

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC137996.18 + phase: 0
(647 letters)
Database: MTGI
36,976 sequences; 27,044,181 total letters
Searching..................................................done
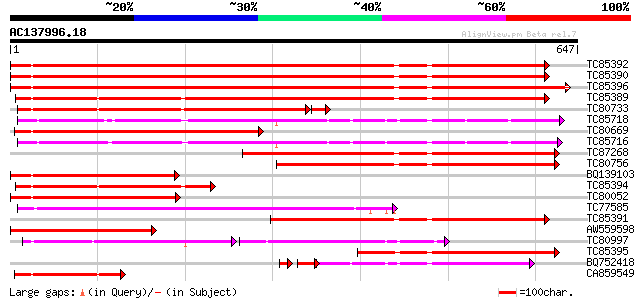
Score E
Sequences producing significant alignments: (bits) Value
TC85392 homologue to PIR|JC4786|JC4786 dnaK-type molecular chape... 745 0.0
TC85390 homologue to SP|P27322|HS72_LYCES Heat shock cognate 70 ... 745 0.0
TC85396 homologue to GP|6911551|emb|CAB72129.1 heat shock protei... 738 0.0
TC85389 homologue to SP|Q03684|BIP4_TOBAC Luminal binding protei... 573 e-164
TC80733 homologue to SP|Q01233|HS70_NEUCR Heat shock 70 kDa prot... 428 e-123
TC85718 homologue to SP|Q01899|HS7M_PHAVU Heat shock 70 kDa prot... 429 e-120
TC80669 homologue to PIR|S53498|S53498 dnaK-type molecular chape... 427 e-120
TC85716 homologue to PIR|S19140|S19140 dnaK-type molecular chape... 415 e-116
TC87268 homologue to PIR|S53500|S44168 dnaK-type molecular chape... 350 1e-96
TC80756 homologue to PIR|S53498|S53498 dnaK-type molecular chape... 298 4e-81
BQ139103 homologue to GP|2655420|gb| heat shock cognate protein ... 289 3e-78
TC85394 homologue to GP|2642238|gb|AAB86942.1| endoplasmic retic... 279 3e-75
TC80052 similar to PIR|JC4786|JC4786 dnaK-type molecular chapero... 256 2e-68
TC77585 similar to GP|17473863|gb|AAL38353.1 putative heat-shock... 248 5e-66
TC85391 homologue to PIR|T06598|T06598 dnaK-type molecular chape... 243 2e-64
AW559598 weakly similar to SP|P09189|HS7C Heat shock cognate 70 ... 236 3e-62
TC80997 similar to PIR|B84729|B84729 70kD heat shock protein [im... 125 4e-54
TC85395 homologue to GP|6911553|emb|CAB72130.1 heat shock protei... 181 8e-46
BQ752418 homologue to SP|P78695|GR78 78 kDa glucose-regulated pr... 162 2e-45
CA859549 similar to GP|20260807|gb Hsp70 protein 1 {Rhizopus sto... 172 3e-43
>TC85392 homologue to PIR|JC4786|JC4786 dnaK-type molecular chaperone
hsc70-3 - tomato, complete
Length = 2295
Score = 745 bits (1924), Expect = 0.0
Identities = 389/619 (62%), Positives = 480/619 (76%), Gaps = 3/619 (0%)
Frame = +1
Query: 1 MEEKYEGVAIGIDLGTTYSCVGVWHEQNDRVEIIHNDQGNRTTPSCVAFTNSQRLIGDAA 60
M K EG AIGIDLGTTYSCVGVW Q+DRVEII NDQGNRTTPS VAFT+++RLIGDAA
Sbjct: 130 MAGKGEGPAIGIDLGTTYSCVGVW--QHDRVEIIANDQGNRTTPSYVAFTDTERLIGDAA 303
Query: 61 KNQAATNPSNTIFDTKRLIGRKYSDSIVQKDCLLWPFKVIAGTNDKPTIVVNYKGEEKRF 120
KNQ A NP+NT+FD KRLIGR+ SD VQ D LWPFKVI G DKP IVVNYK EEK+F
Sbjct: 304 KNQVAMNPTNTVFDAKRLIGRRISDVSVQSDMKLWPFKVIPGPADKPMIVVNYKAEEKQF 483
Query: 121 VAEEISSVILSKMREIAEAFLQTPVKNAVISVPAYFNDSQRRATKDAGAIAGLNVMQIIN 180
AEEISS++L KM+EIAEA+L T +KNAV++VPAYFNDSQR+ATKDAG I+GLNVM+IIN
Sbjct: 484 SAEEISSMVLMKMKEIAEAYLGTTIKNAVVTVPAYFNDSQRQATKDAGVISGLNVMRIIN 663
Query: 181 EPTAAALAYGLQKRANCVDKRNIFIFDLGGGTFDVSILTVKNNVFEVKATAGDTHLGGQD 240
EPTAAA+AYGL K+A ++N+ IFDLGGGTFDVS+LT++ +FEVKATAGDTHLGG+D
Sbjct: 664 EPTAAAIAYGLDKKATSTGEKNVLIFDLGGGTFDVSLLTIEEGIFEVKATAGDTHLGGED 843
Query: 241 FDNIMVDHFVKEFRNKYKEDISENSKALRRLRTACEKAKRTLSYDTDATIELDAIYKGID 300
FDN MV+HFV+EF+ K K+DIS N +ALRR+RTACE+AKRTLS TIE+D++++GID
Sbjct: 844 FDNRMVNHFVQEFKRKNKKDISGNPRALRRVRTACERAKRTLSSTAQTTIEIDSLFEGID 1023
Query: 301 FCSSITRAKFEEMNMNLFEKCMNTVNICLADAKMDKNSVDDVVLVGGSSRIPKVRQLLQE 360
F ++ITRA+FEE+NM+LF KCM V CL DAKMDKNSVDDVVLVGGS+RIPKV+QLLQ+
Sbjct: 1024FYTTITRARFEELNMDLFRKCMEPVEKCLRDAKMDKNSVDDVVLVGGSTRIPKVQQLLQD 1203
Query: 361 VFKGKELCKSINPDEAVAYGAVVQAALLS-EGSKNVPNLVLRDVTPLSFGVSELGNLMSV 419
F GKELCKSINPDEAVAYGA VQAA+LS EG++ V +L+L DVTPLS G+ G +M+V
Sbjct: 1204FFNGKELCKSINPDEAVAYGAAVQAAILSGEGNEKVQDLLLLDVTPLSLGLETAGGVMTV 1383
Query: 420 MIPRNTSIPVTVEKTKTYYECYDYYNKYSVTVKVYQGERITAYDNKLLGLFNFS-VPPIK 478
+IPRNT+IP E+ + Y N+ V ++VY+GER DN LLG F S +PP
Sbjct: 1384LIPRNTTIPTKKEQVFSTYS----DNQPGVLIQVYEGERTRTRDNNLLGKFELSGIPP-- 1545
Query: 479 GQLTPRG-HLIHLSFSIDVDGILNVSAKEETSGYKKGIAMTNQYCRLSTQEIARMIQEAE 537
PRG I + F ID +GILNVSA+++T+G K I +TN RLS ++I +M+QEAE
Sbjct: 1546---APRGVPQITVCFDIDANGILNVSAEDKTTGQKNKITITNDKGRLSKEDIEKMVQEAE 1716
Query: 538 IFKAQDMEFKKKVTAINALDDYLHNARKLMKDDGVGSKLTPVDKEKINSAMIKGESLIDG 597
+K++D E K+KV A NAL++Y +N R +KDD + SKL+ DK+KI A+ +DG
Sbjct: 1717KYKSEDEEHKRKVEAKNALENYAYNMRNTIKDDKIASKLSADDKKKIEDAIEGAIQWLDG 1896
Query: 598 NQQEDMSVFVDLLKELESI 616
NQ + F D +KELE I
Sbjct: 1897NQLGEADEFEDKMKELEGI 1953
>TC85390 homologue to SP|P27322|HS72_LYCES Heat shock cognate 70 kDa protein
2. [Tomato] {Lycopersicon esculentum}, complete
