

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC137667.13 - phase: 0 /pseudo
(267 letters)
Database: MTGI
36,976 sequences; 27,044,181 total letters
Searching..................................................done
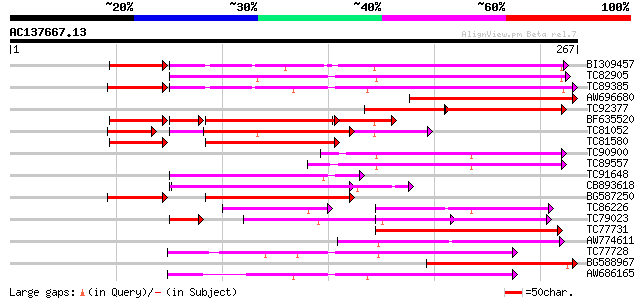
Score E
Sequences producing significant alignments: (bits) Value
BI309457 weakly similar to GP|15787901|gb resistance gene analog... 121 1e-33
TC82905 weakly similar to GP|3947733|emb|CAA08797.1 NL25 {Solanu... 118 3e-27
TC89385 similar to GP|19774173|gb|AAL99063.1 NBS-LRR-Toll resist... 113 4e-27
AW696680 similar to GP|19774149|gb NBS-LRR-Toll resistance gene ... 102 2e-22
TC92377 similar to GP|19774149|gb|AAL99051.1 NBS-LRR-Toll resist... 71 4e-22
BF635520 weakly similar to GP|3947735|emb| NL27 {Solanum tuberos... 73 7e-19
TC81052 weakly similar to GP|10178211|dbj|BAB11635. TMV resistan... 82 4e-18
TC81580 weakly similar to GP|7341113|gb|AAF61210.1| unknown {Gly... 79 2e-17
TC90900 weakly similar to PIR|D96753|D96753 Similar to disease r... 84 4e-17
TC89557 weakly similar to GP|10178211|dbj|BAB11635. TMV resistan... 74 8e-14
TC91648 weakly similar to GP|23477201|emb|CAD36199. NLS-TIR-NBS ... 72 2e-13
CB893618 weakly similar to GP|3947733|emb| NL25 {Solanum tuberos... 70 1e-12
BG587250 weakly similar to GP|3947735|emb| NL27 {Solanum tuberos... 60 2e-11
TC86226 weakly similar to GP|18033111|gb|AAL56987.1 functional c... 52 3e-11
TC79023 weakly similar to GP|18033111|gb|AAL56987.1 functional c... 63 1e-10
TC77731 weakly similar to GP|18033111|gb|AAL56987.1 functional c... 62 3e-10
AW774611 similar to GP|5817345|gb|A putative NBS-LRR type diseas... 60 7e-10
TC77728 similar to GP|12056928|gb|AAG48132.1 putative resistance... 59 2e-09
BG588967 similar to GP|7107246|gb| unknown {Cicer arietinum}, pa... 59 2e-09
AW686165 weakly similar to GP|18033111|gb functional candidate r... 49 2e-06
>BI309457 weakly similar to GP|15787901|gb resistance gene analog NBS7
{Helianthus annuus}, partial (45%)
Length = 806
Score = 121 bits (303), Expect(2) = 1e-33
Identities = 79/197 (40%), Positives = 109/197 (55%), Gaps = 9/197 (4%)
Frame = +3
Query: 76 SLIFIVVFSKNYASSTIGACKN*NPSFTALKSPKGMFCLFSMMLILMR*DIKK------E 129
S +F+ VFS NYASST C G L + + D++K E
Sbjct: 234 SQVFVAVFSINYASSTW--CLQELEKICECVKGSGKHVL-PVFYDVDPSDVRKQSGIYGE 404
Query: 130 IMLKPFSNMNKDSNKTLRWCKDGGKLKHKSPISGWDVRHND--AQIEKIVEEIMNILGYK 187
+K ++ K +W +D LK ISGWD+R +I+KIV+ I+NIL YK
Sbjct: 405 AFIKHEQRFQQEFQKVSKW-RDA--LKQVGSISGWDLRDKPQAGEIKKIVQTILNILKYK 575
Query: 188 STSLPNYLAGMDSLTEELEKHLLLDSVDDVRVVGVCGMGGIGKKAIATALYNKIFHQFPV 247
