

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC137823.16 + phase: 0
(621 letters)
Database: MTGI
36,976 sequences; 27,044,181 total letters
Searching..................................................done
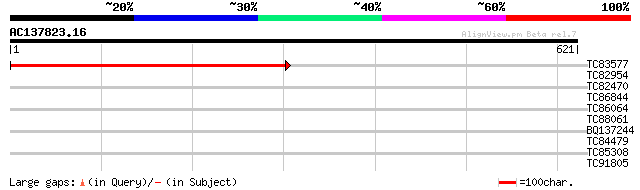
Score E
Sequences producing significant alignments: (bits) Value
TC83577 similar to GP|9758065|dbj|BAB08644.1 gene_id:K15E6.9~unk... 600 e-172
TC82954 weakly similar to GP|8809591|dbj|BAA97142.1 emb|CAB82814... 32 0.56
TC82470 homologue to GP|23497460|gb|AAN37003.1 hypothetical prot... 32 0.56
TC86844 similar to GP|16797791|gb|AAL27150.1 bZIP transcription ... 30 2.8
TC86064 similar to SP|P54767|DCE_LYCES Glutamate decarboxylase (... 30 2.8
TC88061 homologue to SP|Q43117|KPYA_RICCO Pyruvate kinase isozym... 30 3.6
BQ137244 similar to GP|20161222|dbj Epstein-Barr virus EBNA-1-li... 29 6.1
TC84479 ENBP1 29 6.1
TC85308 homologue to SP|O65735|ALF_CICAR Fructose-bisphosphate a... 28 8.0
TC91805 weakly similar to GP|9758374|dbj|BAB08823.1 receptor-lik... 28 8.0
>TC83577 similar to GP|9758065|dbj|BAB08644.1 gene_id:K15E6.9~unknown
protein {Arabidopsis thaliana}, partial (34%)
Length = 1060
Score = 600 bits (1547), Expect = e-172
Identities = 307/307 (100%), Positives = 307/307 (100%)
Frame = +1
Query: 1 MLEAYDRQCDEASRIFAEYHKRLCYYINQARDAQRSGVDSSVEMVNSFSAKNEKEAVYST 60
MLEAYDRQCDEASRIFAEYHKRLCYYINQARDAQRSGVDSSVEMVNSFSAKNEKEAVYST
Sbjct: 139 MLEAYDRQCDEASRIFAEYHKRLCYYINQARDAQRSGVDSSVEMVNSFSAKNEKEAVYST 318
Query: 61 VKGSKSSDDVIVIETTREKNIRKACESLVAYMVDKIQSSFPAYEGSGVLSNPQAEAAKLG 120
VKGSKSSDDVIVIETTREKNIRKACESLVAYMVDKIQSSFPAYEGSGVLSNPQAEAAKLG
Sbjct: 319 VKGSKSSDDVIVIETTREKNIRKACESLVAYMVDKIQSSFPAYEGSGVLSNPQAEAAKLG 498
Query: 121 FDFDGQIPDEVRTVIVNCLKSPPLLLQAITAYTSHLKSQISREIEKIDVRADAEILRYKY 180
FDFDGQIPDEVRTVIVNCLKSPPLLLQAITAYTSHLKSQISREIEKIDVRADAEILRYKY
Sbjct: 499 FDFDGQIPDEVRTVIVNCLKSPPLLLQAITAYTSHLKSQISREIEKIDVRADAEILRYKY 678
Query: 181 ENNIVMDVSSSDGSSPLQYPLYGNGKLGADVPPGGSQNQLLERQKAHVQQFLATEDALNN 240
ENNIVMDVSSSDGSSPLQYPLYGNGKLGADVPPGGSQNQLLERQKAHVQQFLATEDALNN
Sbjct: 679 ENNIVMDVSSSDGSSPLQYPLYGNGKLGADVPPGGSQNQLLERQKAHVQQFLATEDALNN 858
Query: 241 AAEARDLCEKLMKRLHGGTDVTSRSIGIGATSQNVGSLRQLQLDVWAKEREVSGLKASLN 300
AAEARDLCEKLMKRLHGGTDVTSRSIGIGATSQNVGSLRQLQLDVWAKEREVSGLKASLN
Sbjct: 859 AAEARDLCEKLMKRLHGGTDVTSRSIGIGATSQNVGSLRQLQLDVWAKEREVSGLKASLN 1038
Query: 301 TLMSEIQ 307
TLMSEIQ
Sbjct: 1039TLMSEIQ 1059
>TC82954 weakly similar to GP|8809591|dbj|BAA97142.1
emb|CAB82814.1~gene_id:MNB8.8~similar to unknown protein
{Arabidopsis thaliana}, partial (7%)
Length = 783
Score = 32.3 bits (72), Expect = 0.56
Identities = 33/129 (25%), Positives = 51/129 (38%), Gaps = 8/129 (6%)
Frame = +3
Query: 249 EKLMKRLHGGTDVTSRSIGIGATSQNVGSLRQLQLDVWAKEREVSGLKASLNTLMSEIQR 308
E L K H + +GI A + W ER L+ + L++E++
Sbjct: 315 ENLKKIRHEDAKANEKVVGIFAAQEQS----------WFSERRK--LRQQIGALLNELRV 458