Length = 2278
Score = 745 bits (1923), Expect = 0.0
Identities = 389/619 (62%), Positives = 479/619 (76%), Gaps = 3/619 (0%)
Frame = +1
Query: 1 MEEKYEGVAIGIDLGTTYSCVGVWHEQNDRVEIIHNDQGNRTTPSCVAFTNSQRLIGDAA 60
M K EG AIGIDLGTTYSCVGVW Q+DRVEII NDQGNRTTPS VAFT+++RLIGDAA
Sbjct: 136 MAGKGEGPAIGIDLGTTYSCVGVW--QHDRVEIIANDQGNRTTPSYVAFTDTERLIGDAA 309
Query: 61 KNQAATNPSNTIFDTKRLIGRKYSDSIVQKDCLLWPFKVIAGTNDKPTIVVNYKGEEKRF 120
KNQ A NP+NT+FD KRLIGR++ D VQ D LWPFKVI G DKP IVVNYK EEK+F
Sbjct: 310 KNQVAMNPTNTVFDAKRLIGRRFGDVSVQSDMKLWPFKVIPGPADKPMIVVNYKAEEKQF 489
Query: 121 VAEEISSVILSKMREIAEAFLQTPVKNAVISVPAYFNDSQRRATKDAGAIAGLNVMQIIN 180
AEEISS++L KM+EIAEA+L T +KNAV++VPAYFNDSQR+ATKDAG I+GLNV++IIN
Sbjct: 490 SAEEISSMVLMKMKEIAEAYLGTTIKNAVVTVPAYFNDSQRQATKDAGVISGLNVLRIIN 669
Query: 181 EPTAAALAYGLQKRANCVDKRNIFIFDLGGGTFDVSILTVKNNVFEVKATAGDTHLGGQD 240
EPTAAA+AYGL K+A ++N+ IFDLGGGTFDVS+LT++ +FEVKATAGDTHLGG+D
Sbjct: 670 EPTAAAIAYGLDKKATSTGEKNVLIFDLGGGTFDVSLLTIEEGIFEVKATAGDTHLGGED 849
Query: 241 FDNIMVDHFVKEFRNKYKEDISENSKALRRLRTACEKAKRTLSYDTDATIELDAIYKGID 300
FDN MV+HFV+EF+ K K+DIS N +ALRRLRTACE+AKRTLS TIE+D++++GID
Sbjct: 850 FDNRMVNHFVQEFKRKNKKDISGNPRALRRLRTACERAKRTLSSTAQTTIEIDSLFEGID 1029
Query: 301 FCSSITRAKFEEMNMNLFEKCMNTVNICLADAKMDKNSVDDVVLVGGSSRIPKVRQLLQE 360
F ++ITRA+FEE+NM+LF KCM V CL DAKMDKNSV DVVLVGGS+RIPKV+QLLQ+
Sbjct: 1030FYTTITRARFEELNMDLFRKCMEPVEKCLRDAKMDKNSVHDVVLVGGSTRIPKVQQLLQD 1209
Query: 361 VFKGKELCKSINPDEAVAYGAVVQAALLS-EGSKNVPNLVLRDVTPLSFGVSELGNLMSV 419
F GKELCKSINPDEAVAYGA VQAA+LS EG++ V +L+L DVTPLS G+ G +M+V
Sbjct: 1210FFNGKELCKSINPDEAVAYGAAVQAAILSGEGNEKVQDLLLLDVTPLSLGLETAGGVMTV 1389
Query: 420 MIPRNTSIPVTVEKTKTYYECYDYYNKYSVTVKVYQGERITAYDNKLLGLFNFS-VPPIK 478
+IPRNT+IP E+ + Y N+ V ++VY+GER DN LLG F S +PP
Sbjct: 1390LIPRNTTIPTKKEQVFSTYS----DNQPGVLIQVYEGERTRTRDNNLLGKFELSGIPP-- 1551
Query: 479 GQLTPRG-HLIHLSFSIDVDGILNVSAKEETSGYKKGIAMTNQYCRLSTQEIARMIQEAE 537
PRG I + F ID +GILNVSA+++T+G K I +TN RLS +EI +M+QEAE
Sbjct: 1552---APRGVPQITVCFDIDANGILNVSAEDKTTGQKNKITITNDKGRLSKEEIEKMVQEAE 1722
Query: 538 IFKAQDMEFKKKVTAINALDDYLHNARKLMKDDGVGSKLTPVDKEKINSAMIKGESLIDG 597
+K++D E KKKV A NAL++Y +N R +KDD + SKL+ DK+KI A+ +DG
Sbjct: 1723KYKSEDEEHKKKVEAKNALENYAYNMRNTIKDDKIASKLSADDKKKIEDAIEGAIQWLDG 1902
Query: 598 NQQEDMSVFVDLLKELESI 616
NQ + F D +KELE I
Sbjct: 1903NQLGEADEFEDKMKELEGI 1959
>TC85396 homologue to GP|6911551|emb|CAB72129.1 heat shock protein 70
{Cucumis sativus}, complete
Length = 2320
Score = 738 bits (1905), Expect = 0.0
Identities = 391/643 (60%), Positives = 489/643 (75%), Gaps = 3/643 (0%)
Frame = +2
Query: 1 MEEKYEGVAIGIDLGTTYSCVGVWHEQNDRVEIIHNDQGNRTTPSCVAFTNSQRLIGDAA 60
M K EG AIGIDLGTTYSCVGVW Q+DRVEII NDQGNRTTPS VAFT+S+RLIGDAA
Sbjct: 107 MAGKGEGPAIGIDLGTTYSCVGVW--QHDRVEIIANDQGNRTTPSYVAFTDSERLIGDAA 280
Query: 61 KNQAATNPSNTIFDTKRLIGRKYSDSIVQKDCLLWPFKVIAGTNDKPTIVVNYKGEEKRF 120
KNQ A NP+NT+FD KRLIGR+ SD+ VQ D LWPFKVIAG +KP I VNYKGEEK F
Sbjct: 281 KNQVAMNPTNTVFDAKRLIGRRISDASVQSDMKLWPFKVIAGPGEKPMIGVNYKGEEKLF 460
Query: 121 VAEEISSVILSKMREIAEAFLQTPVKNAVISVPAYFNDSQRRATKDAGAIAGLNVMQIIN 180
+EEISS++L KMREIAEA+L +KNAV++VPAYFNDSQR+ATKDAG IAGLNV++IIN
Sbjct: 461 ASEEISSMVLIKMREIAEAYLGVTIKNAVVTVPAYFNDSQRQATKDAGVIAGLNVLRIIN 640
Query: 181 EPTAAALAYGLQKRANCVDKRNIFIFDLGGGTFDVSILTVKNNVFEVKATAGDTHLGGQD 240
EPTAAA+AYGL K+A V ++N+ IFDLGGGTFDVS+LT++ +FEVKATAGDTHLGG+D
Sbjct: 641 EPTAAAIAYGLDKKATSVGEKNVLIFDLGGGTFDVSLLTIEEGIFEVKATAGDTHLGGED 820
Query: 241 FDNIMVDHFVKEFRNKYKEDISENSKALRRLRTACEKAKRTLSYDTDATIELDAIYKGID 300
FDN MV HFV+EF+ K K+DIS N +ALRRLRTACE+AKRTLS TIE+D++++G+D
Sbjct: 821 FDNRMVTHFVQEFKRKNKKDISGNPRALRRLRTACERAKRTLSSTAQTTIEIDSLFEGVD 1000
Query: 301 FCSSITRAKFEEMNMNLFEKCMNTVNICLADAKMDKNSVDDVVLVGGSSRIPKVRQLLQE 360
F ++ITRA+FEE+NM+LF KCM V CL DAKMDK++V DVVLVGGS+RIPKV+QLLQ+
Sbjct: 1001FYTTITRARFEELNMDLFRKCMEPVEKCLRDAKMDKSTVHDVVLVGGSTRIPKVQQLLQD 1180
Query: 361 VFKGKELCKSINPDEAVAYGAVVQAALLS-EGSKNVPNLVLRDVTPLSFGVSELGNLMSV 419
F GKELCKSINPDEAVAYGA VQAA+LS EG++ V +L+L DVTPLS G+ G +M+V
Sbjct: 1181FFNGKELCKSINPDEAVAYGAAVQAAILSGEGNEKVQDLLLLDVTPLSQGLETAGGVMTV 1360
Query: 420 MIPRNTSIPVTVEKTKTYYECYDYYNKYSVTVKVYQGERITAYDNKLLGLFNFS-VPPIK 478
+IPRNT+IP E+ + Y N+ V ++VY+GER DN LLG F S +PP
Sbjct: 1361LIPRNTTIPTKKEQVFSTYS----DNQPGVLIQVYEGERTRTKDNNLLGKFELSGIPP-- 1522
Query: 479 GQLTPRG-HLIHLSFSIDVDGILNVSAKEETSGYKKGIAMTNQYCRLSTQEIARMIQEAE 537
PRG I + F ID +GILNVSA+++T+G K I +TN RLS +EI +M+QEAE
Sbjct: 1523---APRGVPQITVCFDIDANGILNVSAEDKTTGQKNKITITNDKGRLSKEEIEKMVQEAE 1693