S+ L G+DS + L+ HLLLDSVD VR +G+CGMGGIGK +A ALY++I H+F
Sbjct: 576 SSCFSKDLVGIDSRLDGLQNHLLLDSVDSVRAIGICGMGGIGKTTLAMALYDQISHRFSA 755
Query: 248 LFLIDDLRKIY-RHDGP 263
+ IDD+++ Y HDGP
Sbjct: 756 SWFIDDVKQNY*LHDGP 806
Score = 38.9 bits (89), Expect(2) = 1e-33
Identities = 18/27 (66%), Positives = 22/27 (80%)
Frame = +2
Query: 48 KRYFCI*G*F*AQQR*IHWIRAPWSNR 74
KR++CI G* *+ +R*IHW RAP SNR
Sbjct: 149 KRHYCIFG*H*SSKR*IHWTRAPSSNR 229
>TC82905 weakly similar to GP|3947733|emb|CAA08797.1 NL25 {Solanum
tuberosum}, partial (22%)
Length = 832
Score = 118 bits (295), Expect = 3e-27
Identities = 76/195 (38%), Positives = 108/195 (54%), Gaps = 6/195 (3%)
Frame = +1
Query: 76 SLIFIVVFSKNYASSTIGACKN*NPSFTALKSPKGMFCLF---SMMLILMR*DIKKEIML 132
S +++ +FSKNYASST + S K + +F + + I E +
Sbjct: 250 SQVYVAIFSKNYASSTWCLQELEKICECIKGSGKHVLPVFYDVDPSEVRKQSGIYSEAFV 429
Query: 133 KPFSNMNKDSNKTLRWCKDGGKLKHKSPISGWDVRHNDA--QIEKIVEEIMNILGYKSTS 190
K +DS K RW + L+ ISGWD+R +I++IV++I+NIL K +
Sbjct: 430 KHEQRFQQDSMKVSRWRE---ALEQVGSISGWDLRDEPLAREIKEIVQKIINILECKYSC 600
Query: 191 LPNYLAGMDSLTEELEKHLLLDSVDDVRVVGVCGMGGIGKKAIATALYNKIFHQFPVLFL 250
+ L G+DS + L+ HLLL+SVD VR +G+CGMGGIGK +AT LY +I HQF
Sbjct: 601 VSKDLVGIDSPIQALQNHLLLNSVDGVRAIGICGMGGIGKTTLATTLYGQISHQFSASCF 780
Query: 251 IDDLRKIY-RHDGPI 264
IDD+ KIY HD P+
Sbjct: 781 IDDVTKIYGLHDDPL 825
>TC89385 similar to GP|19774173|gb|AAL99063.1 NBS-LRR-Toll resistance gene
analog protein {Medicago ruthenica}, partial (98%)
Length = 1285
Score = 113 bits (282), Expect(2) = 4e-27
Identities = 77/201 (38%), Positives = 109/201 (53%), Gaps = 9/201 (4%)
Frame = +3
Query: 76 SLIFIVVFSKNYASSTIGACKN*NPSFTALKSPKGMFCLFSMMLILMR*DIKKEIML--- 132
S +F+ V S+NYASST C G + L + + ++KK+ +
Sbjct: 231 SQVFVAVLSRNYASSTW--CLQELEKICECIKGSGKYVL-PIFYGVDPSEVKKQSGIYWD 401
Query: 133 ---KPFSNMNKDSNKTLRWCKDGGKLKHKSPISGWDVR--HNDAQIEKIVEEIMNILGYK 187
K +D +K RW + L I+GWD+R ++EKIV+ I+NIL K
Sbjct: 402 DFAKHEQRFKQDPHKVSRWRE---ALNQVGSIAGWDLRDKQQSVEVEKIVQTILNILKCK 572
Query: 188 STSLPNYLAGMDSLTEELEKHLLLDSVDDVRVVGVCGMGGIGKKAIATALYNKIFHQFPV 247
S+ + L G++S TE L+ LLL+SVD VRV+G+ GMGG GK +A LY +I H+F
Sbjct: 573 SSFVSKDLVGINSRTEALKHQLLLNSVDGVRVIGIWGMGGKGKTTLAMNLYGQICHRFDA 752
Query: 248 LFLIDDLRKIYR-HDGPISAQ 267
IDD+ KI+R HDGPI AQ
Sbjct: 753 SCFIDDVSKIFRLHDGPIDAQ 815
Score = 25.4 bits (54), Expect(2) = 4e-27
Identities = 14/28 (50%), Positives = 17/28 (60%)
Frame = +2
Query: 47 KKRYFCI*G*F*AQQR*IHWIRAPWSNR 74
KKRY C G* + + *IH AP +NR
Sbjct: 143 KKRYLCFQG*HKSSKG*IHRT*APSNNR 226
>AW696680 similar to GP|19774149|gb NBS-LRR-Toll resistance gene analog
