

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC137703.1 + phase: 0
(233 letters)
Database: MTGI
36,976 sequences; 27,044,181 total letters
Searching..................................................done
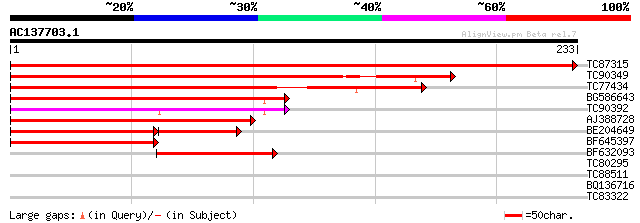
Score E
Sequences producing significant alignments: (bits) Value
TC87315 homologue to GP|21553886|gb|AAM62979.1 unknown {Arabidop... 478 e-135
TC90349 similar to GP|21553886|gb|AAM62979.1 unknown {Arabidopsi... 265 1e-71
TC77434 similar to GP|21554191|gb|AAM63270.1 unknown {Arabidopsi... 173 4e-44
BG586643 similar to GP|21553886|gb unknown {Arabidopsis thaliana... 167 3e-42
TC90392 similar to GP|21553886|gb|AAM62979.1 unknown {Arabidopsi... 149 9e-37
AJ388728 similar to PIR|H96694|H96 hypothetical protein F5A8.2 [... 139 1e-33
BE204649 similar to GP|21553886|gb| unknown {Arabidopsis thalian... 125 1e-29
BF645397 homologue to PIR|H96694|H966 hypothetical protein F5A8.... 100 4e-22
BF632093 similar to GP|20804498|dbj contains ESTs C97108(C52055)... 81 3e-16
TC80295 similar to SP|O82660|H136_ARATH Photosystem II stability... 30 1.0
TC88511 weakly similar to GP|20259567|gb|AAM14126.1 unknown prot... 28 3.0
BQ136716 similar to PIR|S51590|S515 mitochondrial processing pep... 27 8.7
TC83322 similar to PIR|E84706|E84706 hypothetical protein At2g30... 27 8.7
>TC87315 homologue to GP|21553886|gb|AAM62979.1 unknown {Arabidopsis
thaliana}, partial (50%)
Length = 1171
Score = 478 bits (1229), Expect = e-135
Identities = 233/233 (100%), Positives = 233/233 (100%)
Frame = +3
Query: 1 MSCNGCRVLRKGCSESCILRPCLQWIETPEAQGHATVFVAKFFGRAGLMSFISNVPEPQR 60
MSCNGCRVLRKGCSESCILRPCLQWIETPEAQGHATVFVAKFFGRAGLMSFISNVPEPQR
Sbjct: 102 MSCNGCRVLRKGCSESCILRPCLQWIETPEAQGHATVFVAKFFGRAGLMSFISNVPEPQR 281
Query: 61 PALFQSLLFEACGRTVNPVNGAVGLLWTGNWHVCQAAVETVLRGGTLRPLPELTLLDAPL 120
PALFQSLLFEACGRTVNPVNGAVGLLWTGNWHVCQAAVETVLRGGTLRPLPELTLLDAPL
Sbjct: 282 PALFQSLLFEACGRTVNPVNGAVGLLWTGNWHVCQAAVETVLRGGTLRPLPELTLLDAPL 461
Query: 121 TATDEASEAEDGGADAMWKIREPNFGLASSRSSNKVCSGSKRRRSEEFVKVPAAINLDLR 180
TATDEASEAEDGGADAMWKIREPNFGLASSRSSNKVCSGSKRRRSEEFVKVPAAINLDLR
Sbjct: 462 TATDEASEAEDGGADAMWKIREPNFGLASSRSSNKVCSGSKRRRSEEFVKVPAAINLDLR 641
Query: 181 LTPIFQQKAVEERRHGSPSMTSEESVTTTACLETGIGDRWSHGGDRKVLNLFI 233
LTPIFQQKAVEERRHGSPSMTSEESVTTTACLETGIGDRWSHGGDRKVLNLFI
Sbjct: 642 LTPIFQQKAVEERRHGSPSMTSEESVTTTACLETGIGDRWSHGGDRKVLNLFI 800
>TC90349 similar to GP|21553886|gb|AAM62979.1 unknown {Arabidopsis
thaliana}, partial (46%)
Length = 676
Score = 265 bits (677), Expect = 1e-71