Query: 538 IFKAQDMEFKKKVTAINALDDYLHNARKLMKDDGVGSKLTPVDKEKINSAMIKGESLIDG 597
+K++D E KKKV A N+L++Y +N R +KD+ + SKL+ DK++I A+ +D
Sbjct: 1694KYKSEDEEHKKKVEAKNSLENYAYNMRNTIKDEKISSKLSGGDKKQIEDAIEGAIQWLDA 1873
Query: 598 NQQEDMSVFVDLLKELESIFESSKNKINKGYSDEESDWVLVDD 640
NQ + F D +KELE+I K+ +G + E + VDD
Sbjct: 1874NQLAEADEFEDKMKELETICNPIIAKMYQGGAGEGPE---VDD 1993
>TC85389 homologue to SP|Q03684|BIP4_TOBAC Luminal binding protein 4
precursor (BiP 4) (78 kDa glucose-regulated protein
homolog 4) (GRP 78-4)., partial (94%)
Length = 2265
Score = 573 bits (1478), Expect = e-164
Identities = 306/615 (49%), Positives = 435/615 (69%), Gaps = 5/615 (0%)
Frame = +2
Query: 7 GVAIGIDLGTTYSCVGVWHEQNDRVEIIHNDQGNRTTPSCVAFTNSQRLIGDAAKNQAAT 66
G IGIDLGTTYSCVGV+ +N VEII NDQGNR TPS V+FT+ +RLIG+AAKN AA
Sbjct: 155 GTVIGIDLGTTYSCVGVY--KNGHVEIIANDQGNRITPSWVSFTDDERLIGEAAKNLAAV 328
Query: 67 NPSNTIFDTKRLIGRKYSDSIVQKDCLLWPFKVIAGTNDKPTIVVNYK-GEEKRFVAEEI 125
NP TIFD KRLIGRK++D VQ+D L P+K++ + KP I V K GE K F EE+
Sbjct: 329 NPERTIFDVKRLIGRKFADKEVQRDMKLVPYKIV-NKDGKPYIQVRVKDGETKVFSPEEV 505
Query: 126 SSVILSKMREIAEAFLQTPVKNAVISVPAYFNDSQRRATKDAGAIAGLNVMQIINEPTAA 185
S++IL+KM+E AEAFL +++AV++VPAYFND+QR+ATKDAG IAGLNV +IINEPTAA
Sbjct: 506 SAMILTKMKETAEAFLGKTIRDAVVTVPAYFNDAQRQATKDAGVIAGLNVARIINEPTAA 685
Query: 186 ALAYGLQKRANCVDKRNIFIFDLGGGTFDVSILTVKNNVFEVKATAGDTHLGGQDFDNIM 245
A+AYGL K+ ++NI +FDLGGGTFDVSILT+ N VFEV +T GDTHLGG+DFD +
Sbjct: 686 AIAYGLDKKGG---EKNILVFDLGGGTFDVSILTIDNGVFEVLSTNGDTHLGGEDFDQRI 856
Query: 246 VDHFVKEFRNKYKEDISENSKALRRLRTACEKAKRTLSYDTDATIELDAIYKGIDFCSSI 305
+++F+K + K+ +DIS++++AL +LR E+AKR LS +E+++++ G+DF +
Sbjct: 857 MEYFIKLIKKKHSKDISKDNRALGKLRRESERAKRALSSQHQVRVEIESLFDGVDFSEPL 1036
Query: 306 TRAKFEEMNMNLFEKCMNTVNICLADAKMDKNSVDDVVLVGGSSRIPKVRQLLQEVFKGK 365
TRA+FEE+N +LF K M V + DA + KN +D++VLVGGS+RIPKV+QLL++ F GK
Sbjct: 1037TRARFEELNNDLFRKTMGPVKKAMDDAGLQKNQIDEIVLVGGSTRIPKVQQLLKDYFDGK 1216
Query: 366 ELCKSINPDEAVAYGAVVQAALLS-EGSKNVPNLVLRDVTPLSFGVSELGNLMSVMIPRN 424
E K +NPDEAVA+GA VQ ++LS EG + +++L DV PL+ G+ +G +M+ +IPRN
Sbjct: 1217EPNKGVNPDEAVAFGAAVQGSILSGEGGEETKDILLLDVAPLTLGIETVGGVMTKLIPRN 1396
Query: 425 TSIPVTVEKTKTYYECYDYYNKYSVTVKVYQGERITAYDNKLLGLFNFS-VPPIKGQLTP 483
T IP + T Y+ + +V+++V++GER D +LLG F+ S +PP P
Sbjct: 1397TVIPTKKSQVFTTYQ----DQQTTVSIQVFEGERSLTKDCRLLGNFDLSGIPP-----AP 1549
Query: 484 RG-HLIHLSFSIDVDGILNVSAKEETSGYKKGIAMTNQYCRLSTQEIARMIQEAEIFKAQ 542
RG I ++F +D +GILNV A+++ +G + I +TN+ RLS +EI RM++EAE F +
Sbjct: 1550RGTPQIEVTFEVDANGILNVRAEDKGTGKSEKITITNEKGRLSQEEIDRMVREAEEFAEE 1729
Query: 543 DMEFKKKVTAINALDDYLHNARKLMKD-DGVGSKLTPVDKEKINSAMIKGESLIDGNQQE 601
D + K+++ A NAL+ Y++N + + D D + KL +KEKI +A+ + +D NQ
Sbjct: 1730DKKVKERIDARNALETYVYNMKNQISDKDKLADKLESDEKEKIEAAVKEALEWLDDNQTV 1909
Query: 602 DMSVFVDLLKELESI 616
+ F + LKE+E++
Sbjct: 1910EKEEFEEKLKEVEAV 1954
>TC80733 homologue to SP|Q01233|HS70_NEUCR Heat shock 70 kDa protein
(HSP70). {Neurospora crassa}, partial (54%)
Length = 1258
Score = 428 bits (1101), Expect(2) = e-123
Identities = 217/335 (64%), Positives = 271/335 (80%)
Frame = +2
Query: 9 AIGIDLGTTYSCVGVWHEQNDRVEIIHNDQGNRTTPSCVAFTNSQRLIGDAAKNQAATNP 68
AIGIDLGTTYSCVG + ND++EII NDQGNRTTPS VAF +++RLIGDAAKNQ A NP
Sbjct: 188 AIGIDLGTTYSCVGRY--ANDKIEIIANDQGNRTTPSYVAFNDTERLIGDAAKNQVAMNP 361
Query: 69 SNTIFDTKRLIGRKYSDSIVQKDCLLWPFKVIAGTNDKPTIVVNYKGEEKRFVAEEISSV 128
NT+FD KRLIGRK+SDS VQ D +PFKVI KP I V +KGE K F EEIS++
Sbjct: 362 HNTVFDAKRLIGRKFSDSEVQADMKHFPFKVI-DKGGKPNIEVEFKGENKTFTPEEISAM 538
Query: 129 ILSKMREIAEAFLQTPVKNAVISVPAYFNDSQRRATKDAGAIAGLNVMQIINEPTAAALA 188
+L KMRE AEA+L V NAVI+VPAYFNDSQR+ATKDAG IAGLNV++IINEPTAAA+A
Sbjct: 539 VLVKMRETAEAYLGGQVTNAVITVPAYFNDSQRQATKDAGLIAGLNVLRIINEPTAAAIA 718
Query: 189 YGLQKRANCVDKRNIFIFDLGGGTFDVSILTVKNNVFEVKATAGDTHLGGQDFDNIMVDH 248
YGL K+A +RN+ IFDLGGGTFDVS+LT++ +FEVK+TAGDTHLGG+DFDN +V+H
Sbjct: 719 YGLDKKAE--GERNVLIFDLGGGTFDVSLLTIEEGIFEVKSTAGDTHLGGEDFDNRLVNH 892
Query: 249 FVKEFRNKYKEDISENSKALRRLRTACEKAKRTLSYDTDATIELDAIYKGIDFCSSITRA 308
FV EF+ K+K+D+S N++ALRRLRTACE+AKRTLS +IE+D++++GIDF +SITRA
Sbjct: 893 FVNEFKRKHKKDLSSNARALRRLRTACERAKRTLSSSAQTSIEIDSLFEGIDFYTSITRA 1072
Query: 309 KFEEMNMNLFEKCMNTVNICLADAKMDKNSVDDVV 343
+FEE+ +LF + V+ L+DAK+DK+ V ++V
Sbjct: 1073RFEELCQDLFRSTIQPVDRVLSDAKIDKSQVHEIV 1177
Score = 32.7 bits (73), Expect(2) = e-123
Identities = 12/22 (54%), Positives = 19/22 (85%)
Frame = +3
Query: 345 VGGSSRIPKVRQLLQEVFKGKE 366
VGGS+RIP++++L+ + F GKE
