

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC137822.15 + phase: 0
(279 letters)
Database: MTGI
36,976 sequences; 27,044,181 total letters
Searching..................................................done
Score E
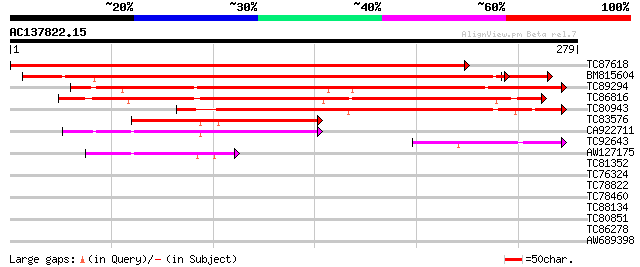
Sequences producing significant alignments: (bits) Value
TC87618 similar to GP|10176982|dbj|BAB10214. contains similarity... 478 e-135
BM815604 similar to GP|10176982|db contains similarity to unknow... 392 e-112
TC89294 similar to GP|10176982|dbj|BAB10214. contains similarity... 236 9e-63
TC86816 similar to GP|10176982|dbj|BAB10214. contains similarity... 218 2e-57
TC80943 similar to GP|15451150|gb|AAK96846.1 Unknown protein {Ar... 179 1e-45
TC83576 similar to GP|15293013|gb|AAK93617.1 unknown protein {Ar... 96 1e-20
CA922711 similar to GP|20197959|gb| expressed protein {Arabidops... 87 7e-18
TC92643 similar to GP|15293013|gb|AAK93617.1 unknown protein {Ar... 50 7e-07
AW127175 similar to GP|15293013|gb| unknown protein {Arabidopsis... 41 6e-04
TC81352 similar to GP|6633847|gb|AAF19706.1| F2K11.19 {Arabidops... 32 0.21
TC76324 similar to GP|7242813|emb|CAB77393.1 germin-like protein... 29 1.7
TC78822 similar to PIR|T02962|T02962 peroxidase (EC 1.11.1.7) is... 29 1.7
TC78460 similar to GP|21554579|gb|AAM63621.1 flavonol synthase-l... 29 2.3
TC88134 weakly similar to SP|Q9MB73|LGT_CITUN Limonoid UDP-gluco... 29 2.3
TC80851 weakly similar to GP|21644692|dbj|BAC01248. hypothetical... 27 6.6
TC86278 similar to GP|17473703|gb|AAL38307.1 germin-like protein... 27 8.7
AW689398 similar to GP|21593065|gb| NAM / CUC2-like protein {Ara... 27 8.7
>TC87618 similar to GP|10176982|dbj|BAB10214. contains similarity to unknown
protein~emb|CAB89322.1~gene_id:MYH19.8 {Arabidopsis
thaliana}, partial (46%)
Length = 1037
Score = 478 bits (1229), Expect = e-135
Identities = 226/226 (100%), Positives = 226/226 (100%)
Frame = +2
Query: 1 MVEQGRERVGHVNKVCYVKRVIAKKRNKPYNHRRVKKHVPKALQELFDSCKQTFKGPGTV 60
MVEQGRERVGHVNKVCYVKRVIAKKRNKPYNHRRVKKHVPKALQELFDSCKQTFKGPGTV
Sbjct: 359 MVEQGRERVGHVNKVCYVKRVIAKKRNKPYNHRRVKKHVPKALQELFDSCKQTFKGPGTV 538
Query: 61 PSPRDVHKLCHILDNMKPEDVGLSRDLQFFKPGNIIKENQRVTYTTVYKCDNFSLCIFFL 120
PSPRDVHKLCHILDNMKPEDVGLSRDLQFFKPGNIIKENQRVTYTTVYKCDNFSLCIFFL
Sbjct: 539 PSPRDVHKLCHILDNMKPEDVGLSRDLQFFKPGNIIKENQRVTYTTVYKCDNFSLCIFFL 718