protein {Medicago sativa}, partial (96%)
Length = 641
Score = 102 bits (253), Expect = 2e-22
Identities = 49/79 (62%), Positives = 59/79 (74%)
Frame = +2
Query: 189 TSLPNYLAGMDSLTEELEKHLLLDSVDDVRVVGVCGMGGIGKKAIATALYNKIFHQFPVL 248
+SLP L GM+S +++ LLLDS+DDVRVVG+CGMGGIGK +ATALY +I HQF
Sbjct: 8 SSLPKELVGMNSHIDKVANLLLLDSIDDVRVVGICGMGGIGKTTLATALYGQISHQFDAR 187
Query: 249 FLIDDLRKIYRHDGPISAQ 267
IDDL KIYRHDG + AQ
Sbjct: 188 CFIDDLSKIYRHDGQVGAQ 244
>TC92377 similar to GP|19774149|gb|AAL99051.1 NBS-LRR-Toll resistance gene
analog protein {Medicago sativa}, partial (57%)
Length = 945
Score = 70.9 bits (172), Expect(2) = 4e-22
Identities = 32/57 (56%), Positives = 44/57 (77%)
Frame = +1
Query: 206 EKHLLLDSVDDVRVVGVCGMGGIGKKAIATALYNKIFHQFPVLFLIDDLRKIYRHDG 262
+K LLL+S D ++VVG+ GMGGIGK +ATALY++I HQF IDD+ K+YR++G
Sbjct: 223 KKCLLLESTDIIQVVGIYGMGGIGKTTLATALYDRISHQFDACCFIDDISKVYRNEG 393
Score = 50.8 bits (120), Expect(2) = 4e-22
Identities = 26/40 (65%), Positives = 30/40 (75%)
Frame = +3
Query: 168 HNDAQIEKIVEEIMNILGYKSTSLPNYLAGMDSLTEELEK 207
H IEKIVEEI+N LGY+ +SLP L G+DSL EELEK
Sbjct: 108 HQYIVIEKIVEEIINRLGYRFSSLPKDLVGIDSLIEELEK 227
>BF635520 weakly similar to GP|3947735|emb| NL27 {Solanum tuberosum}, partial
(8%)
Length = 577
Score = 72.8 bits (177), Expect(4) = 7e-19
Identities = 38/63 (60%), Positives = 44/63 (69%)
Frame = +2
Query: 93 GACKN*NPSFTALKSPKGMFCLFSMMLILMR*DIKKEIMLKPFSNMNKDSNKTLRWCKDG 152
GA KN* AL+ K MF LF MMLIL+R + K E + K SNMNKDSNKT +WC+DG
Sbjct: 299 GAFKN*RRFVNALRD*KNMFFLFFMMLILLRYESKVEFIGKLLSNMNKDSNKTFKWCQDG 478
Query: 153 GKL 155
GKL
Sbjct: 479 GKL 487
Score = 28.9 bits (63), Expect(4) = 7e-19
Identities = 15/32 (46%), Positives = 20/32 (61%), Gaps = 2/32 (6%)
Frame = +3
Query: 153 GKLKHKSPISGWDVRHND--AQIEKIVEEIMN 182
G K ISGWD+R A I+KIV++I+N
Sbjct: 480 GSSKQVGSISGWDLRDKPQAAXIKKIVQKIIN 575
Score = 26.2 bits (56), Expect(4) = 7e-19
Identities = 10/16 (62%), Positives = 14/16 (87%)
Frame = +1
Query: 76 SLIFIVVFSKNYASST 91
S +++ VFS+NYASST
Sbjct: 250 SHVYVAVFSRNYASST 297
Score = 21.9 bits (45), Expect(4) = 7e-19
Identities = 14/27 (51%), Positives = 19/27 (69%)
Frame = +3
Query: 48 KRYFCI*G*F*AQQR*IHWIRAPWSNR 74
KR++ I G* *+ +R*I+ I A SNR
Sbjct: 165 KRHYGISG*H*SSKR*IYRI*ASSSNR 245
>TC81052 weakly similar to GP|10178211|dbj|BAB11635. TMV resistance protein
N {Arabidopsis thaliana}, partial (8%)
Length = 656
Score = 82.4 bits (202), Expect(2) = 4e-18
Identities = 44/71 (61%), Positives = 51/71 (70%)
Frame = +3
Query: 92 IGACKN*NPSFTALKSPKGMFCLFSMMLILMR*DIKKEIMLKPFSNMNKDSNKTLRWCKD 151