Sbjct: 1182 VGGSTRIPRIQKLISDYFNGKE 1247
>TC85718 homologue to SP|Q01899|HS7M_PHAVU Heat shock 70 kDa protein
mitochondrial precursor. [Kidney bean French bean]
{Phaseolus vulgaris}, complete
Length = 2371
Score = 429 bits (1102), Expect = e-120
Identities = 267/632 (42%), Positives = 377/632 (59%), Gaps = 8/632 (1%)
Frame = +2
Query: 10 IGIDLGTTYSCVGVWHEQNDRVEIIHNDQGNRTTPSCVAFTNS-QRLIGDAAKNQAATNP 68
IGIDLGTT SCV V +N +V + N +G RTTPS VAFT + L+G AK QA TNP
Sbjct: 263 IGIDLGTTNSCVSVMEGKNPKV--VENSEGARTTPSVVAFTQKGELLVGTPAKRQAVTNP 436
Query: 69 SNTIFDTKRLIGRKYSDSIVQKDCLLWPFKVIAGTNDKPTIVVNYKGEEKRFVAEEISSV 128
NTI KRLIGR++ D QK+ + P+K++ N V KG++ + +I +
Sbjct: 437 ENTISGAKRLIGRRFDDPQTQKEMKMVPYKIVKAPNGDAW--VEAKGQQ--YSPSQIGAF 604
Query: 129 ILSKMREIAEAFLQTPVKNAVISVPAYFNDSQRRATKDAGAIAGLNVMQIINEPTAAALA 188
+L+KM+E AEA+L V AVI+VPAYFND+QR+ATKDAG IAGL V++IINEPTAAAL+
Sbjct: 605 VLTKMKETAEAYLGKTVPKAVITVPAYFNDAQRQATKDAGRIAGLEVLRIINEPTAAALS 784
Query: 189 YGLQKRANCVDKRNIFIFDLGGGTFDVSILTVKNNVFEVKATAGDTHLGGQDFDNIMVDH 248
YG+ K I +FDLGGGTFDVSIL + N VFEVKAT GDT LGG+DFDN ++D
Sbjct: 785 YGMNKEGL------IAVFDLGGGTFDVSILEISNGVFEVKATNGDTFLGGEDFDNALLDF 946
Query: 249 FVKEFRNKYKEDISENSKALRRLRTACEKAKRTLSYDTDATIELDAIYKGIDFCS----S 304
V EF+ D+S++ AL+RLR A EKAK LS + I L I +
Sbjct: 947 LVNEFKRSESIDLSKDKLALQRLREAAEKAKIELSSTSQTEINLPFITADASGAKHLNIT 1126
Query: 305 ITRAKFEEMNMNLFEKCMNTVNICLADAKMDKNSVDDVVLVGGSSRIPKVRQLLQEVFKG 364
+TR+KFE + NL E+ CL DA + +D+V+LVGG +R+PKV++++ +F G
Sbjct: 1127LTRSKFEALVNNLIERTKAPCQSCLKDANISTKDIDEVLLVGGMTRVPKVQEVVSTIF-G 1303
Query: 365 KELCKSINPDEAVAYGAVVQAALLSEGSKNVPNLVLRDVTPLSFGVSELGNLMSVMIPRN 424
K CK +NPDEAVA GA +Q +L +V L+L DVTPLS G+ LG + + +I RN
Sbjct: 1304KSPCKGVNPDEAVAMGAALQGGIL---RGDVKELLLLDVTPLSLGIETLGGIFTRLINRN 1474
Query: 425 TSIPVTVEKTKTYYECYDYYNKYSVTVKVYQGERITAYDNKLLGLFNF-SVPPIKGQLTP 483
T+IP +K++ + D N+ V +KV QGER A DNK+LG F +PP P
Sbjct: 1475TTIP--TKKSQVFSTAAD--NQTQVGIKVLQGEREMASDNKMLGEFELVGLPP-----AP 1627
Query: 484 RG-HLIHLSFSIDVDGILNVSAKEETSGYKKGIAMTNQYCRLSTQEIARMIQEAEIFKAQ 542
RG I ++F ID +GI+ VSAK++++G ++ I + + LS EI M++EAE+ +
Sbjct: 1628RGMPQIEVTFDIDANGIVTVSAKDKSTGKEQQITIKSS-GGLSDDEIQNMVKEAELHAQK 1804
Query: 543 DMEFKKKVTAINALDDYLHNARKLMKDDGVGSKLTPVDKEKINSAMIKGESLIDGNQQED 602
D E K + N+ D +++ K + + K+ ++I A+ S + G+ ++
Sbjct: 1805DQERKSLIDIRNSADTSIYSIEKSLSE--YREKIPAEVAKEIEDAVSDLRSAMAGDSADE 1978
Query: 603 MSVFVDLL-KELESIFESSKNKINKGYSDEES 633
+ +D K + I E + G S + S
Sbjct: 1979IKSKLDAANKAVSKIGEHMSGGSSSGPSSDGS 2074
>TC80669 homologue to PIR|S53498|S53498 dnaK-type molecular chaperone
HSP71.2 - garden pea, partial (44%)
Length = 955
Score = 427 bits (1098), Expect = e-120
Identities = 211/284 (74%), Positives = 245/284 (85%)
Frame = +2
Query: 6 EGVAIGIDLGTTYSCVGVWHEQNDRVEIIHNDQGNRTTPSCVAFTNSQRLIGDAAKNQAA 65
EG AIGIDLGTTYSCVGVW QNDRVEII NDQGNRTTPS VAFT+++RLIGDAAKNQ A
Sbjct: 110 EGKAIGIDLGTTYSCVGVW--QNDRVEIIPNDQGNRTTPSYVAFTDTERLIGDAAKNQVA 283
Query: 66 TNPSNTIFDTKRLIGRKYSDSIVQKDCLLWPFKVIAGTNDKPTIVVNYKGEEKRFVAEEI 125
NP NT+FD KRLIGR++SD VQ D LWPFKV+ G +KP IVVNYKGEEK+F AEEI
Sbjct: 284 MNPQNTVFDAKRLIGRRFSDESVQNDMKLWPFKVVPGPAEKPMIVVNYKGEEKKFAAEEI 463
Query: 126 SSVILSKMREIAEAFLQTPVKNAVISVPAYFNDSQRRATKDAGAIAGLNVMQIINEPTAA 185
SS++L KMRE+AEAFL PVKNAV++VPAYFNDSQR+ATKDAGAI+GLNV++IINEPTAA
Sbjct: 464 SSMVLIKMREVAEAFLGHPVKNAVVTVPAYFNDSQRQATKDAGAISGLNVLRIINEPTAA 643
Query: 186 ALAYGLQKRANCVDKRNIFIFDLGGGTFDVSILTVKNNVFEVKATAGDTHLGGQDFDNIM 245
A+AYGL K+A+ ++N+ IFDLGGGTFDVS+LT++ +FEVKATAGDTHLGG+DFDN M
Sbjct: 644 AIAYGLDKKASRKGEQNVLIFDLGGGTFDVSLLTIEEGIFEVKATAGDTHLGGEDFDNRM 823
Query: 246 VDHFVKEFRNKYKEDISENSKALRRLRTACEKAKRTLSYDTDAT 289
V+HFV EFR K K+DIS N++ALRRLRTACE+AKRTLS T
Sbjct: 824 VNHFVSEFRRKNKKDISGNARALRRLRTACERAKRTLSSTAQTT 955
>TC85716 homologue to PIR|S19140|S19140 dnaK-type molecular chaperone PHSP1
precursor mitochondrial - garden pea, partial (98%)
Length = 2422
Score = 415 bits (1066), Expect = e-116
Identities = 253/629 (40%), Positives = 373/629 (59%), Gaps = 8/629 (1%)
Frame = +1
Query: 10 IGIDLGTTYSCVGVWHEQNDRVEIIHNDQGNRTTPSCVAFTNS-QRLIGDAAKNQAATNP 68
IGIDLGTT SCV + +N +V I N +G RTTPS VAF + L+G AK QA TNP
Sbjct: 247 IGIDLGTTNSCVSLMEGKNPKV--IENSEGARTTPSVVAFNQKGELLVGTPAKRQAVTNP 420
Query: 69 SNTIFDTKRLIGRKYSDSIVQKDCLLWPFKVIAGTNDKPTIVVNYKGEEKRFVAEEISSV 128
+NT+F TKRLIGR++ D QK+ + P+K++ N + +N ++++ +I +
Sbjct: 421 TNTLFGTKRLIGRRFDDPQTQKEMKMVPYKIVKAPNGDAWVEIN----KQQYSPSQIGAF 588
Query: 129 ILSKMREIAEAFLQTPVKNAVISVPAYFNDSQRRATKDAGAIAGLNVMQIINEPTAAALA 188
