: 190 GLDKRSNYDGK-RNILVFDLGGGTFDVSILTIKGDVFDVKATAGNTHLGGEDFDNRMVYY 248
G+ K + + N+ D+G + V I K V + + + LGG DFD + ++
Sbjct: 781 GIYKTDLPENEWLNVAFVDVGHASMQVCIAGFKKGQLHVLSHSYDRSLGGRDFDEALFHH 960
Query: 249 FVEEFKKKNKVEISGNPKSLRRLRTACERAKRILSFTFVTTVEVDALFMGIDFSSSITRA 308
F +FK++ K+++ N ++ RLR ACE+ K++LS + ++ L D I R
Sbjct: 961 FAAKFKEEYKIDVYQNARACLRLRAACEKLKKVLSANPEAPLNIECLMDEKDVRGFIKRD 1140
Query: 309 KFEEINMDFFNECINIVDSCLRDSKIYKNDIDDVVLVGGSSRIPKVQDLLLEFFKGKALF 368
FE++++ ++ L ++ + +I V +VG SR+P + +L EFFK K
Sbjct: 1141DFEQLSLPILERVKGPLEKALAEAGLTVENIHMVEVVGSGSRVPAINKILTEFFK-KEPR 1317
Query: 369 MNINPDEAIAYGAAVQAAILSEDFKNVPNLVLRDVTPLSLGIATATYWENIMD------- 421
+N E +A GAA+Q AILS FK V + + P S+ ++ + D
Sbjct: 1318RTMNASECVARGAALQCAILSPTFK-VREFQVNESFPFSVSLSWKYSGSDAPDSESDNKQ 1494
Query: 422 --VVIPRNTSIP 431
+V P+ IP
Sbjct: 1495STIVFPKGNPIP 1530
>TC85391 homologue to PIR|T06598|T06598 dnaK-type molecular chaperone BiP-A
- soybean, partial (52%)
Length = 1231
Score = 236 bits (602), Expect = 2e-62
Identities = 144/328 (43%), Positives = 203/328 (60%), Gaps = 5/328 (1%)
Frame = +1
Query: 298 GIDFSSSITRAKFEEINMDFFNECINIVDSCLRDSKIYKNDIDDVVLVGGSSRIPKVQDL 357
G+ FS +TRA+FEE+N D F + + V + D+ + KN ID++VLVGGS+RIPKVQ L
Sbjct: 4 GVAFSEPLTRARFEELNNDLFRKTMGPVKKAMDDAGLQKNQIDEIVLVGGSTRIPKVQQL 183
Query: 358 LLEFFKGKALFMNINPDEAIAYGAAVQAAILS-EDFKNVPNLVLRDVTPLSLGIATATYW 416
L ++F GK +NPDEA+A+GAAVQ +ILS E +++L DV PL+LGI T
Sbjct: 184 LKDYFDGKEPNKGVNPDEAVAFGAAVQGSILSGEGGAETKDILLLDVAPLTLGIETV--- 354
Query: 417 ENIMDVVIPRNTSIPVKNTKGYCTAID-NCGASIIVYEGERPRASDNNLLGLFTLSCL-P 474
+M +IPRNT IP K ++ + T D SI V+EGER D LLG F LS + P
Sbjct: 355 GGVMTKLIPRNTVIPTKKSQVFTTYQDQQTTVSIQVFEGERSLTKDCRLLGKFDLSGIPP 534
Query: 475 GPRGQP-LEVCFSIDENGILTVSGKDITTGILNEITITNEKERLSKFEIEKKIEEAEKYR 533
PRG P +EV F +D NGIL V +D TG +ITITNEK RLS+ EIE+ + EAE++
Sbjct: 535 APRGTPQIEVTFEVDANGILNVKAEDKGTGKSEKITITNEKGRLSQEEIERMVREAEEFA 714
Query: 534 VEDMKFLRKAKVISALDSCVYNIKSALNMKD-VDLILSPQESEQINNAIIVAMNLLEMNK 592
ED K + +AL++ VYN+K+ ++ KD + L E E+I A+ A+ L+
Sbjct: 715 EEDKKVKERIDARNALETYVYNMKNQVSDKDKLADKLESDEKEKIETAVKEALEWLD--- 885