Query: 121 PERGVIPLHNHPGMTVFSKLLLGQMHIKSYDWVDHEASHNLLQPSSKLRLAKLKANKTFT 180
PERGVIPLHNHPGMTVFSKLLLGQMHIKSYDWVDHEASHNLLQPSSKLRLAKLKANKTFT
Sbjct: 719 PERGVIPLHNHPGMTVFSKLLLGQMHIKSYDWVDHEASHNLLQPSSKLRLAKLKANKTFT 898
Query: 181 APCDTSVLYPTTGGNIHEFTAITPCAVLDVIGPPYSKEDGRDCSYY 226
APCDTSVLYPTTGGNIHEFTAITPCAVLDVIGPPYSKEDGRDCSYY
Sbjct: 899 APCDTSVLYPTTGGNIHEFTAITPCAVLDVIGPPYSKEDGRDCSYY 1036
>BM815604 similar to GP|10176982|db contains similarity to unknown
protein~emb|CAB89322.1~gene_id:MYH19.8 {Arabidopsis
thaliana}, partial (45%)
Length = 793
Score = 392 bits (1008), Expect(2) = e-112
Identities = 190/241 (78%), Positives = 214/241 (87%), Gaps = 1/241 (0%)
Frame = +2
Query: 7 ERVGHVNKVCYVKRVIAKKRNKPYNHRRVKKHVP-KALQELFDSCKQTFKGPGTVPSPRD 65
ERV +V KV Y+KRVI KKR KPY+ +VKK + KALQ+LF SCK+TFKGP TVPSP++
Sbjct: 2 ERVRNVEKVGYMKRVIVKKR-KPYHSNKVKKPMSNKALQKLFVSCKETFKGPNTVPSPQN 178
Query: 66 VHKLCHILDNMKPEDVGLSRDLQFFKPGNIIKENQRVTYTTVYKCDNFSLCIFFLPERGV 125
VHKLCHILDNMKPEDVGLS+DLQFFKP +I +EN RVTYTT+YKCDNFSLCIFFLP +GV
Sbjct: 179 VHKLCHILDNMKPEDVGLSKDLQFFKPESIFRENPRVTYTTIYKCDNFSLCIFFLPSKGV 358
Query: 126 IPLHNHPGMTVFSKLLLGQMHIKSYDWVDHEASHNLLQPSSKLRLAKLKANKTFTAPCDT 185
IPLHNHPGMTVFSKLLLGQMHIKSYDWVD + SHNLL+ S+LRLAKLKANK FT+PCDT
Sbjct: 359 IPLHNHPGMTVFSKLLLGQMHIKSYDWVDPDVSHNLLKQPSQLRLAKLKANKVFTSPCDT 538
Query: 186 SVLYPTTGGNIHEFTAITPCAVLDVIGPPYSKEDGRDCSYYKDYPYNAFPNEEKIGEVKD 245
SVLYP TGGNIHEFTAITPCAVLDVIGPPYSK+DGRDCSYY+D+ Y AFPN E I E+K+
Sbjct: 539 SVLYPKTGGNIHEFTAITPCAVLDVIGPPYSKDDGRDCSYYRDHLYTAFPNGE-IAELKE 715
Query: 246 K 246
+
Sbjct: 716 E 718
Score = 29.3 bits (64), Expect(2) = e-112
Identities = 12/25 (48%), Positives = 19/25 (76%)
Frame = +1
Query: 243 VKDKDDSYGLLEEIDMPENCQMDGI 267
V+ +++SYG LEEI+MPE + G+
Sbjct: 706 VERRNESYGWLEEIEMPETLKWMGL 780
>TC89294 similar to GP|10176982|dbj|BAB10214. contains similarity to unknown
protein~emb|CAB89322.1~gene_id:MYH19.8 {Arabidopsis
thaliana}, partial (82%)
Length = 838
Score = 236 bits (601), Expect = 9e-63
Identities = 124/250 (49%), Positives = 164/250 (65%), Gaps = 6/250 (2%)
Frame = +3
Query: 31 NHRRVKKHVPKALQELFDSCKQTF--KGPGTVPSPRDVHKLCHILDNMKPEDVGLSRDLQ 88
N RR KK P +Q+LF++CK+ F G G VP D+ KL +LD +KPEDV L ++
Sbjct: 6 NRRRQKKMPP--VQKLFETCKEVFASSGTGIVPPAEDIDKLRSVLDGIKPEDVDLDPNMP 179
Query: 89 FFKPGNIIKENQRVTYTTVYKCDNFSLCIFFLPERGVIPLHNHPGMTVFSKLLLGQMHIK 148
+F+ N ++TY +Y+C+ FS+ IF LP GVIPLHNHPGMTVFSKLL G MHIK