+L+KM+E AEA+L + AV++VPAYFND+QR+ATKDAG IAGL V +IINEPTAAAL+
Sbjct: 589 VLTKMKETAEAYLGKTISKAVVTVPAYFNDAQRQATKDAGRIAGLEVKRIINEPTAAALS 768
Query: 189 YGLQKRANCVDKRNIFIFDLGGGTFDVSILTVKNNVFEVKATAGDTHLGGQDFDNIMVDH 248
YG+ + I +FDLGGGTFDVSIL + N VFEVKAT GDT LGG+DFDN ++D
Sbjct: 769 YGMNNKEGL-----IAVFDLGGGTFDVSILEISNGVFEVKATNGDTFLGGEDFDNALLDF 933
Query: 249 FVKEFRNKYKEDISENSKALRRLRTACEKAKRTLSYDTDATIELDAIYKGIDFCS----S 304
V EF+ D++++ AL+RLR A EKAK LS + I L I +
Sbjct: 934 LVSEFKRTDSIDLAKDKLALQRLREAAEKAKIELSSTSQTEINLPFITADASGAKHLNIT 1113
Query: 305 ITRAKFEEMNMNLFEKCMNTVNICLADAKMDKNSVDDVVLVGGSSRIPKVRQLLQEVFKG 364
+TR+KFE + NL E+ CL DA + VD+V+LVGG +R+PKV++++ E+F G
Sbjct: 1114LTRSKFEALVNNLIERTKAPCKSCLKDANISIKDVDEVLLVGGMTRVPKVQEVVSEIF-G 1290
Query: 365 KELCKSINPDEAVAYGAVVQAALLSEGSKNVPNLVLRDVTPLSFGVSELGNLMSVMIPRN 424
K K +NPDEAVA GA +Q +L +V L+L DVTPLS G+ LG + + +I RN
Sbjct: 1291KSPSKGVNPDEAVAMGAALQGGIL---RGDVKELLLLDVTPLSLGIETLGGIFTRLISRN 1461
Query: 425 TSIPVTVEKTKTYYECYDYYNKYSVTVKVYQGERITAYDNKLLGLFNF-SVPPIKGQLTP 483
T+IP +K++ + D + +G R A DNK LG F+ +PP P
Sbjct: 1462TTIP--TKKSQVFSTAADNQTQRGYXRCSQEGXREMAADNKSLGEFDLVGIPP-----AP 1620
Query: 484 RG-HLIHLSFSIDVDGILNVSAKEETSGYKKGIAMTNQYCRLSTQEIARMIQEAEIFKAQ 542
RG I ++F ID +GI+ VSAK++++G ++ I + + LS EI M++EAE+ +
Sbjct: 1621RGLPQIEVTFDIDANGIVTVSAKDKSTGKEQQITIRSS-GGLSDDEINNMVKEAELHAQR 1797
Query: 543 DMEFKKKVTAINALDDYLHNARKLMKDDGVGSKLTPVDKEKINSAMIKGESLIDGNQQED 602
D E K + N+ D +++ K + + K+ ++I +++ + ++G ++
Sbjct: 1798DQERKALIDIKNSADTSIYSIEKSLSE--YREKIPSEVAKEIENSVSDLRTAMEGESVDE 1971
Query: 603 MSVFVDLL-KELESIFESSKNKINKGYSD 630
+ +D K + I + + G SD
Sbjct: 1972IKTKLDAANKAVSKIGQHMSGGSSGGSSD 2058
>TC87268 homologue to PIR|S53500|S44168 dnaK-type molecular chaperone
HSC71.0 - garden pea, partial (59%)
Length = 1336
Score = 350 bits (898), Expect = 1e-96
Identities = 193/365 (52%), Positives = 252/365 (68%), Gaps = 3/365 (0%)
Frame = +2
Query: 266 KALRRLRTACEKAKRTLSYDTDATIELDAIYKGIDFCSSITRAKFEEMNMNLFEKCMNTV 325
+ALRRLRTACE+AKRTLS TIE+D++++GIDF S ITRA+FEE+NM+LF KCM V
Sbjct: 2 RALRRLRTACERAKRTLSSTAQTTIEIDSLFEGIDFYSPITRARFEELNMDLFRKCMEPV 181
Query: 326 NICLADAKMDKNSVDDVVLVGGSSRIPKVRQLLQEVFKGKELCKSINPDEAVAYGAVVQA 385
CL DAKMDK SV DVVLVGGS+RIPKV+QLLQ+ F GKELCKSINPDEAVAYGA VQA
Sbjct: 182 EKCLRDAKMDKKSVHDVVLVGGSTRIPKVQQLLQDFFNGKELCKSINPDEAVAYGAAVQA 361
Query: 386 ALLS-EGSKNVPNLVLRDVTPLSFGVSELGNLMSVMIPRNTSIPVTVEKTKTYYECYDYY 444
A+LS EG++ V +L+L DVTPLS G+ G +M+V+IPRNT+IP E+ + Y
Sbjct: 362 AILSGEGNEKVQDLLLLDVTPLSLGLETAGGVMTVLIPRNTTIPTKKEQVFSTYS----D 529
Query: 445 NKYSVTVKVYQGERITAYDNKLLGLFNFS-VPPIKGQLTPRG-HLIHLSFSIDVDGILNV 502
N+ V ++V++GER DN LLG F S +PP PRG I + F ID +GILNV
Sbjct: 530 NQPGVLIQVFEGERTRTRDNNLLGKFELSGIPP-----APRGVPQITVCFDIDANGILNV 694
Query: 503 SAKEETSGYKKGIAMTNQYCRLSTQEIARMIQEAEIFKAQDMEFKKKVTAINALDDYLHN 562
SA+++T+G K I +TN RLS ++I +M+QEAE +K++D E KKKV A NAL++Y +N
Sbjct: 695 SAEDKTTGQKNKITITNDKGRLSKEDIEKMVQEAEKYKSEDEEHKKKVEAKNALENYAYN 874
Query: 563 ARKLMKDDGVGSKLTPVDKEKINSAMIKGESLIDGNQQEDMSVFVDLLKELESIFESSKN 622
R +KD+ + KL DK+KI + +D NQ + F D +KELE +
Sbjct: 875 MRNTIKDEKIAGKLDSDDKKKIEDTIEAAIQWLDANQLAEADEFEDKMKELEGVCNPIIA 1054
Query: 623 KINKG 627
K+ +G
Sbjct: 1055KMYQG 1069
>TC80756 homologue to PIR|S53498|S53498 dnaK-type molecular chaperone
HSP71.2 - garden pea, partial (53%)
Length = 1189
Score = 298 bits (763), Expect = 4e-81
Identities = 165/326 (50%), Positives = 223/326 (67%), Gaps = 3/326 (0%)
Frame = +3
Query: 305 ITRAKFEEMNMNLFEKCMNTVNICLADAKMDKNSVDDVVLVGGSSRIPKVRQLLQEVFKG 364
ITRA+FEE+NM+LF KCM V CL DAK+DK+ V +VVLVGGS+RIPKV+QLLQ+ F G
Sbjct: 3 ITRARFEELNMDLFRKCMEPVEKCLRDAKIDKSHVHEVVLVGGSTRIPKVQQLLQDFFNG 182
Query: 365 KELCKSINPDEAVAYGAVVQAALLS-EGSKNVPNLVLRDVTPLSFGVSELGNLMSVMIPR 423
KELCKSINPDEAVAYGA VQAA+LS EG + V +L+L DVTPLS G+ G +M+ +IPR
Sbjct: 183 KELCKSINPDEAVAYGAAVQAAILSGEGDEKVQDLLLLDVTPLSLGLETAGGVMTTLIPR 362
Query: 424 NTSIPVTVEKTKTYYECYDYYNKYSVTVKVYQGERITAYDNKLLGLFNFS-VPPIKGQLT 482
NT+IP E+ + Y N+ V ++V++GER DN LLG F + +PP
Sbjct: 363 NTTIPTKKEQIFSTYS----DNQPGVLIQVFEGERARTKDNNLLGKFELTGIPP-----A 515
Query: 483 PRG-HLIHLSFSIDVDGILNVSAKEETSGYKKGIAMTNQYCRLSTQEIARMIQEAEIFKA 541
PRG I++ F ID +GILNVSA+++T+G K I +TN RLS +EI +M+++AE +KA
Sbjct: 516 PRGVPQINVCFDIDANGILNVSAEDKTAGVKNKITITNDKGRLSKEEIEKMVKDAEKYKA 695
Query: 542 QDMEFKKKVTAINALDDYLHNARKLMKDDGVGSKLTPVDKEKINSAMIKGESLIDGNQQE 601
+D E KKKV A N++++Y +N R +KD+ +G KL+ DKEKI A+ ++GNQ