Query: 593 NNQHIEIDVLEHHLKEMNSMLNMLLKIV 620
+NQ +E + E LKE+ ++ N ++ V
Sbjct: 886 DNQSVEKEEYEEKLKEVEAVCNPIITAV 969
>TC80997 similar to PIR|B84729|B84729 70kD heat shock protein [imported] -
Arabidopsis thaliana, partial (91%)
Length = 1717
Score = 134 bits (338), Expect(2) = 7e-61
Identities = 88/236 (37%), Positives = 133/236 (56%), Gaps = 6/236 (2%)
Frame = +1
Query: 266 KSLRRLRTACERAKRILSFTFVTTVEVDALFMGIDFSSSITRAKFEEINMDFFNECINIV 325
KS+ LR A + A LS V+VD L + + RA+FEE+N + F +C +++
Sbjct: 772 KSMALLRVATQDAIHQLSTQSSVQVDVD-LGNSLKICKVVDRAEFEEVNKEVFEKCESLI 948
Query: 326 DSCLRDSKIYKNDIDDVVLVGGSSRIPKVQDLLLEFFKGKALFMNINPDEAIAYGAAVQA 385
CL D+K+ DI+DV+LVGG S IP+V++L+ K ++ INP E GA ++
Sbjct: 949 IQCLHDAKVEVGDINDVILVGGCSYIPRVENLVTNLCKITEVYKGINPLEGAVCGATMEG 1128
Query: 386 AI---LSEDFKNVPNLVLRDVTPLSLGIATATYWENIMDVVIPRNTSIPVKNTKGYCTAI 442
A+ +S+ F N+ L ++ TPL++GI N VIPRNT++P + + T
Sbjct: 1129 AVASGISDPFGNLDLLTIQ-ATPLAIGIRAD---GNKFVPVIPRNTTMPARKDLLFTTIH 1296
Query: 443 DN-CGASIIVYEGERPRASDNNLLGLFTLSCLP-GPRGQP-LEVCFSIDENGILTV 495
DN A I+VYEGE +A +N+LLG F + +P P+G P + VC ID +L V
Sbjct: 1297 DNQTEALILVYEGEGKKAEENHLLGYFKIMGIPAAPKGVPEISVCMDIDAANVLRV 1464
Score = 118 bits (296), Expect(2) = 7e-61
Identities = 81/249 (32%), Positives = 131/249 (52%), Gaps = 11/249 (4%)
Frame = +3
Query: 15 GTTYSCVAVWLDDHNRVEIIHNDQGNRTTPSVVAFTDYQRLIGNGAENQSATNPE----N 70
GT+ VAVW + + VE++ N + + S V F D G +Q + E +
Sbjct: 3 GTSQCSVAVW--NGSEVELLKNKRNQKLMKSFVTFKD--EAPSGGVTSQFSHEHEMLFGD 170
Query: 71 TVFDAKRLIGRKYSDPIVQKDIMLWPFMV-TSGVNDKPMITVKYKGQEKQFCAEEISSAV 129
T+F+ KRLIGR +DP+V L PF+V T + +P I + EE+ +
Sbjct: 171 TIFNMKRLIGRVDTDPVVHASKNL-PFLVQTLDIGVRPFIAALVNNVWRSTTPEEVLAMF 347
Query: 130 LTKMREIAEAYLGSPVKNVVVTVPAYFNDSQRKATIDAGAIAGLNVIRILNEPTAAAIAY 189
L ++R + E +L P++NVV+TVP F+ Q A A+AGL+V+R++ EPTA A+ Y
Sbjct: 348 LVELRLMTETHLKRPIRNVVLTVPVSFSRFQLTRLERACAMAGLHVLRLMPEPTAVALLY 527
Query: 190 GLDKRS------NYDGKRNILVFDLGGGTFDVSILTIKGDVFDVKATAGNTHLGGEDFDN 243
G ++ ++ L+F++G G DV++ G V +KA AG+T +GGED
Sbjct: 528 GQQQQKASQETMGSGSEKIALIFNMGAGYCDVAVTATAGGVSQIKALAGST-IGGEDLLQ 704
Query: 244 RMVYYFVEE 252