Identities = 137/184 (74%), Positives = 150/184 (81%), Gaps = 1/184 (0%)
Frame = +3
Query: 1 MSCNGCRVLRKGCSESCILRPCLQWIETPEAQGHATVFVAKFFGRAGLMSFISNVPEPQR 60
MSCNGCRVLRKGCSESCILRPC+QWI+TPEAQGHATVFVAKFFGRA LMSFISNVP PQR
Sbjct: 108 MSCNGCRVLRKGCSESCILRPCIQWIDTPEAQGHATVFVAKFFGRADLMSFISNVPLPQR 287
Query: 61 PALFQSLLFEACGRTVNPVNGAVGLLWTGNWHVCQAAVETVLRGGTLRPLPELTLLDAPL 120
PALFQSLLFEACGRTVNPVNGAVGLLWTGNWHVCQAAVETVLRGGTL+PLPE LDA
Sbjct: 288 PALFQSLLFEACGRTVNPVNGAVGLLWTGNWHVCQAAVETVLRGGTLKPLPEFLGLDALA 467
Query: 121 TATDEASEAEDGGADAMWKIREPNFGLASSRSSNKVCSGSKRRRS-EEFVKVPAAINLDL 179
+ D++SE E DA KIR+PN + SG KR+RS E +K+PA +L+L
Sbjct: 468 STVDDSSEGEVTCNDAR-KIRDPN------PKVRFMSSGGKRKRSGGEVLKLPATTDLNL 626
Query: 180 RLTP 183
RLTP
Sbjct: 627 RLTP 638
>TC77434 similar to GP|21554191|gb|AAM63270.1 unknown {Arabidopsis
thaliana}, partial (71%)
Length = 1362
Score = 173 bits (439), Expect = 4e-44
Identities = 87/175 (49%), Positives = 119/175 (67%), Gaps = 4/175 (2%)
Frame = +3
Query: 1 MSCNGCRVLRKGCSESCILRPCLQWIETPEAQGHATVFVAKFFGRAGLMSFISNVPEPQR 60
MSCNGCRVLRKGCSE+C +RPCLQWI++PE+Q +ATVF+AKF+GRAGLM+ ++ PE R
Sbjct: 216 MSCNGCRVLRKGCSENCSIRPCLQWIKSPESQANATVFLAKFYGRAGLMNLVNAGPEHLR 395
Query: 61 PALFQSLLFEACGRTVNPVNGAVGLLWTGNWHVCQAAVETVLRGGTLRPLPELTLLDAPL 120
PA+F+SLL+EACGR VNP+ G+VGLLW+G+W +CQAAVE VL+G + P+
Sbjct: 396 PAIFRSLLYEACGRIVNPIYGSVGLLWSGSWQLCQAAVEAVLKGAPITPI---------- 545
Query: 121 TATDEASEAEDGGADAMWKIR----EPNFGLASSRSSNKVCSGSKRRRSEEFVKV 171
T EA+ G + IR E N ++ + +V S++R S +K+
Sbjct: 546 --TSEAAANGRGPPLKAYDIRHVSKEENSAASTELTQQRVKPRSRKRSSLAKLKI 704
>BG586643 similar to GP|21553886|gb unknown {Arabidopsis thaliana}, partial
(42%)
Length = 647
Score = 167 bits (423), Expect = 3e-42
Identities = 81/117 (69%), Positives = 90/117 (76%), Gaps = 2/117 (1%)
Frame = +1
Query: 1 MSCNGCRVLRKGCSESCILRPCLQWIETPEAQGHATVFVAKFFGRAGLMSFISNVPEPQR 60
MSCNGCRVLRKGCSE C+LR CL WI+ P+AQ +ATVFV KFFGRA LMSF+S V QR
Sbjct: 67 MSCNGCRVLRKGCSEDCMLRDCLTWIQNPQAQANATVFVTKFFGRATLMSFLSPVHPNQR 246
Query: 61 PALFQSLLFEACGRTVNPVNGAVGLLWTGNWHVCQAAVETVLR--GGTLRPLPELTL 115
LFQSL++EA GRT+NPVNGAVGLLWTG W CQ VE VLR GG L LP+ L
Sbjct: 247 SCLFQSLMYEAVGRTINPVNGAVGLLWTGKWQFCQLGVEQVLRGNGGALTALPDQLL 417
>TC90392 similar to GP|21553886|gb|AAM62979.1 unknown {Arabidopsis
thaliana}, partial (39%)
Length = 870
Score = 149 bits (376), Expect = 9e-37
Identities = 81/153 (52%), Positives = 90/153 (57%), Gaps = 38/153 (24%)
Frame = +1
Query: 1 MSCNGCRVLRKGCSESCILRPCLQWIETPEAQGHATVFVAKFFGRAGLMSFISNVPEPQR 60
MSCNGCRVLRKGCSE C+LR CL WI+ P+AQ +ATVFV KFFGRA LMSF+S V QR