+GAC+N S TA+K + +F LF MMLIL+R*DIKKE +KPF NMNK SN TL W K
Sbjct: 318 LGACENWYTSLTAVKFLEDVFFLFFMMLILLR*DIKKEFTVKPFPNMNKHSNMTLMWYKV 497
Query: 152 GGKLKHKSPIS 162
GGKL HK IS
Sbjct: 498 GGKL*HKWGIS 530
Score = 51.6 bits (122), Expect = 3e-07
Identities = 44/129 (34%), Positives = 63/129 (48%), Gaps = 5/129 (3%)
Frame = +2
Query: 76 SLIFIVVFSKNYASSTIGACKN*NPSFTALKSPKGMFCLF---SMMLILMR*DIKKEIML 132
S +FI V SKNY+SST + + + S + + +F + + I E
Sbjct: 272 SQVFIAVLSKNYSSSTWCLRELVHILDCSQVSGRRVLPVFYDVDPSEVRHQKGIYGEAFS 451
Query: 133 KPFSNMNKDSNKTLRWCKDGGKLKHKSPISGWDVRHND--AQIEKIVEEIMNILGYKSTS 190
K DS+ W + L ISGWD+R A+I+KIVEEI+NILG+ +S
Sbjct: 452 KHEQTFQHDSHVVQSWRE---ALTQVGNISGWDLRDKPQYAEIKKIVEEILNILGHNFSS 622
Query: 191 LPNYLAGMD 199
LP L GM+
Sbjct: 623 LPKELVGMN 649
Score = 25.8 bits (55), Expect(2) = 4e-18
Identities = 14/23 (60%), Positives = 17/23 (73%)
Frame = +1
Query: 47 KKRYFCI*G*F*AQQR*IHWIRA 69
KKR+FCI* *+ +R*IH RA
Sbjct: 184 KKRHFCI*RRR*SSKR*IHTTRA 252
>TC81580 weakly similar to GP|7341113|gb|AAF61210.1| unknown {Glycine max},
partial (84%)
Length = 827
Score = 79.3 bits (194), Expect(2) = 2e-17
Identities = 38/63 (60%), Positives = 46/63 (72%)
Frame = +3
Query: 93 GACKN*NPSFTALKSPKGMFCLFSMMLILMR*DIKKEIMLKPFSNMNKDSNKTLRWCKDG 152
GACKN* A + P+ +F LF MMLIL+ + K ++KP SNMNKDSNKTLRWC+DG
Sbjct: 324 GACKN*RRYVNAFRDPENIFFLFFMMLILLWFESKVGFIVKPLSNMNKDSNKTLRWCQDG 503
Query: 153 GKL 155
GKL
Sbjct: 504 GKL 512
Score = 36.2 bits (82), Expect = 0.014
Identities = 29/95 (30%), Positives = 43/95 (44%), Gaps = 3/95 (3%)
Frame = +2
Query: 76 SLIFIVVFSKNYASSTIGACKN*NPSFTALKSPKGMFCLF---SMMLILMR*DIKKEIML 132
S +F+ VFS+NYASST + +S K + +F ++ + I E +
Sbjct: 275 SRVFVAVFSRNYASSTWCLQELEKICKCVQRSRKHILPVFYDVDPSVVRKQSGIYCEAFV 454
Query: 133 KPFSNMNKDSNKTLRWCKDGGKLKHKSPISGWDVR 167
K +D RW + LKH ISGWD+R
Sbjct: 455 KHEQRFQQDFEMVSRWRE---ALKHVGSISGWDLR 550
Score = 26.2 bits (56), Expect(2) = 2e-17
Identities = 15/27 (55%), Positives = 19/27 (69%)
Frame = +1
Query: 48 KRYFCI*G*F*AQQR*IHWIRAPWSNR 74
KR+FCI G* *+ + *I+ AP SNR
Sbjct: 190 KRHFCISG*H*SSKG*IYRT*APSSNR 270
>TC90900 weakly similar to PIR|D96753|D96753 Similar to disease resistance
proteins [imported] - Arabidopsis thaliana, partial (7%)
Length = 826
Score = 84.3 bits (207), Expect = 4e-17
Identities = 50/119 (42%), Positives = 70/119 (58%), Gaps = 3/119 (2%)
Frame = +2
Query: 147 RWCKDGGKLKHKSPISGWDVRHNDA--QIEKIVEEIMNILGYKSTSLPNYLAGMDSLTEE 204
RW K G+L + + GWDVR+ +IE IV+E++ LG+K + + L EE
Sbjct: 5 RWTKAMGRL---AELVGWDVRNKPEFREIENIVQEVIKTLGHKFSGFADDLIATQPRVEE 175
