

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
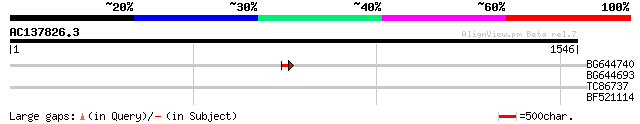
Query= AC137826.3 - phase: 0 /pseudo
(1546 letters)
Database: MTGI
36,976 sequences; 27,044,181 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
BG644740 similar to PIR|A84460|A84 probable retroelement pol pol... 47 6e-05
BG644693 weakly similar to GP|18767374|g Putative 22 kDa kafirin... 37 0.002
TC86737 weakly similar to GP|6683624|dbj|BAA89272.1 Pol {Alterna... 33 1.1
BF521114 similar to PIR|T45723|T457 hypothetical protein F1P2.18... 30 7.3
>BG644740 similar to PIR|A84460|A84 probable retroelement pol polyprotein
[imported] - Arabidopsis thaliana, partial (4%)
Length = 754
Score = 47.0 bits (110), Expect = 6e-05
Identities = 20/34 (58%), Positives = 26/34 (75%)
Frame = -1
Query: 740 DSSMIMCKDYRQLNRVTTRNKYTLPLIDDLFNQL 773
D MC DYRQ N+VTT+NKY LP ID+LF+++
Sbjct: 211 DGYFRMCIDYRQFNKVTTKNKYPLPRIDNLFDKI 110
>BG644693 weakly similar to GP|18767374|g Putative 22 kDa kafirin cluster;
Ty3-Gypsy type {Oryza sativa}, partial (15%)
Length = 716
Score = 37.4 bits (85), Expect(2) = 0.002
Identities = 18/34 (52%), Positives = 22/34 (63%)
Frame = +2
Query: 740 DSSMIMCKDYRQLNRVTTRNKYTLPLIDDLFNQL 773
D + M DY QLN V + KY LPLID+LF+ L
Sbjct: 212 DGFLRMSIDYPQLNNVNIKIKYPLPLIDELFDNL 313
Score = 23.9 bits (50), Expect(2) = 0.002
Identities = 9/21 (42%), Positives = 14/21 (65%)
Frame = +1
Query: 806 LWAL*VSSDIVWVDKCSFGCH 826
+W+L* SD+ WV+K G +
Sbjct: 412 IWSL*NPSDVFWVNKSPDGIY 474
>TC86737 weakly similar to GP|6683624|dbj|BAA89272.1 Pol {Alternaria
alternata}, partial (21%)
Length = 1540
Score = 32.7 bits (73), Expect = 1.1
Identities = 12/28 (42%), Positives = 19/28 (67%)
Frame = +1
Query: 746 CKDYRQLNRVTTRNKYTLPLIDDLFNQL 773
C DYR LN +T +++Y LPLI + ++
Sbjct: 622 CVDYRALNAITKKDRYPLPLISETLRRV 705
>BF521114 similar to PIR|T45723|T457 hypothetical protein F1P2.180 -
Arabidopsis thaliana, partial (24%)
Length = 421
Score = 30.0 bits (66), Expect = 7.3
Identities = 11/27 (40%), Positives = 18/27 (65%)
Frame = -3
Query: 171 GSESEDTFKFILHYYERLNKLVIVKQY 197
G ESED F+ ++HYY K++++ Y
Sbjct: 197 GEESEDNFESLVHYYPYRYKMMVMPTY 117
Database: MTGI
Posted date: Oct 22, 2004 3:39 PM
Number of letters in database: 27,044,181
Number of sequences in database: 36,976
Lambda K H
0.359 0.161 0.583
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 52,228,474
Number of Sequences: 36976
Number of extensions: 811056
Number of successful extensions: 7389
Number of sequences better than 10.0: 8
Number of HSP's better than 10.0 without gapping: 2120
Number of HSP's successfully gapped in prelim test: 289
Number of HSP's that attempted gapping in prelim test: 5128
Number of HSP's gapped (non-prelim): 2716
length of query: 1546
length of database: 9,014,727
effective HSP length: 109
effective length of query: 1437
effective length of database: 4,984,343
effective search space: 7162500891
effective search space used: 7162500891
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 14 ( 7.2 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 37 (21.8 bits)
S2: 65 (29.6 bits)
Medicago: description of AC137826.3