Query: 309 LNK--------LCAERKEAEDSLKKKWKKIEEFDGRRSELESIYTALLKANTDAASFWSQ 360
K L + KE E +++K KKIEE + +R ELE KA DA
Sbjct: 459 FEKKRDLAISDLNQKLKEMEGLVEEKDKKIEEEEKKRKELEE---KAKKAEKDAEELRES 629
Query: 361 QPSTAREYA 369
+E++
Sbjct: 630 SKREGQEHS 656
>TC82470 homologue to GP|23497460|gb|AAN37003.1 hypothetical protein
{Plasmodium falciparum 3D7}, partial (8%)
Length = 1064
Score = 32.3 bits (72), Expect = 0.56
Identities = 38/152 (25%), Positives = 75/152 (49%), Gaps = 25/152 (16%)
Frame = +2
Query: 216 SQNQLLERQKAHVQQFL--ATEDALNNAAEARDLCE--KLMKRLHGGTD-----VTSRSI 266
S+ Q+ ER+ +++ + ED N E RD+ + K++ G D ++
Sbjct: 167 SKTQVEERRLQSIKRDIEECCEDLENKKKEIRDVGRIIEARKKMQGKIDECVKDFVAKEG 346
Query: 267 GIGATSQNVGS-----------LRQLQLDVWAKEREV-SGLKASLNTLMSEIQR----LN 310
+G +G LRQ+ +D +K++E+ S +K +N L+S+ + +
Sbjct: 347 QLGLMEDLIGEHKKELKTKELELRQV-MDNISKQKELESQVKELVNDLVSKQKHFESHIK 523
Query: 311 KLCAERKEAEDSLKKKWKKIEEFDGRRSELES 342
+L ++ ++ E LK+ + +EF+GR +ELES
Sbjct: 524 ELESKERQLEGRLKEHELEEKEFEGRMNELES 619
>TC86844 similar to GP|16797791|gb|AAL27150.1 bZIP transcription factor
{Nicotiana tabacum}, partial (47%)
Length = 1609
Score = 30.0 bits (66), Expect = 2.8
Identities = 30/137 (21%), Positives = 60/137 (42%)
Frame = +1
Query: 304 SEIQRLNKLCAERKEAEDSLKKKWKKIEEFDGRRSELESIYTALLKANTDAASFWSQQPS 363
++ +R+ ++ + R+ A S ++K + E + + SEL ++LLK TD ++
Sbjct: 664 TDAKRVRRMLSNRESARRSRRRKQAHLTELETQVSELRGENSSLLKRLTDVTQKFNNSAV 843
Query: 364 TAREYALSTIIPACSAVVETSNSAKDLIEKEVSAFYRSPDNSLYMLPSSPQALLEAIGSS 423
R A VET + + E+ V F S N ++ S + ++G S
Sbjct: 844 DNR---------ILKADVETLRAKVKMAEETVKRFTGS--NPVFNAMSE----VSSMGMS 978
Query: 424 GSSGQEAVANAEISAAI 440
G + ++A+ S +
Sbjct: 979 LFDGSPSESSADASVPV 1029
>TC86064 similar to SP|P54767|DCE_LYCES Glutamate decarboxylase (EC 4.1.1.15)
(GAD) (ERT D1). [Tomato] {Lycopersicon esculentum},
partial (87%)
Length = 1759
Score = 30.0 bits (66), Expect = 2.8
Identities = 15/44 (34%), Positives = 25/44 (56%)
Frame = +1
Query: 135 IVNCLKSPPLLLQAITAYTSHLKSQISREIEKIDVRADAEILRY 178
+VN L + P + + A+ + + S+ S E++K AEILRY
Sbjct: 1399 VVNLLDTLPSSINSKDAHVAVIASETSEEVKKDITETQAEILRY 1530
>TC88061 homologue to SP|Q43117|KPYA_RICCO Pyruvate kinase isozyme A
chloroplast precursor (EC 2.7.1.40). [Castor bean]
{Ricinus communis}, partial (84%)
Length = 1765
Score = 29.6 bits (65), Expect = 3.6
Identities = 24/97 (24%), Positives = 43/97 (43%)
Frame = +3
Query: 116 AAKLGFDFDGQIPDEVRTVIVNCLKSPPLLLQAITAYTSHLKSQISREIEKIDVRADAEI 175
++K D D I + V + ++ +KS ++ +HLKS I+ D+ A+I
Sbjct: 897 SSKDWLDIDFGIAEGVDFIAISFVKSAEVI--------THLKSYIAARSRDSDISVIAKI 1052
Query: 176 LRYKYENNIVMDVSSSDGSSPLQYPLYGNGKLGADVP 212
N+ + +SDG+ + G LGA +P
Sbjct: 1053ESIDSLKNLEEIIQASDGA------MVARGDLGAQIP 1145
>BQ137244 similar to GP|20161222|dbj Epstein-Barr virus EBNA-1-like protein
{Oryza sativa (japonica cultivar-group)}, partial (5%)
Length = 1050
Score = 28.9 bits (63), Expect = 6.1