Query: 205 LEKHLLLDSVDD-VRVVGVCGMGGIGKKAIATALYNKIFHQFPVLFLIDDLRKIYRHDG 262
LE L L S DD +RVVG+ GM GIGK +A+ LY++I QF I+++ KIYR G
Sbjct: 176 LESLLKLSSDDDELRVVGIWGMAGIGKTTLASVLYDRISSQFDASCFIENVSKIYRDGG 352
>TC89557 weakly similar to GP|10178211|dbj|BAB11635. TMV resistance protein
N {Arabidopsis thaliana}, partial (8%)
Length = 1567
Score = 73.6 bits (179), Expect = 8e-14
Identities = 49/125 (39%), Positives = 65/125 (51%), Gaps = 3/125 (2%)
Frame = +1
Query: 141 DSNKTLRWCKDGGKLKHKSPISGWDVRHNDA--QIEKIVEEIMNILGYKSTSLPNYLAGM 198
+S K RW + + +GWDVR IEKIVE ++ LG + + L G+
Sbjct: 631 NSGKVARWRTT---MTYLGGTAGWDVRVEPEFEMIEKIVEAVIKKLGRTFSGSTDDLIGI 801
Query: 199 DSLTEELEKHLLLDSVDD-VRVVGVCGMGGIGKKAIATALYNKIFHQFPVLFLIDDLRKI 257
E LE L L S DD RV+G+ GM GIGK +AT LY+KI QF I+++ KI
Sbjct: 802 QPHIEALENLLKLSSKDDGCRVLGIWGMDGIGKTTLATVLYDKISFQFDACCFIENVSKI 981
Query: 258 YRHDG 262
Y G
Sbjct: 982 YEDGG 996
>TC91648 weakly similar to GP|23477201|emb|CAD36199. NLS-TIR-NBS disease
resistance protein {Populus tremula}, partial (17%)
Length = 666
Score = 72.4 bits (176), Expect = 2e-13
Identities = 44/96 (45%), Positives = 56/96 (57%), Gaps = 4/96 (4%)
Frame = +3
Query: 76 SLIFIVVFSKNYASSTIGACKN*NPSFTALKSPKGMFCLFSMMLILMR*DIKKEIMLKPF 135
+L F++ FS G C+N + F A + +FCL SMMLI R DIKK++MLKP
Sbjct: 330 TLRFLLWFSLRTMLPQFGVCENWSAYFKAFNYLENVFCLCSMMLIHQRCDIKKDVMLKP* 509
Query: 136 SNMNKDSNKTL----RWCKDGGKLKHKSPISGWDVR 167
NM KDSNKTL RW + L + +SGWDVR
Sbjct: 510 LNMKKDSNKTLKIXQRWRE---ALTQVANLSGWDVR 608
Score = 31.2 bits (69), Expect = 0.44
Identities = 14/16 (87%), Positives = 15/16 (93%)
Frame = +2
Query: 75 DSLIFIVVFSKNYASS 90
DS IF+VVFSKNYASS
Sbjct: 329 DSQIFVVVFSKNYASS 376
>CB893618 weakly similar to GP|3947733|emb| NL25 {Solanum tuberosum}, partial
(18%)
Length = 927
Score = 69.7 bits (169), Expect = 1e-12
Identities = 46/117 (39%), Positives = 66/117 (56%), Gaps = 2/117 (1%)
Frame = +3
Query: 76 SLIFIVVFSKNYASSTIGACKN*NPSFTALKSPKGMFCLFSMMLILMR*DIKKEIMLKPF 135
+L F +++S +G C+N*+ S TA+ + FCLF MMLI +R + K +M+ P
Sbjct: 144 ALRFSLLYSLKTMLLQLGVCEN*HTSLTAVYCMENAFCLFFMMLIFLRCESKVVVMVNPL 323
Query: 136 SNMNKDSNKTLRWCKDGGKLKHKSPIS--GWDVRHNDAQIEKIVEEIMNILGYKSTS 190
+ M +DS T WCK+G KL IS G V + ++E I+E I NILG K S
Sbjct: 324 TIMERDSKITQIWCKEGEKLYS*WVISLAGIYVISHIMELENIIEHI-NILGCKFPS 491
Score = 53.9 bits (128), Expect = 6e-08
Identities = 35/86 (40%), Positives = 45/86 (51%)
Frame = +1
Query: 77 LIFIVVFSKNYASSTIGACKN*NPSFTALKSPKGMFCLFSMMLILMR*DIKKEIMLKPFS 136
L F +++S +G C+N N SF A+K+ + F LFSMMLIL+R K + P
Sbjct: 682 LRFTLLYSLRTMLPQLGVCENWNTSFIAVKNMENTFFLFSMMLILLRYKSKVVATVTPCP 861