M+ + + +
Sbjct: 705 NMMRHLLPD 731
>BQ752418 homologue to SP|P78695|GR78 78 kDa glucose-regulated protein
homolog precursor (GRP 78), partial (47%)
Length = 1032
Score = 168 bits (426), Expect(3) = 3e-48
Identities = 105/245 (42%), Positives = 149/245 (59%), Gaps = 4/245 (1%)
Frame = -1
Query: 351 IPKVQDLLLEFFKGKALFMNINPDEAIAYGAAVQAAILSEDFKNVPNLVLRDVTPLSLGI 410
IPKVQ L+ E+F GK INPDEA+A+GAAVQA +LS + + +VL DV PL+LGI
Sbjct: 900 IPKVQSLIEEYFGGKKASKGINPDEAVAFGAAVQAGVLSGE-EGTEEIVLMDVNPLTLGI 724
Query: 411 ATATYWENIMDVVIPRNTSIPVKNTKGYCTAIDNCGASII-VYEGERPRASDNNLLGLFT 469
T +M +I RNT IP + ++ + TA DN +I V+EGER DNN LG F
Sbjct: 723 ETTG---GVMTKLITRNTPIPTRKSQIFSTAADNQPVVLIQVFEGERSLTKDNNQLGKFE 553
Query: 470 LSCLP-GPRGQP-LEVCFSIDENGILTVSGKDITTGILNEITITNEKERLSKFEIEKKIE 527
L+ +P PRG P +EV F +D NGIL VS D TG ITITN+K RL+K EI++ +E
Sbjct: 552 LTGIPPAPRGVPQIEVSFELDANGILKVSAHDKGTGKQESITITNDKGRLTKEEIDRMVE 373
Query: 528 EAEKYRVEDMKFLRKAKVISALDSCVYNIKSALNMKD-VDLILSPQESEQINNAIIVAMN 586
EAEKY ED + + + L++ +++K+ +N ++ + + ++ E I A+ +
Sbjct: 372 EAEKYAEEDKATRERIEARNGLENYAFSLKNQVNDEEGLGGKIDDEDKETILEAVKETND 193
Query: 587 LLEMN 591
LE N
Sbjct: 192 WLEEN 178
Score = 40.8 bits (94), Expect(3) = 3e-48
Identities = 16/29 (55%), Positives = 24/29 (82%)
Frame = -3
Query: 325 VDSCLRDSKIYKNDIDDVVLVGGSSRIPK 353
V+ L+D+K+ K D+DD+VLVGGS+R P+
Sbjct: 979 VEQVLKDAKVKKEDVDDIVLVGGSTRYPQ 893
Score = 22.3 bits (46), Expect(3) = 3e-48
Identities = 8/15 (53%), Positives = 11/15 (73%)
Frame = -2
Query: 308 AKFEEINMDFFNECI 322
AKFEE+NMD + +
Sbjct: 1031 AKFEELNMDLLKKTL 987
>TC85395 homologue to GP|6911553|emb|CAB72130.1 heat shock protein 70
{Cucumis sativus}, partial (35%)
Length = 723
Score = 182 bits (463), Expect = 3e-46
Identities = 107/224 (47%), Positives = 148/224 (65%), Gaps = 3/224 (1%)
Frame = +3
Query: 397 NLVLRDVTPLSLGIATATYWENIMDVVIPRNTSIPVKNTKGYCTAIDNC-GASIIVYEGE 455
+L+L DVTPLSLG+ TA +M V+IPRNT+IP K + + T DN G I VYEGE
Sbjct: 6 DLLLLDVTPLSLGLETAG---GVMTVLIPRNTTIPTKKEQVFSTYSDNQPGVLIQVYEGE 176
Query: 456 RPRASDNNLLGLFTLSCLP-GPRGQP-LEVCFSIDENGILTVSGKDITTGILNEITITNE 513
