

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
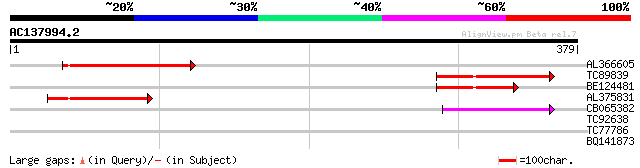
Query= AC137994.2 + phase: 0 /pseudo
(379 letters)
Database: MTGI
36,976 sequences; 27,044,181 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
AL366605 87 1e-17
TC89839 homologue to SP|P23466|CYAA_SACKL Adenylate cyclase (EC ... 84 1e-16
BE124481 64 7e-11
AL375831 64 7e-11
CB065382 similar to PIR|T09671|T096 RPE15 protein - alfalfa (fra... 64 1e-10
TC92638 similar to GP|23463525|gb|AAN33391.1 conserved hypotheti... 25 0.92
TC77786 28 5.8
BQ141873 similar to SP|P33485|VNUA Probable nuclear antigen. [st... 27 9.9
>AL366605
Length = 422
Score = 87.0 bits (214), Expect = 1e-17
Identities = 45/89 (50%), Positives = 55/89 (61%)
Frame = -3
Query: 36 EMKALHRKELFGINAYDLCLVSNVVIPPKFKVPDFEKYKGNTFPKIHLVMYVRKMSYQAD 95
E+KA +R DLCLVS V +P KFK+PDF++Y G T P+ H++ YVRKM D
Sbjct: 417 EIKA-NRGNADSFKTQDLCLVSKVDVPKKFKIPDFDRYNGLTCPQNHIIKYVRKMGNYKD 241
Query: 96 NDELFIHYFQDSLTGTALLGYMGLDKSDI 124
ND L IH FQDSL A Y L K+DI
Sbjct: 240 NDSLMIHCFQDSLMEDAAEWYTSLSKNDI 154
>TC89839 homologue to SP|P23466|CYAA_SACKL Adenylate cyclase (EC 4.6.1.1)
(ATP pyrophosphate-lyase) (Adenylyl cyclase). [Yeast],
partial (0%)
Length = 665
Score = 83.6 bits (205), Expect = 1e-16
Identities = 44/80 (55%), Positives = 54/80 (67%), Gaps = 1/80 (1%)
Frame = +3
Query: 286 TQIDLIPMKYADLFPILLERNLVHTKAPLP-VPARLSARYRPNLFCAFHQGASCHDIEHC 344
+Q D IP+KYADL PILL++NL+ T PLP VP L YRP+L C FHQGA HD E C
Sbjct: 369 SQCDPIPVKYADLLPILLKKNLIQT-LPLPRVPNSLPPWYRPDLNCVFHQGAPGHDTEQC 545
Query: 345 FSLKKIVQNLIPENLLPLED 364
+ LK+ VQ LI N+ +D
Sbjct: 546 YPLKEEVQKLIENNVWSFDD 605
>BE124481
Length = 538
Score = 64.3 bits (155), Expect = 7e-11
Identities = 34/56 (60%), Positives = 40/56 (70%), Gaps = 1/56 (1%)
Frame = +3
Query: 286 TQIDLIPMKYADLFPILLERNLVHTKAPLP-VPARLSARYRPNLFCAFHQGASCHD 340
+Q D IP+KYADL PILL++NL+ T PLP VP L YRP+L C FHQGA HD
Sbjct: 372 SQCDPIPVKYADLLPILLKKNLIQT-LPLPRVPNSLPPWYRPDLNCVFHQGAPGHD 536
>AL375831
Length = 467
Score = 64.3 bits (155), Expect = 7e-11
Identities = 30/70 (42%), Positives = 45/70 (63%)
Frame = +2
Query: 26 LQEKYNEIQREMKALHRKELFGINAYDLCLVSNVVIPPKFKVPDFEKYKGNTFPKIHLVM 85
++E E++ E++A +R + DLCLVS V +P KFK+PDF++Y G T P+ H++
Sbjct: 158 MEELAKELRHEIQA-NRGNADSVKTQDLCLVSKVDVPKKFKIPDFDRYNGLTCPQNHIIK 334
Query: 86 YVRKMSYQAD 95
YVRKM D
Sbjct: 335 YVRKMGNYKD 364
>CB065382 similar to PIR|T09671|T096 RPE15 protein - alfalfa (fragment),
partial (9%)
Length = 624