Sbjct: 180 YFR-ANASHRRPKITYLHIYECEKFSMGIFCLPPSGVIPLHNHPGMTVFSKLLFGTMHIK 356
Query: 149 SYDWVDH--EASHNLLQPSSK--LRLAKLKANKTFTAPCDTSVLYPTTGGNIHEFTAITP 204
SYDWV S + +PS LRLAK+K + FTAPC+ S+LYP GGN+H FTA+T
Sbjct: 357 SYDWVVDLPPESPTIFKPSESPDLRLAKVKVDDDFTAPCNPSILYPEDGGNLHCFTAVTA 536
Query: 205 CAVLDVIGPPYSKEDGRDCSYYKDYPYNAFPNEEKIGEVKDKDDSYGLLEEIDMPENCQM 264
CAVLDV+GPPYS DGR C+YY +YP++ F E + +++ Y L+E D E+ ++
Sbjct: 537 CAVLDVLGPPYSDFDGRHCTYYTNYPFSNF-QVEGLSIPEEERSVYEWLQEKDQLEDLKV 713
Query: 265 DGIEYLGPPI 274
+G Y GP I
Sbjct: 714 EGRMYSGPTI 743
>TC86816 similar to GP|10176982|dbj|BAB10214. contains similarity to unknown
protein~emb|CAB89322.1~gene_id:MYH19.8 {Arabidopsis
thaliana}, partial (71%)
Length = 1314
Score = 218 bits (556), Expect = 2e-57
Identities = 119/244 (48%), Positives = 158/244 (63%), Gaps = 4/244 (1%)
Frame = +3
Query: 25 KRNKPYNHRRVKKHVPKALQELFDSCKQTFKGP--GTVPSPRDVHKLCHILDNMKPEDVG 82
++N+ + RR + +Q+LF +CK F G VPS + + L +L +KPED+G
Sbjct: 363 RKNRRHLRRRTEM---TPVQKLFLACKHVFANAAHGIVPSSQHIEMLRSVLAGIKPEDLG 533
Query: 83 LSRDLQFFKPGNIIKENQRVTYTTVYKCDNFSLCIFFLPERGVIPLHNHPGMTVFSKLLL 142
L D+ +F NI ++TY +Y+C+ FS+ IF LP GVIPLHNHPGMTVFSKLL
Sbjct: 534 LKPDMPYFS--NINGGTPKITYLHIYECEKFSMGIFCLPPSGVIPLHNHPGMTVFSKLLF 707
Query: 143 GQMHIKSYDWV-DHEASHNLLQPSSKLRLAKLKANKTFTAPCDTSVLYPTTGGNIHEFTA 201
G MHIKSYDW D A + Q K RLAK+K + FTAPC+ S+LYP GGN+H FTA
Sbjct: 708 GTMHIKSYDWAGDLPADVSQTQIPEK-RLAKIKVDADFTAPCNPSILYPDDGGNMHCFTA 884
Query: 202 ITPCAVLDVIGPPYSKEDGRDCSYYKDYPYNAFPNEE-KIGEVKDKDDSYGLLEEIDMPE 260
+T CAVLDV+GPPYS DGR C+YY+ +P++ FP E I E + KD Y L+E + PE
Sbjct: 885 VTACAVLDVLGPPYSDPDGRHCAYYRSFPFSNFPVEGISIPEEEKKD--YEWLQEREKPE 1058
Query: 261 NCQM 264
+ Q+
Sbjct: 1059SLQV 1070
>TC80943 similar to GP|15451150|gb|AAK96846.1 Unknown protein {Arabidopsis
thaliana}, partial (55%)
Length = 661
Score = 179 bits (454), Expect = 1e-45
Identities = 92/194 (47%), Positives = 123/194 (62%), Gaps = 2/194 (1%)
Frame = +2
Query: 83 LSRDLQFFKPGNIIKENQRVTYTTVYKCDNFSLCIFFLPERGVIPLHNHPGMTVFSKLLL 142
LSR ++ KP +TY +++ D+F++C+F P VIPLH+HP MTVFSKLL
Sbjct: 14 LSRVARWAKP---------ITYVDIHESDSFTMCMFCFPTSSVIPLHDHPQMTVFSKLLY 166
Query: 143 GQMHIKSYDWVDHEASHNLLQPS-SKLRLAKLKANKTFTAPCDTSVLYPTTGGNIHEFTA 201
G +H+K+YDWV+ P +++RLAKL +K APC+TSVLYP GGNIH FTA
Sbjct: 167 GSLHVKAYDWVEPPCIVKSKGPGHAQVRLAKLAVDKVLNAPCETSVLYPNCGGNIHCFTA 346