Identities = 31/92 (33%), Positives = 44/92 (47%), Gaps = 4/92 (4%)
Frame = -1
Query: 405 SLYMLPSSPQALLEAIGSSGSSGQEAVANAEISAAILTARAGARDPSAIPSICRVS-AAL 463
+LY + SP + + SS + + AVA+A ++ + ARA AR PS VS A L
Sbjct: 840 ALYCISRSPAFVPLSPLSSLLAARVAVASAAVARVVARARARARLPSLPVVSSHVSCAVL 661
Query: 464 QYAAGGLEGSDAGLASILESLEFCL---KRRG 492
AA A +S L S C+ +RRG
Sbjct: 660 SPAASFSLARIARASSRLRSSACCVAWSRRRG 565
>TC84479 ENBP1
Length = 5108
Score = 28.9 bits (63), Expect = 6.1
Identities = 33/118 (27%), Positives = 50/118 (41%), Gaps = 4/118 (3%)
Frame = +1
Query: 361 QPSTAREYALSTIIPACSAVVETSNSAKDLIEKEVSAFYRSPDNSLYMLPSSPQALLEAI 420
Q ST + ALSTI+P E+ + KD + KE + S ++L S + +E I
Sbjct: 2242 QKSTRVDGALSTIVPHKHIQEESISPLKDPVNKE--------EKSDFVLECSKDSGIEKI 2397
Query: 421 --GSSGSSGQEAVANAEISAAILTARAGARDPSAIPSIC--RVSAALQYAAGGLEGSD 474
G SG +E +LT ++D + C V A+ + LE SD
Sbjct: 2398 TKGLMSKSGDVHKRCSERLRTLLTDHKNSQDVEVEETFCENEVEEAIDHE---LESSD 2562
>TC85308 homologue to SP|O65735|ALF_CICAR Fructose-bisphosphate aldolase
cytoplasmic isozyme (EC 4.1.2.13). [Chickpea Garbanzo],
partial (98%)
Length = 1507
Score = 28.5 bits (62), Expect = 8.0
Identities = 19/69 (27%), Positives = 36/69 (51%), Gaps = 5/69 (7%)
Frame = -1
Query: 468 GGLEGSDAGLASILESLEFCLKRRGS-----EASVLEDLLKAINLVHIRRDLVQSGHALL 522
G L+ S+ L++ L +C++ RG+ E++ ED++ LV +DL+ + LL
Sbjct: 901 GSLQSSNGVLSNNLGCNLWCIRSRGNHVRLQESAFKEDMVVIQGLVACCKDLLGNSSTLL 722
Query: 523 NHAYFVQQD 531
N + +D
Sbjct: 721 NVMRSINKD 695
>TC91805 weakly similar to GP|9758374|dbj|BAB08823.1 receptor-like protein
kinase {Arabidopsis thaliana}, partial (5%)
Length = 587
Score = 28.5 bits (62), Expect = 8.0
Identities = 20/96 (20%), Positives = 41/96 (41%)
Frame = +3
Query: 123 FDGQIPDEVRTVIVNCLKSPPLLLQAITAYTSHLKSQISREIEKIDVRADAEILRYKYEN 182
F+G++P+ + C LQ +TAY +HL ++ + + +I + +
Sbjct: 75 FEGRLPENL------CYHGE---LQNLTAYENHLSGELPESLGNCSSLLEMKIYKNDFYG 227
Query: 183 NIVMDVSSSDGSSPLQYPLYGNGKLGADVPPGGSQN 218
NI + S+ L Y + + K ++P S +
Sbjct: 228 NIPSGLWRSEN---LGYFMISHNKFNGELPQNSSSS 326
Database: MTGI
Posted date: Oct 22, 2004 3:39 PM
Number of letters in database: 27,044,181
Number of sequences in database: 36,976
Lambda K H
0.314 0.129 0.361
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 16,425,047
Number of Sequences: 36976
Number of extensions: 196858
Number of successful extensions: 702
Number of sequences better than 10.0: 20
Number of HSP's better than 10.0 without gapping: 699
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 702
length of query: 621
length of database: 9,014,727
effective HSP length: 102
effective length of query: 519
effective length of database: 5,243,175
effective search space: 2721207825
effective search space used: 2721207825
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.2 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 42 (22.0 bits)
S2: 61 (28.1 bits)
Medicago: description of AC137823.16