Query: 137 NMNKDSNKTLRWCKDGGKLKHKSPIS 162
NM W KDG KL HK IS
Sbjct: 862 NM------A*IWFKDGEKLSHKWGIS 921
Score = 29.6 bits (65), Expect = 1.3
Identities = 13/16 (81%), Positives = 15/16 (93%)
Frame = +3
Query: 76 SLIFIVVFSKNYASST 91
S ++IVVFSKNYASST
Sbjct: 681 SQVYIVVFSKNYASST 728
>BG587250 weakly similar to GP|3947735|emb| NL27 {Solanum tuberosum}, partial
(9%)
Length = 655
Score = 60.5 bits (145), Expect(2) = 2e-11
Identities = 36/70 (51%), Positives = 43/70 (61%)
Frame = +2
Query: 93 GACKN*NPSFTALKSPKGMFCLFSMMLILMR*DIKKEIMLKPFSNMNKDSNKTLRWCKDG 152
GACKN* A + + MF LFSM+ IL+R* K + NMNKDSNKTL C+DG
Sbjct: 281 GACKN*RRFVNASRDLENMFFLFSMVSILLR*KSKVGFIGTILPNMNKDSNKTLIRCQDG 460
Query: 153 GKLKHKSPIS 162
GKL K +S
Sbjct: 461 GKL*TKWGVS 490
Score = 25.4 bits (54), Expect(2) = 2e-11
Identities = 14/28 (50%), Positives = 17/28 (60%)
Frame = +3
Query: 47 KKRYFCI*G*F*AQQR*IHWIRAPWSNR 74
KKRY C G* + + *IH AP +NR
Sbjct: 144 KKRYLCFQG*HKSSKG*IHRT*APSNNR 227
>TC86226 weakly similar to GP|18033111|gb|AAL56987.1 functional candidate
resistance protein KR1 {Glycine max}, partial (24%)
Length = 1769
Score = 52.0 bits (123), Expect(2) = 3e-11
Identities = 31/85 (36%), Positives = 50/85 (58%), Gaps = 1/85 (1%)
Frame = +3
Query: 173 IEKIVEEIMNILGYKSTSLPNYLAGMDSLTEELEKHLLLDSVDD-VRVVGVCGMGGIGKK 231
I KIV+ I N + +S + Y G+ S +++ K LL + DD V +VG+ G+GG GK
Sbjct: 558 IGKIVKYISNKISRQSLHVATYPVGLQSRVQQV-KSLLDEGPDDGVHMVGIYGIGGSGKS 734
Query: 232 AIATALYNKIFHQFPVLFLIDDLRK 256
+A A+YN + QF L ++ +R+
Sbjct: 735 TLARAIYNFVADQFEGLCFLEQVRE 809
Score = 33.1 bits (74), Expect(2) = 3e-11
Identities = 22/55 (40%), Positives = 30/55 (54%), Gaps = 3/55 (5%)
Frame = +1
Query: 101 SFTALKSPKGMFCLFSMMLILMR*DIKKEIMLKPFSNMN---KDSNKTLRWCKDG 152
SFTA K FCLFS++ L+ DI + M+K + +M K KT R C +G
Sbjct: 322 SFTATKPRVASFCLFSLVWNLLMCDIIQVGMVKHWLSMKIGFKMIRKTWRGCSNG 486
>TC79023 weakly similar to GP|18033111|gb|AAL56987.1 functional candidate
resistance protein KR1 {Glycine max}, partial (37%)
Length = 2072
Score = 62.8 bits (151), Expect = 1e-10
Identities = 33/83 (39%), Positives = 50/83 (59%)
Frame = +3
Query: 173 IEKIVEEIMNILGYKSTSLPNYLAGMDSLTEELEKHLLLDSVDDVRVVGVCGMGGIGKKA 232
I KIVE+I N + + + Y G+ S + ++ HL S D+V +VG+ G GGIGK
Sbjct: 705 IGKIVEDISNRISREPLDVAKYPVGLQSRVQHVKGHLDEKSDDEVHMVGLYGTGGIGKST 884
Query: 233 IATALYNKIFHQFPVLFLIDDLR 255
+A A+YN I QF VL ++++R
Sbjct: 885 LAKAIYNFIADQFEVLCFLENVR 953
Score = 38.9 bits (89), Expect(2) = 2e-05
Identities = 31/106 (29%), Positives = 50/106 (46%), Gaps = 6/106 (5%)
Frame = +2