Query: 202 ITPCAVLDVIGPPYSKEDGRDCSYYKDYPYNAFPNEEKIGEVKDKD-DSYGLLEEIDMPE 260
+TPCA+LDV+ PPY + +GR C+YY DYPY+ F G + D D D Y L E++ P
Sbjct: 347 VTPCAMLDVLAPPYKEYEGRKCTYYHDYPYSTFSAGN--GSLCDGDEDEYAWLAEVE-PS 517
Query: 261 NCQMDGIEYLGPPI 274
N M+ Y GP I
Sbjct: 518 NLYMNSGVYAGPAI 559
>TC83576 similar to GP|15293013|gb|AAK93617.1 unknown protein {Arabidopsis
thaliana}, partial (37%)
Length = 695
Score = 96.3 bits (238), Expect = 1e-20
Identities = 45/101 (44%), Positives = 65/101 (63%), Gaps = 7/101 (6%)
Frame = +3
Query: 61 PSPRDVHKLCHILDNMKPEDVGLSRDLQFFKP--GNIIKENQR-----VTYTTVYKCDNF 113
P + + K+C L+ MKP DVGL ++ Q + G + + N + Y +++CD+F
Sbjct: 333 PCRKAISKVCEKLEKMKPSDVGLEQEAQVVRTWNGQMPESNGNHQPPPIKYLHLHECDSF 512
Query: 114 SLCIFFLPERGVIPLHNHPGMTVFSKLLLGQMHIKSYDWVD 154
S+ IF +P VIPLHNHPGMTV SKL+ G +H+KSYDW+D
Sbjct: 513 SIGIFCMPTSSVIPLHNHPGMTVLSKLIYGTVHVKSYDWID 635
>CA922711 similar to GP|20197959|gb| expressed protein {Arabidopsis
thaliana}, partial (21%)
Length = 806
Score = 87.0 bits (214), Expect = 7e-18
Identities = 45/135 (33%), Positives = 76/135 (55%), Gaps = 7/135 (5%)
Frame = +1
Query: 27 NKPYNHRRVKKHVPKALQELFDSCKQTFKGPGTVPSPRDVHKLCHILDNMKPEDVGLSRD 86
++P N ++ +P +Q+L+D CK + G + S + K+ +LD +KP VGL +
Sbjct: 406 HQPTNVSQLLSTMPY-IQKLYDVCKASLSPEGPI-SEEALEKVRTVLDGLKPSHVGLDHE 579
Query: 87 LQFFKP-------GNIIKENQRVTYTTVYKCDNFSLCIFFLPERGVIPLHNHPGMTVFSK 139
Q + GN E + Y +++CD FS+ +F + +IPLH+HP MTV SK
Sbjct: 580 AQLARSWKRSMNGGNGTPEGPPIKYIHLHECDKFSIGVFCMSPGSLIPLHDHPRMTVLSK 759
Query: 140 LLLGQMHIKSYDWVD 154
L G +H++++DW+D
Sbjct: 760 GLYGSLHVEAFDWID 804
>TC92643 similar to GP|15293013|gb|AAK93617.1 unknown protein {Arabidopsis
thaliana}, partial (7%)
Length = 648
Score = 50.4 bits (119), Expect = 7e-07
Identities = 28/79 (35%), Positives = 45/79 (56%), Gaps = 3/79 (3%)
Frame = +2
Query: 199 FTAITPCAVLDVIGPPYSKED---GRDCSYYKDYPYNAFPNEEKIGEVKDKDDSYGLLEE 255
F AITPCA+ D++ PPYS E+ GR+CSY++ P E++ + + ++ LE+
Sbjct: 32 FKAITPCALFDILTPPYSLEEEVNGRNCSYFRKSLRTDLPVLEELRGMSSSEITW--LEK 205
Query: 256 IDMPENCQMDGIEYLGPPI 274
I P + + +Y GP I
Sbjct: 206 IPPPSDLIIGNGQYRGPNI 262
>AW127175 similar to GP|15293013|gb| unknown protein {Arabidopsis thaliana},
partial (37%)
Length = 452
Score = 40.8 bits (94), Expect = 6e-04
Identities = 25/84 (29%), Positives = 44/84 (51%), Gaps = 8/84 (9%)
Frame = +3