Sbjct: 696 EDEEVKKKVEAKNSIENYAYNMRNTIKDEKIGGKLSHEDKEKIEKAVEDAIQWLEGNQMA 875
Query: 602 DMSVFVDLLKELESIFESSKNKINKG 627
++ F D KELE I K+ +G
Sbjct: 876 EVDEFEDKQKELEGICNPIIAKMYQG 953
>BQ139103 homologue to GP|2655420|gb| heat shock cognate protein HSC70
{Brassica napus}, partial (29%)
Length = 628
Score = 289 bits (739), Expect = 3e-78
Identities = 145/193 (75%), Positives = 166/193 (85%)
Frame = +2
Query: 1 MEEKYEGVAIGIDLGTTYSCVGVWHEQNDRVEIIHNDQGNRTTPSCVAFTNSQRLIGDAA 60
M K EG AIGIDLGTTYSCVGVW Q+DRVEII NDQGNRTTPS VAFT+S+RLIGDAA
Sbjct: 56 MAGKGEGPAIGIDLGTTYSCVGVW--QHDRVEIIANDQGNRTTPSYVAFTDSERLIGDAA 229
Query: 61 KNQAATNPSNTIFDTKRLIGRKYSDSIVQKDCLLWPFKVIAGTNDKPTIVVNYKGEEKRF 120
KNQ A NP NT+FD KRLIGR++SD+ VQ D LWPFK+I+G +KP I VNYKGE+K F
Sbjct: 230 KNQVAMNPINTVFDAKRLIGRRFSDASVQSDMKLWPFKIISGPAEKPLIGVNYKGEDKEF 409
Query: 121 VAEEISSVILSKMREIAEAFLQTPVKNAVISVPAYFNDSQRRATKDAGAIAGLNVMQIIN 180
AEEISS++L KMREIAEA+L + +KNAV++VPAYFNDSQR+ATKDAG IAGLNVM+IIN
Sbjct: 410 AAEEISSMVLMKMREIAEAYLGSAIKNAVVTVPAYFNDSQRQATKDAGVIAGLNVMRIIN 589
Query: 181 EPTAAALAYGLQK 193
EPTAAA+AYGL K
Sbjct: 590 EPTAAAIAYGLDK 628
>TC85394 homologue to GP|2642238|gb|AAB86942.1| endoplasmic reticulum
HSC70-cognate binding protein precursor {Glycine max},
partial (38%)
Length = 962
Score = 279 bits (713), Expect = 3e-75
Identities = 148/230 (64%), Positives = 177/230 (76%), Gaps = 1/230 (0%)
Frame = +1
Query: 7 GVAIGIDLGTTYSCVGVWHEQNDRVEIIHNDQGNRTTPSCVAFTNSQRLIGDAAKNQAAT 66
G IGIDLGTTYSCVGV+ +N VEII NDQGNR TPS V+FT+ +RLIG+AAKN AA
Sbjct: 289 GTVIGIDLGTTYSCVGVY--KNGHVEIIANDQGNRITPSWVSFTDDERLIGEAAKNLAAV 462
Query: 67 NPSNTIFDTKRLIGRKYSDSIVQKDCLLWPFKVIAGTNDKPTIVVNYK-GEEKRFVAEEI 125
NP TIFD KRLIGRK+ D VQ+D L P+K++ + KP I V K GE K F EEI
Sbjct: 463 NPERTIFDVKRLIGRKFEDKEVQRDMKLVPYKIV-NRDGKPYIQVRVKDGETKVFSPEEI 639
Query: 126 SSVILSKMREIAEAFLQTPVKNAVISVPAYFNDSQRRATKDAGAIAGLNVMQIINEPTAA 185
S++IL KM+E AE FL +++AV++VPAYFND+QR+ATKDAG IAGLNV +IINEPTAA
Sbjct: 640 SAMILGKMKETAEGFLGKTIRDAVVTVPAYFNDAQRQATKDAGVIAGLNVARIINEPTAA 819
Query: 186 ALAYGLQKRANCVDKRNIFIFDLGGGTFDVSILTVKNNVFEVKATAGDTH 235
A+AYGL K+ ++NI +FDLGGGTFDVSILT+ N VFEV AT GDTH
Sbjct: 820 AIAYGLDKKGG---EKNILVFDLGGGTFDVSILTIDNGVFEVLATNGDTH 960
>TC80052 similar to PIR|JC4786|JC4786 dnaK-type molecular chaperone hsc70-3
- tomato, partial (27%)
Length = 667
Score = 256 bits (654), Expect = 2e-68
Identities = 133/194 (68%), Positives = 156/194 (79%)
Frame = +3
Query: 1 MEEKYEGVAIGIDLGTTYSCVGVWHEQNDRVEIIHNDQGNRTTPSCVAFTNSQRLIGDAA 60
M + + VAIGIDLGTTYSCV V +N ++II ND G RTTPS VAF +S+R+IGDAA
Sbjct: 81 MSGRGKKVAIGIDLGTTYSCVAVC--KNGEIDIIVNDLGKRTTPSFVAFKDSERMIGDAA 254
Query: 61 KNQAATNPSNTIFDTKRLIGRKYSDSIVQKDCLLWPFKVIAGTNDKPTIVVNYKGEEKRF 120
N AA+NP+NTIFD KRLIGRK+SD IVQ D LWPFKVI NDKP IVVNY EEK F
Sbjct: 255 FNIAASNPTNTIFDAKRLIGRKFSDPIVQSDVKLWPFKVIGDLNDKPMIVVNYNDEEKHF 434
Query: 121 VAEEISSVILSKMREIAEAFLQTPVKNAVISVPAYFNDSQRRATKDAGAIAGLNVMQIIN 180
AEEISS++L KMREIAE FL + V++ VI+VPAYFNDSQR++T+DAGAIAGLNVM+IIN
Sbjct: 435 AAEEISSMVLVKMREIAETFLGSIVEDVVITVPAYFNDSQRQSTRDAGAIAGLNVMRIIN 614
Query: 181 EPTAAALAYGLQKR 194
EPTAAA+AYG +
Sbjct: 615 EPTAAAIAYGFNTK 656
>TC77585 similar to GP|17473863|gb|AAL38353.1 putative heat-shock protein
{Arabidopsis thaliana}, partial (92%)
Length = 3027
Score = 248 bits (633), Expect = 5e-66
Identities = 152/453 (33%), Positives = 240/453 (52%), Gaps = 20/453 (4%)
Frame = +1
Query: 10 IGIDLGTTYSCVGVWHEQNDRVEIIHNDQGNRTTPSCVAFTNSQRLIGDAAKNQAATNPS 69
+G D G V V ++ ++++ ND+ R TP+ V F + QR IG A NP
Sbjct: 247 VGFDFGNESCIVAVARQRG--IDVVLNDESKRETPAIVCFGDKQRFIGTAGAASTMMNPK 420
Query: 70 NTIFDTKRLIGRKYSDSIVQKDCLLWPFKVIAGTNDKPTIVVNYKGEEKRFVAEEISSVI 129
N+I KRLIG+K++D +Q+D PF V G + P I Y GE + F A ++ ++
Sbjct: 421 NSISQIKRLIGKKFADPELQRDLKSLPFNVTEGPDGYPLIHARYLGESREFTATQVFGMM 600
Query: 130 LSKMREIAEAFLQTPVKNAVISVPAYFNDSQRRATKDAGAIAGLNVMQIINEPTAAALAY 189
LS ++EIA+ L V + I +P YF D QRR+ DA IAGL+ + +I+E TA ALAY
Sbjct: 601 LSNLKEIAQKNLNAAVVDCCIGIPVYFTDLQRRSVLDAATIAGLHPLHLIHETTATALAY 780
Query: 190 GLQKRANCVDK-RNIFIFDLGGGTFDVSILTVKNNVFEVKATAGDTHLGGQDFDNIMVDH 248
G+ K ++ N+ D+G + V I K V + + D LGG+DFD + H
Sbjct: 781 GIYKTDLPENEWLNVAFVDVGHASMQVCIAGFKKGQLHVLSHSYDRSLGGRDFDEALFHH 960
Query: 249 FVKEFRNKYKEDISENSKALRRLRTACEKAKRTLSYDTDATIELDAIYKGIDFCSSITRA 308
F +F+ +YK D+ +N++A RLR ACEK K+ LS + +A + ++ + D I R
Sbjct: 961 FAAKFKEEYKIDVYQNARACLRLRAACEKLKKVLSANPEAPLNIECLMDEKDVRGFIKRD 1140
Query: 309 KFEEMNMNLFEKCMNTVNICLADAKMDKNSVDDVVLVGGSSRIPKVRQLLQEVFKGKELC 368