Score = 63.5 bits (153), Expect = 1e-10
Identities = 34/75 (45%), Positives = 44/75 (58%)
Frame = -2
Query: 290 LIPMKYADLFPILLERNLVHTKAPLPVPARLSARYRPNLFCAFHQGASCHDIEHCFSLKK 349
LIPM YA+ P LL R + P P L R+R +L C FHQGA HD+E C++LK
Sbjct: 470 LIPMLYAE*LPTLLLRGHCTIRQGKPPPDPLPPRFRSDLKCDFHQGALGHDVEGCYALKH 291
Query: 350 IVQNLIPENLLPLED 364
IV+ LI + L E+
Sbjct: 290 IVKKLINQGKLTFEN 246
>TC92638 similar to GP|23463525|gb|AAN33391.1 conserved hypothetical protein
{Brucella suis 1330}, partial (6%)
Length = 612
Score = 25.0 bits (53), Expect(2) = 0.92
Identities = 20/56 (35%), Positives = 26/56 (45%), Gaps = 2/56 (3%)
Frame = +2
Query: 302 LLERNLVHTKAPLPVPARLSARYRPNLFCAFHQGAS--CHDIEHCFSLKKIVQNLI 355
L E+NLV T+AP R PN F AS C I+H F+ ++ N I
Sbjct: 296 LFEKNLV*TRAP---------RRMPN*SIYFFSIASIFCFKIDHWFNYCLVMYNTI 436
Score = 24.3 bits (51), Expect(2) = 0.92
Identities = 9/13 (69%), Positives = 11/13 (84%)
Frame = +1
Query: 289 DLIPMKYADLFPI 301
D IPMKYA+L P+
Sbjct: 262 DPIPMKYAELLPV 300
>TC77786
Length = 1094
Score = 28.1 bits (61), Expect = 5.8
Identities = 10/26 (38%), Positives = 17/26 (64%)
Frame = +3
Query: 330 CAFHQGASCHDIEHCFSLKKIVQNLI 355
C +H+ HD E CF L++ +++LI
Sbjct: 825 CCYHRSRG-HDTEECFQLREAIEDLI 899
>BQ141873 similar to SP|P33485|VNUA Probable nuclear antigen. [strain Kaplan
PRV] {Pseudorabies virus}, partial (2%)
Length = 1350
Score = 27.3 bits (59), Expect = 9.9
Identities = 13/38 (34%), Positives = 23/38 (60%)
Frame = -3
Query: 317 PARLSARYRPNLFCAFHQGASCHDIEHCFSLKKIVQNL 354
P R++ +R +L CAFH S + C +++ +V+NL
Sbjct: 1279 PPRITP*HRASLLCAFH---SASHLLSCRTIRVLVRNL 1175
Database: MTGI
Posted date: Oct 22, 2004 3:39 PM
Number of letters in database: 27,044,181
Number of sequences in database: 36,976
Lambda K H
0.340 0.147 0.501
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 14,013,147
Number of Sequences: 36976
Number of extensions: 225826
Number of successful extensions: 1615
Number of sequences better than 10.0: 17
Number of HSP's better than 10.0 without gapping: 1595
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 1614
length of query: 379
length of database: 9,014,727
effective HSP length: 98
effective length of query: 281
effective length of database: 5,391,079
effective search space: 1514893199
effective search space used: 1514893199
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 15 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 39 (21.9 bits)
S2: 59 (27.3 bits)
Medicago: description of AC137994.2