Query: 38 HVPKALQELFDSCKQTFKGPGTVPSPRDVHKLCHILDNMKPEDVGLSRDLQFFK--PGNI 95
++P +Q L+ CK +F G V S V K+ LD +KP DVGL ++ Q + +
Sbjct: 177 NMPYCVQRLYRLCKASFSPDGPV-SEEVVKKVREKLDRIKPSDVGLEQEAQVVRNMSRTV 353
Query: 96 IKEN------QRVTYTTVYKCDNF 113
+++N + Y +++CD+F
Sbjct: 354 LEQNGSHHSLPAIKYLHLHECDSF 425
>TC81352 similar to GP|6633847|gb|AAF19706.1| F2K11.19 {Arabidopsis
thaliana}, partial (20%)
Length = 876
Score = 32.3 bits (72), Expect = 0.21
Identities = 24/79 (30%), Positives = 34/79 (42%), Gaps = 2/79 (2%)
Frame = +2
Query: 153 VDHEASHNLLQPSSKLRLAKLKANKTFTAPCDTSVLYPTTGGNIHEFTAITPCAVLDVIG 212
VD E+ +L+ S K + P + T GNIH F + +L++I
Sbjct: 158 VDFESWKTILERSEKNSGSISSQGAVCVLPNSLEARHLDTKGNIHAFGVL----LLEIIS 325
Query: 213 --PPYSKEDGRDCSYYKDY 229
PPY KE G + KDY
Sbjct: 326 GRPPYCKEKGYLVDWAKDY 382
>TC76324 similar to GP|7242813|emb|CAB77393.1 germin-like protein {Phaseolus
vulgaris}, partial (93%)
Length = 886
Score = 29.3 bits (64), Expect = 1.7
Identities = 28/105 (26%), Positives = 39/105 (36%), Gaps = 7/105 (6%)
Frame = +1
Query: 48 DSCKQTFKGPGTVPSPRDVHKLCHILDNMKPEDVGLSRDLQFFKPGNIIKENQRVTYTTV 107
D C K P T P H C L N+ +D FK GN + T+
Sbjct: 124 DFCVADLKAPDT---PSGYH--CKPLANITSDDFVFHG----FKAGNT-NNSFNAALTSA 273
Query: 108 YKCD-------NFSLCIFFLPERGVIPLHNHPGMTVFSKLLLGQM 145
+ D S + E G IP+H HPG T ++ G++
Sbjct: 274 FVTDFPGLNGLGISAARLDIAENGSIPMHTHPGATELLIIVQGEI 408
>TC78822 similar to PIR|T02962|T02962 peroxidase (EC 1.11.1.7) isozyme 40K
precursor cationic - common tobacco, partial (70%)
Length = 1181
Score = 29.3 bits (64), Expect = 1.7
Identities = 16/44 (36%), Positives = 21/44 (47%)
Frame = +2
Query: 159 HNLLQPSSKLRLAKLKANKTFTAPCDTSVLYPTTGGNIHEFTAI 202
H +P +L +L + F CD SVL +T GN E AI
Sbjct: 182 HVSSRPELPAKLIRLHFHDCFVRGCDASVLLESTAGNTAEKDAI 313
>TC78460 similar to GP|21554579|gb|AAM63621.1 flavonol synthase-like protein
{Arabidopsis thaliana}, partial (86%)
Length = 1442
Score = 28.9 bits (63), Expect = 2.3
Identities = 20/71 (28%), Positives = 32/71 (44%), Gaps = 12/71 (16%)
Frame = -1
Query: 117 IFFLPERGVIPLHNHPGMTVFSKLLLGQMHIKSYDWV------------DHEASHNLLQP 164
I FLPE L++HPG T L ++HI+ + W+ DH+ H ++
Sbjct: 821 IGFLPEEH---LYHHPGSTKSINHNLCELHIQCHHWIHQGNQLYASPKMDHQLDHLQIRM 651
Query: 165 SSKLRLAKLKA 175
L L + +A
Sbjct: 650 VIHLMLKQFRA 618
>TC88134 weakly similar to SP|Q9MB73|LGT_CITUN Limonoid
UDP-glucosyltransferase (EC 2.4.1.210) (Limonoid