FE++++ + E+ + LA+A + ++ V +VG SR+P + ++L E FK KE
Sbjct: 1141DFEQLSLPILERVKGPLEKALAEAGLTVENIHMVEVVGSGSRVPAINKILTEFFK-KEPR 1317
Query: 369 KSINPDEAVAYGAVVQAALLSEGSKNVPNLVLRDVTPLSFGV-----------SELGNLM 417
+++N E VA GA +Q A+LS K V + + P S + SE N
Sbjct: 1318RTMNASECVARGAALQCAILSPTFK-VREFQVNESFPFSVSLSWKYSGSDAPDSESDNKQ 1494
Query: 418 S-VMIPRNTSIP----VTVEKTKTY---YECYD 442
S ++ P+ IP +T +T T+ +C+D
Sbjct: 1495STIVFPKGNPIPSSKVLTFFRTGTFSVDVQCHD 1593
>TC85391 homologue to PIR|T06598|T06598 dnaK-type molecular chaperone BiP-A
- soybean, partial (52%)
Length = 1231
Score = 243 bits (620), Expect = 2e-64
Identities = 134/323 (41%), Positives = 209/323 (64%), Gaps = 4/323 (1%)
Frame = +1
Query: 298 GIDFCSSITRAKFEEMNMNLFEKCMNTVNICLADAKMDKNSVDDVVLVGGSSRIPKVRQL 357
G+ F +TRA+FEE+N +LF K M V + DA + KN +D++VLVGGS+RIPKV+QL
Sbjct: 4 GVAFSEPLTRARFEELNNDLFRKTMGPVKKAMDDAGLQKNQIDEIVLVGGSTRIPKVQQL 183
Query: 358 LQEVFKGKELCKSINPDEAVAYGAVVQAALLS-EGSKNVPNLVLRDVTPLSFGVSELGNL 416
L++ F GKE K +NPDEAVA+GA VQ ++LS EG +++L DV PL+ G+ +G +
Sbjct: 184 LKDYFDGKEPNKGVNPDEAVAFGAAVQGSILSGEGGAETKDILLLDVAPLTLGIETVGGV 363
Query: 417 MSVMIPRNTSIPVTVEKTKTYYECYDYYNKYSVTVKVYQGERITAYDNKLLGLFNFS-VP 475
M+ +IPRNT IP + T Y+ + +V+++V++GER D +LLG F+ S +P
Sbjct: 364 MTKLIPRNTVIPTKKSQVFTTYQ----DQQTTVSIQVFEGERSLTKDCRLLGKFDLSGIP 531
Query: 476 PIKGQLTPRG-HLIHLSFSIDVDGILNVSAKEETSGYKKGIAMTNQYCRLSTQEIARMIQ 534
P PRG I ++F +D +GILNV A+++ +G + I +TN+ RLS +EI RM++
Sbjct: 532 P-----APRGTPQIEVTFEVDANGILNVKAEDKGTGKSEKITITNEKGRLSQEEIERMVR 696
Query: 535 EAEIFKAQDMEFKKKVTAINALDDYLHNARKLMKD-DGVGSKLTPVDKEKINSAMIKGES 593
EAE F +D + K+++ A NAL+ Y++N + + D D + KL +KEKI +A+ +
Sbjct: 697 EAEEFAEEDKKVKERIDARNALETYVYNMKNQVSDKDKLADKLESDEKEKIETAVKEALE 876
Query: 594 LIDGNQQEDMSVFVDLLKELESI 616
+D NQ + + + LKE+E++
Sbjct: 877 WLDDNQSVEKEEYEEKLKEVEAV 945
>AW559598 weakly similar to SP|P09189|HS7C Heat shock cognate 70 kDa protein.
[Petunia] {Petunia hybrida}, partial (24%)
Length = 532
Score = 236 bits (601), Expect = 3e-62
Identities = 112/167 (67%), Positives = 138/167 (82%)
Frame = +1
Query: 1 MEEKYEGVAIGIDLGTTYSCVGVWHEQNDRVEIIHNDQGNRTTPSCVAFTNSQRLIGDAA 60
M E A+GIDLGTTYSCV VW E+ +RVEIIHND+G++ TPS VAFT+ QRL+G AA
Sbjct: 31 MANNCECYAVGIDLGTTYSCVAVWLEEKNRVEIIHNDEGSKITPSFVAFTDGQRLVGAAA 210
Query: 61 KNQAATNPSNTIFDTKRLIGRKYSDSIVQKDCLLWPFKVIAGTNDKPTIVVNYKGEEKRF 120
K+QAA NP NT+FD KRLIGRK+SD +VQKD LLWPFKV++G NDKP I + +KG+EK
Sbjct: 211 KDQAAINPQNTVFDAKRLIGRKFSDPVVQKDILLWPFKVVSGVNDKPMISLKFKGQEKLL 390
Query: 121 VAEEISSVILSKMREIAEAFLQTPVKNAVISVPAYFNDSQRRATKDA 167
AEEISS++L+ MREIAE +L++PVKNA I+VPAYFND+QR+AT DA
Sbjct: 391 CAEEISSIVLTNMREIAEMYLESPVKNAGITVPAYFNDAQRKATIDA 531
>TC80997 similar to PIR|B84729|B84729 70kD heat shock protein [imported] -
Arabidopsis thaliana, partial (91%)
Length = 1717
Score = 125 bits (314), Expect(2) = 4e-54
Identities = 83/242 (34%), Positives = 125/242 (51%), Gaps = 2/242 (0%)
Frame = +1
Query: 263 ENSKALRRLRTACEKAKRTLSYDTDATIELDAIYKGIDFCSSITRAKFEEMNMNLFEKCM 322
E K++ LR A + A LS + +++D + + C + RA+FEE+N +FEKC
Sbjct: 763 EEIKSMALLRVATQDAIHQLSTQSSVQVDVD-LGNSLKICKVVDRAEFEEVNKEVFEKCE 939
Query: 323 NTVNICLADAKMDKNSVDDVVLVGGSSRIPKVRQLLQEVFKGKELCKSINPDEAVAYGAV 382
+ + CL DAK++ ++DV+LVGG S IP+V L+ + K E+ K INP E GA
Sbjct: 940 SLIIQCLHDAKVEVGDINDVILVGGCSYIPRVENLVTNLCKITEVYKGINPLEGAVCGAT 1119
Query: 383 VQAALLSEGSKNVPNLVLRDV--TPLSFGVSELGNLMSVMIPRNTSIPVTVEKTKTYYEC 440
++ A+ S S NL L + TPL+ G+ GN +IPRNT++P K +
Sbjct: 1120MEGAVASGISDPFGNLDLLTIQATPLAIGIRADGNKFVPVIPRNTTMP--ARKDLLFTTI 1293
Query: 441 YDYYNKYSVTVKVYQGERITAYDNKLLGLFNFSVPPIKGQLTPRGHLIHLSFSIDVDGIL 500
+D N+ + VY+GE A +N LLG F P + P I + ID +L
Sbjct: 1294HD--NQTEALILVYEGEGKKAEENHLLGYFKIMGIPAAPKGVPE---ISVCMDIDAANVL 1458
Query: 501 NV 502
V
Sbjct: 1459RV 1464
Score = 105 bits (261), Expect(2) = 4e-54
Identities = 76/253 (30%), Positives = 129/253 (50%), Gaps = 9/253 (3%)
Frame = +3
Query: 15 GTTYSCVGVWHEQNDRVEIIHNDQGNRTTPSCVAFTNSQRLIGDAAK--NQAATNPSNTI 72
GT+ V VW+ VE++ N + + S V F + G ++ ++ +TI
Sbjct: 3 GTSQCSVAVWN--GSEVELLKNKRNQKLMKSFVTFKDEAPSGGVTSQFSHEHEMLFGDTI 176
Query: 73 FDTKRLIGRKYSDSIVQKDCLLWPFKV-IAGTNDKPTIVVNYKGEEKRFVAEEISSVILS 131
F+ KRLIGR +D +V L PF V +P I + EE+ ++ L
Sbjct: 177 FNMKRLIGRVDTDPVVHASKNL-PFLVQTLDIGVRPFIAALVNNVWRSTTPEEVLAMFLV 353
Query: 132 KMREIAEAFLQTPVKNAVISVPAYFNDSQRRATKDAGAIAGLNVMQIINEPTAAALAYGL 191
++R + E L+ P++N V++VP F+ Q + A A+AGL+V++++ EPTA AL YG