Sbjct: 82 MSCNGCRVLRKGCSEDCMLRDCLTWIQNPQAQANATVFVTKFFGRATLMSFLSPVHPNQR 261
Query: 61 ------------------------------------PALFQSLLFEACGRTVNPVNGAVG 84
LFQSL++EA GRT+NPVNGAVG
Sbjct: 262 SCIYIYIYTTRINKS*CIWFKRIYKFLTCIN*FNYITGLFQSLMYEAVGRTINPVNGAVG 441
Query: 85 LLWTGNWHVCQAAVETVLR--GGTLRPLPELTL 115
LLWTG W CQ VE VLR GG L LP+ L
Sbjct: 442 LLWTGKWQFCQLGVEQVLRGNGGALTALPDQLL 540
>AJ388728 similar to PIR|H96694|H96 hypothetical protein F5A8.2 [imported] -
Arabidopsis thaliana, partial (44%)
Length = 567
Score = 139 bits (349), Expect = 1e-33
Identities = 62/101 (61%), Positives = 80/101 (78%)
Frame = +3
Query: 1 MSCNGCRVLRKGCSESCILRPCLQWIETPEAQGHATVFVAKFFGRAGLMSFISNVPEPQR 60
MSCNGCRVLRKGCSE+C +RPCLQWI++PE+Q +ATVF+AKF+GRAGLM+ + PE R
Sbjct: 141 MSCNGCRVLRKGCSENCSIRPCLQWIKSPESQANATVFLAKFYGRAGLMNLGNAGPEHLR 320
Query: 61 PALFQSLLFEACGRTVNPVNGAVGLLWTGNWHVCQAAVETV 101
A+F+S++ EACG V P G++ LLW +W +CQA VE V
Sbjct: 321 XAIFRSVVXEACGPIVXPXYGSIRLLWXRSWQLCQAXVEAV 443
>BE204649 similar to GP|21553886|gb| unknown {Arabidopsis thaliana},
partial (38%)
Length = 498
Score = 125 bits (315), Expect = 1e-29
Identities = 57/61 (93%), Positives = 59/61 (96%)
Frame = +2
Query: 1 MSCNGCRVLRKGCSESCILRPCLQWIETPEAQGHATVFVAKFFGRAGLMSFISNVPEPQR 60
MSCNGCRVLRKGCSESCILRPC+QWI+TPEAQGHATVFVAKFFGRA LMSFISNVP PQR
Sbjct: 38 MSCNGCRVLRKGCSESCILRPCIQWIDTPEAQGHATVFVAKFFGRADLMSFISNVPLPQR 217
Query: 61 P 61
P
Sbjct: 218P 220
Score = 78.2 bits (191), Expect = 3e-15
Identities = 34/34 (100%), Positives = 34/34 (100%)
Frame = +1
Query: 62 ALFQSLLFEACGRTVNPVNGAVGLLWTGNWHVCQ 95
ALFQSLLFEACGRTVNPVNGAVGLLWTGNWHVCQ
Sbjct: 397 ALFQSLLFEACGRTVNPVNGAVGLLWTGNWHVCQ 498
>BF645397 homologue to PIR|H96694|H966 hypothetical protein F5A8.2 [imported]
- Arabidopsis thaliana, partial (33%)
Length = 624
Score = 100 bits (250), Expect = 4e-22
Identities = 43/61 (70%), Positives = 54/61 (88%)
Frame = +2
Query: 1 MSCNGCRVLRKGCSESCILRPCLQWIETPEAQGHATVFVAKFFGRAGLMSFISNVPEPQR 60
MSCNGCRVLRKGCSE+C +RPCLQWI++PE+Q +ATVF+AKF+GRAGLM+ ++ PE R
Sbjct: 83 MSCNGCRVLRKGCSENCSIRPCLQWIKSPESQANATVFLAKFYGRAGLMNLVNAGPEHLR 262
Query: 61 P 61
P
Sbjct: 263 P 265
Score = 28.5 bits (62), Expect = 2.3
Identities = 11/17 (64%), Positives = 14/17 (81%)
Frame = +1
Query: 62 ALFQSLLFEACGRTVNP 78
A+F+SLL+EACG NP
Sbjct: 574 AIFRSLLYEACGXIXNP 624
>BF632093 similar to GP|20804498|dbj contains ESTs C97108(C52055)
AU085724(C52055) C25099(S5247)~similar to Oryza sativa
chromosome 1, partial (20%)
Length = 357
Score = 81.3 bits (199), Expect = 3e-16