glucosyltransferase) (Limonoid GTase), partial (27%)
Length = 1669
Score = 28.9 bits (63), Expect = 2.3
Identities = 12/35 (34%), Positives = 19/35 (54%)
Frame = -1
Query: 85 RDLQFFKPGNIIKENQRVTYTTVYKCDNFSLCIFF 119
R L FF P I + + TV++C ++L +FF
Sbjct: 1042 RQLNFFSPNIITLHHPEKIFVTVFQCTCYALHLFF 938
>TC80851 weakly similar to GP|21644692|dbj|BAC01248. hypothetical
protein~similar to Oryza sativa chromosome 1
P0665A11.10, partial (5%)
Length = 1527
Score = 27.3 bits (59), Expect = 6.6
Identities = 10/24 (41%), Positives = 18/24 (74%)
Frame = -3
Query: 145 MHIKSYDWVDHEASHNLLQPSSKL 168
+H+KS+ ++H +SHN++ SKL
Sbjct: 880 LHVKSFS-IEHNSSHNIVHSHSKL 812
>TC86278 similar to GP|17473703|gb|AAL38307.1 germin-like protein
{Arabidopsis thaliana}, partial (88%)
Length = 908
Score = 26.9 bits (58), Expect = 8.7
Identities = 28/80 (35%), Positives = 32/80 (40%), Gaps = 9/80 (11%)
Frame = +2
Query: 77 KPEDVGLSRDLQF---FKPGNIIKE-NQRVTYTTVYKCD-----NFSLCIFFLPERGVIP 127
KP S D F PGNI N VT V + S L GVIP
Sbjct: 158 KPASTVTSDDFAFEGLIAPGNITNIINAAVTPAFVAQFPAVNGLGLSAARLDLGPAGVIP 337
Query: 128 LHNHPGMTVFSKLLLGQMHI 147
LH HPG + L++ Q HI
Sbjct: 338 LHTHPGAS--ELLVVTQGHI 391
>AW689398 similar to GP|21593065|gb| NAM / CUC2-like protein {Arabidopsis
thaliana}, partial (13%)
Length = 651
Score = 26.9 bits (58), Expect = 8.7
Identities = 14/41 (34%), Positives = 21/41 (51%), Gaps = 2/41 (4%)
Frame = +2
Query: 229 YPYNAFPNEEK-IGEVKDK-DDSYGLLEEIDMPENCQMDGI 267
Y PN+ + ++ K DD YG L +D+ EN DG+
Sbjct: 386 YSRETNPNDNSTVTQISSKGDDGYGFLWNMDLEENSLEDGV 508
Database: MTGI
Posted date: Oct 22, 2004 3:39 PM
Number of letters in database: 27,044,181
Number of sequences in database: 36,976
Lambda K H
0.320 0.140 0.433
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 9,869,904
Number of Sequences: 36976
Number of extensions: 157919
Number of successful extensions: 760
Number of sequences better than 10.0: 34
Number of HSP's better than 10.0 without gapping: 749
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 752
length of query: 279
length of database: 9,014,727
effective HSP length: 95
effective length of query: 184
effective length of database: 5,502,007
effective search space: 1012369288
effective search space used: 1012369288
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.8 bits)
S2: 58 (26.9 bits)
Medicago: description of AC137822.15