Sbjct: 354 ELRLMTETHLKRPIRNVVLTVPVSFSRFQLTRLERACAMAGLHVLRLMPEPTAVALLYGQ 533
Query: 192 QKRANCVD------KRNIFIFDLGGGTFDVSILTVKNNVFEVKATAGDTHLGGQDFDNIM 245
Q++ + ++ IF++G G DV++ V ++KA AG T +GG+D M
Sbjct: 534 QQQKASQETMGSGSEKIALIFNMGAGYCDVAVTATAGGVSQIKALAGST-IGGEDLLQNM 710
Query: 246 VDHFVKEFRNKYK 258
+ H + + N +K
Sbjct: 711 MRHLLPDSENIFK 749
>TC85395 homologue to GP|6911553|emb|CAB72130.1 heat shock protein 70
{Cucumis sativus}, partial (35%)
Length = 723
Score = 181 bits (459), Expect = 8e-46
Identities = 105/233 (45%), Positives = 148/233 (63%), Gaps = 2/233 (0%)
Frame = +3
Query: 397 NLVLRDVTPLSFGVSELGNLMSVMIPRNTSIPVTVEKTKTYYECYDYYNKYSVTVKVYQG 456
+L+L DVTPLS G+ G +M+V+IPRNT+IP E+ + Y N+ V ++VY+G
Sbjct: 6 DLLLLDVTPLSLGLETAGGVMTVLIPRNTTIPTKKEQVFSTYSD----NQPGVLIQVYEG 173
Query: 457 ERITAYDNKLLGLFNFS-VPPIKGQLTPRG-HLIHLSFSIDVDGILNVSAKEETSGYKKG 514
ER DN LLG F S +PP PRG I + F ID +GILNVSA+++T+G K
Sbjct: 174 ERARTRDNNLLGKFELSGIPP-----APRGVPQITVCFDIDANGILNVSAEDKTTGQKNK 338
Query: 515 IAMTNQYCRLSTQEIARMIQEAEIFKAQDMEFKKKVTAINALDDYLHNARKLMKDDGVGS 574
I +TN RLS +EI +M++EAE +KA+D E KKKV A NAL++Y +N R +KD+ +GS
Sbjct: 339 ITITNDKGRLSKEEIEKMVEEAEKYKAEDEEHKKKVDAKNALENYAYNMRNTIKDEKIGS 518
Query: 575 KLTPVDKEKINSAMIKGESLIDGNQQEDMSVFVDLLKELESIFESSKNKINKG 627
KL+P DK+KI+ A+ +D NQ + F D +KELES+ K+ +G
Sbjct: 519 KLSPEDKKKIDDAIDAAIQWLDSNQLAEADEFQDKMKELESLCNPIIAKMYQG 677
>BQ752418 homologue to SP|P78695|GR78 78 kDa glucose-regulated protein
homolog precursor (GRP 78), partial (47%)
Length = 1032
Score = 162 bits (409), Expect(3) = 2e-45
Identities = 97/251 (38%), Positives = 151/251 (59%), Gaps = 3/251 (1%)
Frame = -1
Query: 351 IPKVRQLLQEVFKGKELCKSINPDEAVAYGAVVQAALLSEGSKNVPNLVLRDVTPLSFGV 410
IPKV+ L++E F GK+ K INPDEAVA+GA VQA +LS G + +VL DV PL+ G+
Sbjct: 900 IPKVQSLIEEYFGGKKASKGINPDEAVAFGAAVQAGVLS-GEEGTEEIVLMDVNPLTLGI 724
Query: 411 SELGNLMSVMIPRNTSIPVTVEKTKTYYECYDYYNKYSVTVKVYQGERITAYDNKLLGLF 470
G +M+ +I RNT IP K++ + D N+ V ++V++GER DN LG F
Sbjct: 723 ETTGGVMTKLITRNTPIPT--RKSQIFSTAAD--NQPVVLIQVFEGERSLTKDNNQLGKF 556
Query: 471 NFS-VPPIKGQLTPRG-HLIHLSFSIDVDGILNVSAKEETSGYKKGIAMTNQYCRLSTQE 528
+ +PP PRG I +SF +D +GIL VSA ++ +G ++ I +TN RL+ +E
Sbjct: 555 ELTGIPP-----APRGVPQIEVSFELDANGILKVSAHDKGTGKQESITITNDKGRLTKEE 391
Query: 529 IARMIQEAEIFKAQDMEFKKKVTAINALDDYLHNARKLMKD-DGVGSKLTPVDKEKINSA 587
I RM++EAE + +D ++++ A N L++Y + + + D +G+G K+ DKE I A
Sbjct: 390 IDRMVEEAEKYAEEDKATRERIEARNGLENYAFSLKNQVNDEEGLGGKIDDEDKETILEA 211
Query: 588 MIKGESLIDGN 598
+ + ++ N
Sbjct: 210 VKETNDWLEEN 178
Score = 36.6 bits (83), Expect(3) = 2e-45
Identities = 16/25 (64%), Positives = 20/25 (80%)
Frame = -3
Query: 329 LADAKMDKNSVDDVVLVGGSSRIPK 353
L DAK+ K VDD+VLVGGS+R P+
Sbjct: 967 LKDAKVKKEDVDDIVLVGGSTRYPQ 893
Score = 23.9 bits (50), Expect(3) = 2e-45
Identities = 9/15 (60%), Positives = 13/15 (86%)
Frame = -2
Query: 308 AKFEEMNMNLFEKCM 322
AKFEE+NM+L +K +
Sbjct: 1031 AKFEELNMDLLKKTL 987
>CA859549 similar to GP|20260807|gb Hsp70 protein 1 {Rhizopus stolonifer},
partial (19%)
Length = 527
Score = 172 bits (437), Expect = 3e-43
Identities = 89/127 (70%), Positives = 101/127 (79%)
Frame = +3
Query: 6 EGVAIGIDLGTTYSCVGVWHEQNDRVEIIHNDQGNRTTPSCVAFTNSQRLIGDAAKNQAA 65
EG A+GIDLGTTYSCVGVW QNDRVEII NDQGNRTTPS VAFT+++RLIGDAAKNQ A
Sbjct: 156 EGKAVGIDLGTTYSCVGVW--QNDRVEIIANDQGNRTTPSYVAFTDTERLIGDAAKNQVA 329
Query: 66 TNPSNTIFDTKRLIGRKYSDSIVQKDCLLWPFKVIAGTNDKPTIVVNYKGEEKRFVAEEI 125
NP NT+FD KRLIGR+++D VQ D WPFKVI KP I V YKGE K+F EEI
Sbjct: 330 MNPYNTVFDAKRLIGRRFADQEVQSDMKHWPFKVI-DKAAKPYIQVEYKGETKQFTPEEI 506
Query: 126 SSVILSK 132
SS++L+K
Sbjct: 507 SSMVLTK 527
Database: MTGI
Posted date: Oct 22, 2004 3:39 PM
Number of letters in database: 27,044,181
Number of sequences in database: 36,976
Lambda K H
0.317 0.134 0.377
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 16,067,442
Number of Sequences: 36976
Number of extensions: 189427
Number of successful extensions: 1006
Number of sequences better than 10.0: 69
Number of HSP's better than 10.0 without gapping: 923
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 927
length of query: 647
length of database: 9,014,727
effective HSP length: 102
effective length of query: 545
effective length of database: 5,243,175
effective search space: 2857530375
effective search space used: 2857530375
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.3 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.6 bits)
S2: 61 (28.1 bits)
Medicago: description of AC137996.18