Identities = 33/50 (66%), Positives = 44/50 (88%)
Frame = +1
Query: 61 PALFQSLLFEACGRTVNPVNGAVGLLWTGNWHVCQAAVETVLRGGTLRPL 110
PA+F+SLL+EACGR VNP+ G+VGLLW+G+WH+CQAAVE VL+G + P+
Sbjct: 1 PAIFRSLLYEACGRIVNPIYGSVGLLWSGSWHLCQAAVEAVLKGEPITPI 150
>TC80295 similar to SP|O82660|H136_ARATH Photosystem II stability/assembly
factor HCF136 chloroplast precursor. [Mouse-ear cress],
partial (50%)
Length = 879
Score = 29.6 bits (65), Expect = 1.0
Identities = 11/27 (40%), Positives = 19/27 (69%)
Frame = +2
Query: 46 AGLMSFISNVPEPQRPALFQSLLFEAC 72
A L+ +SN+PEP+ +L + LL++ C
Sbjct: 146 ASLLKLLSNLPEPEDSSLQKPLLYQCC 226
>TC88511 weakly similar to GP|20259567|gb|AAM14126.1 unknown protein
{Arabidopsis thaliana}, partial (22%)
Length = 1060
Score = 28.1 bits (61), Expect = 3.0
Identities = 12/21 (57%), Positives = 15/21 (71%), Gaps = 2/21 (9%)
Frame = -3
Query: 4 NGCRVLRKGCSESCI--LRPC 22
NGCR+L +GC+ CI LR C
Sbjct: 854 NGCRLLCRGCNRLCIILLRQC 792
>BQ136716 similar to PIR|S51590|S515 mitochondrial processing peptidase (EC
3.4.24.64) alpha-II chain precursor - potato, partial
(3%)
Length = 958
Score = 26.6 bits (57), Expect = 8.7
Identities = 17/49 (34%), Positives = 23/49 (46%)
Frame = +2
Query: 162 RRRSEEFVKVPAAINLDLRLTPIFQQKAVEERRHGSPSMTSEESVTTTA 210
RRR + + +P I TP Q + R SPS+ + S TTTA
Sbjct: 614 RRR*DPLIYLPPPIITSTISTPELQLSIYTDNRAHSPSIPTYISQTTTA 760
>TC83322 similar to PIR|E84706|E84706 hypothetical protein At2g30280
[imported] - Arabidopsis thaliana, partial (19%)
Length = 725
Score = 26.6 bits (57), Expect = 8.7
Identities = 10/34 (29%), Positives = 19/34 (55%)
Frame = +1
Query: 124 DEASEAEDGGADAMWKIREPNFGLASSRSSNKVC 157
++ S A+D + +WK R+ N G A + ++C
Sbjct: 364 EKESSAKDARFEQIWKSRKVNKGTADENALQEIC 465
Database: MTGI
Posted date: Oct 22, 2004 3:39 PM
Number of letters in database: 27,044,181
Number of sequences in database: 36,976
Lambda K H
0.319 0.134 0.416
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 7,971,883
Number of Sequences: 36976
Number of extensions: 105382
Number of successful extensions: 542
Number of sequences better than 10.0: 26
Number of HSP's better than 10.0 without gapping: 539
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 541
length of query: 233
length of database: 9,014,727
effective HSP length: 93
effective length of query: 140
effective length of database: 5,575,959
effective search space: 780634260
effective search space used: 780634260
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.8 bits)
S2: 57 (26.6 bits)
Medicago: description of AC137703.1