R R DNNLLG F LS +P PRG P + VCF ID NGIL VS +D TTG N+ITITN+
Sbjct: 177 RARTRDNNLLGKFELSGIPPAPRGVPQITVCFDIDANGILNVSAEDKTTGQKNKITITND 356
Query: 514 KERLSKFEIEKKIEEAEKYRVEDMKFLRKAKVISALDSCVYNIKSALNMKDVDLILSPQE 573
K RLSK EIEK +EEAEKY+ ED + +K +AL++ YN+++ + + + LSP++
Sbjct: 357 KGRLSKEEIEKMVEEAEKYKAEDEEHKKKVDAKNALENYAYNMRNTIKDEKIGSKLSPED 536
Query: 574 SEQINNAIIVAMNLLEMNKNNQHIEIDVLEHHLKEMNSMLNMLL 617
++I++AI A+ L+ +NQ E D + +KE+ S+ N ++
Sbjct: 537 KKKIDDAIDAAIQWLD---SNQLAEADEFQDKMKELESLCNPII 659
>AW695262 homologue to GP|170386|gb|A glucose-regulated protein 78
{Lycopersicon esculentum}, partial (39%)
Length = 607
Score = 165 bits (418), Expect = 5e-41
Identities = 88/158 (55%), Positives = 116/158 (72%), Gaps = 2/158 (1%)
Frame = +2
Query: 7 GRAVGIDLGTTYSCVAVWLDDHNRVEIIHNDQGNRTTPSVVAFTDYQRLIGNGAENQSAT 66
G +GIDLGTTYSCV V+ + H VEII NDQGNR TPS V+FTD +RLIG A+N +A
Sbjct: 128 GTVIGIDLGTTYSCVGVYKNGH--VEIIANDQGNRITPSWVSFTDDERLIGEAAKNLAAV 301
Query: 67 NPENTVFDAKRLIGRKYSDPIVQKDIMLWPFMVTSGVNDKPMITVKYK-GQEKQFCAEEI 125
NPE T+FD KRLIGRK++D VQ+D+ L P+ + + + KP I V+ K G+ K F EE+
Sbjct: 302 NPERTIFDVKRLIGRKFADKEVQRDMKLVPYKIVN-KDGKPYIQVRVKDGETKVFSPEEV 478
Query: 126 SSAVLTKMREIAEAYLGSPVKNVVVTVPAYF-NDSQRK 162
S+ +LTKM+E AEA+LG +++ VVTVPA ND+QR+
Sbjct: 479 SAMILTKMKETAEAFLGKTIRDAVVTVPALLSNDAQRQ 592
Database: MTGI
Posted date: Oct 22, 2004 3:39 PM
Number of letters in database: 27,044,181
Number of sequences in database: 36,976
Lambda K H
0.319 0.136 0.382
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 18,352,386
Number of Sequences: 36976
Number of extensions: 222475
Number of successful extensions: 1223
Number of sequences better than 10.0: 67
Number of HSP's better than 10.0 without gapping: 1111
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 1146
length of query: 713
length of database: 9,014,727
effective HSP length: 103
effective length of query: 610
effective length of database: 5,206,199
effective search space: 3175781390
effective search space used: 3175781390
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.7 bits)
S2: 62 (28.5 bits)
Medicago: description of AC137996.5