Query: 111 MFCLFSMMLILMR*DIKKEIMLKPFSNMNKDSNK---TLRWCKDGGKLKHKSPISGWDVR 167
+FCLF ++ IL+ DI + M+K + K TLR C +G KL K +
Sbjct: 359 LFCLFLLVWILLMCDITQVDMVKH*LCIKKSFKMIRITLRGCSNGKKL*AKRLTCLASII 538
Query: 168 HNDAQIE---KIVEEIMNILGYKSTSLPNYLAGMDSLTEELEKHLL 210
+ D + E KIVE+I N + K + Y + + ++ HL+
Sbjct: 539 NMDIEYEFIGKIVEDISNRINRKPQDVAKYHVELQLQVQHIKTHLI 676
Score = 25.8 bits (55), Expect(2) = 2e-05
Identities = 12/16 (75%), Positives = 14/16 (87%)
Frame = +1
Query: 76 SLIFIVVFSKNYASST 91
S IFI VFS+NYASS+
Sbjct: 256 SRIFIPVFSENYASSS 303
>TC77731 weakly similar to GP|18033111|gb|AAL56987.1 functional candidate
resistance protein KR1 {Glycine max}, partial (23%)
Length = 1651
Score = 61.6 bits (148), Expect = 3e-10
Identities = 32/88 (36%), Positives = 54/88 (61%)
Frame = +1
Query: 173 IEKIVEEIMNILGYKSTSLPNYLAGMDSLTEELEKHLLLDSVDDVRVVGVCGMGGIGKKA 232
IEKIVE+I N + + ++ Y G+ S E+++ L + S D+VR+VG+ G GG+GK
Sbjct: 514 IEKIVEDISNNINHVFLNVAKYPVGLQSRIEQVKLLLDMGSEDEVRMVGLFGTGGMGKST 693
Query: 233 IATALYNKIFHQFPVLFLIDDLRKIYRH 260
+A A+YN + QF + + ++R+ H
Sbjct: 694 LAKAVYNFVADQFEGVCFLHNVRENSTH 777
Score = 28.9 bits (63), Expect = 2.2
Identities = 20/59 (33%), Positives = 30/59 (49%), Gaps = 3/59 (5%)
Frame = +3
Query: 97 N*NPSFTALKSPKGMFCLFSMMLILMR*DIKKEIMLKPFSNMN---KDSNKTLRWCKDG 152
N + SFTA + +C FSMM L+ DI+ +M+ +M K + KT C +G
Sbjct: 30 NLSTSFTATRQRVA*YCPFSMMWNLLTYDIRVGVMVSILLSMKRGFKITRKTWSDCVNG 206
>AW774611 similar to GP|5817345|gb|A putative NBS-LRR type disease resistance
protein {Pisum sativum}, partial (49%)
Length = 718
Score = 60.5 bits (145), Expect = 7e-10
Identities = 35/110 (31%), Positives = 62/110 (55%), Gaps = 3/110 (2%)
Frame = +1
Query: 155 LKHKSPISGWDVRHNDAQ---IEKIVEEIMNILGYKSTSLPNYLAGMDSLTEELEKHLLL 211
L + +SGWD + ++ ++KIV+E++ L S+ + G++S EEL + +
Sbjct: 241 LNQAANLSGWDANNFRSEADLVKKIVKEVLTKLDSTHLSITEFPVGLESRVEELIE-FID 417
Query: 212 DSVDDVRVVGVCGMGGIGKKAIATALYNKIFHQFPVLFLIDDLRKIYRHD 261
D + V ++G+ GMGG GK A A+YN+I +F I+++R+I D
Sbjct: 418 DQSNKVCMIGIWGMGGSGKTTTAKAIYNQINRKFADRSFIENIREICEKD 567
>TC77728 similar to GP|12056928|gb|AAG48132.1 putative resistance protein
{Glycine max}, partial (12%)
Length = 765
Score = 58.9 bits (141), Expect = 2e-09
Identities = 54/178 (30%), Positives = 85/178 (47%), Gaps = 13/178 (7%)
Frame = +1
Query: 75 DSLIFIVVFSKNYASSTIGACKN*NPSFTALKSPKGMFCLFSMML-------ILMR*DIK 127
+S IFI +FS NYASS+ + + K CL + I + I
Sbjct: 247 ESRIFIPIFSANYASSSFCL----DELVHIIHCYKTKSCLVFPVFYDVEPTHIRNQSGIY 414
Query: 128 KEIMLKP---FSNMNKDSNKTLRWCKDGGKLKHKSPISGWDVRHNDAQ---IEKIVEEIM 181
E + K F N K+ + +W L + +SG+ + + IEKIVE+I
Sbjct: 415 GEHLTKHEERFQNNEKNMERLRQWKI---ALIQAANLSGYHYSPHGYEYKFIEKIVEDIS 585
Query: 182 NILGYKSTSLPNYLAGMDSLTEELEKHLLLDSVDDVRVVGVCGMGGIGKKAIATALYN 239
N + + ++ Y G+ S EE++ L + S D+VR+VG+ G GG+GK +A A+YN
Sbjct: 586 NNINHVFLNVAKYPVGLQSRIEEVKLLLDMGSEDEVRMVGLFGTGGMGKSTLAKAVYN 759
>BG588967 similar to GP|7107246|gb| unknown {Cicer arietinum}, partial (85%)
Length = 699
Score = 58.9 bits (141), Expect = 2e-09
Identities = 28/72 (38%), Positives = 46/72 (63%), Gaps = 1/72 (1%)
Frame = +3
Query: 197 GMDSLTEELEKHLLLDSVDDVRVVGVCGMGGIGKKAIATALYNKIFHQFPVLFLIDDLRK 256
G++S +E+ + L + ++ RV+G+CG GGIGK IA A+YNKI H F + ++R+
Sbjct: 126 GVESRVQEVIQLLNTEPSEETRVIGICGTGGIGKTTIAKAVYNKIHHHFEAKSFLLNVRQ 305
Query: 257 IYRHD-GPISAQ 267
++ D G +S Q
Sbjct: 306 VWEQDNGEVSLQ 341
>AW686165 weakly similar to GP|18033111|gb functional candidate resistance
protein KR1 {Glycine max}, partial (7%)
Length = 551
Score = 48.9 bits (115), Expect = 2e-06
Identities = 50/193 (25%), Positives = 79/193 (40%), Gaps = 28/193 (14%)
Frame = +2
Query: 75 DSLIFIVVFSKNYASSTIGACKN*NPSFTALKSPKGMFCLFSMMLILMR*DIKKEIML-- 132
+S IFI VFS NYASS FCL ++ I+ K ++L
Sbjct: 8 ESRIFIPVFSINYASSK--------------------FCLDELVHIIHCYKTKGRLVLPV 127
Query: 133 ------------------------KPFSNMNKDSNKTLRWCKDGGKLKHKSPISGWDVR- 167
+ F N K+ + +W L + +SG+
Sbjct: 128 FYGVDPTQIRHQSGSYGEHLTKHEESFQNNKKNKERLHQWKL---ALTQAANLSGYHYSP 298
Query: 168 -HNDAQIEKIVEEIMNILGYKSTSLPNYLAGMDSLTEELEKHLLLDSVDDVRVVGVCGMG 226
+ I KIVE+I N + + Y G++S E+++ L +S + V +VG+ G G
Sbjct: 299 GYEYKFIGKIVEDISNKINRVILHVAKYPVGLESRLEQVKLLLDKESDEGVHMVGLYGTG 478
Query: 227 GIGKKAIATALYN 239
G+GK +A A+YN
Sbjct: 479 GLGKSTLAKAIYN 517
Database: MTGI
Posted date: Oct 22, 2004 3:39 PM
Number of letters in database: 27,044,181
Number of sequences in database: 36,976
Lambda K H
0.339 0.149 0.474
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 8,254,218
Number of Sequences: 36976
Number of extensions: 107546
Number of successful extensions: 921
Number of sequences better than 10.0: 114
Number of HSP's better than 10.0 without gapping: 888
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 907
length of query: 267
length of database: 9,014,727
effective HSP length: 94
effective length of query: 173
effective length of database: 5,538,983
effective search space: 958244059
effective search space used: 958244059
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 15 ( 7.3 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 39 (21.8 bits)
S2: 57 (26.6 bits)
Medicago: description of AC137667.13


