

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
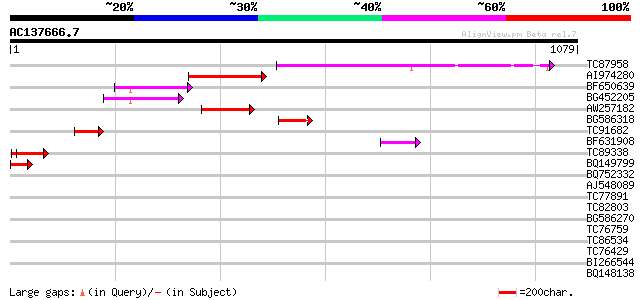
Query= AC137666.7 - phase: 0
(1079 letters)
Database: MTGI
36,976 sequences; 27,044,181 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
TC87958 weakly similar to GP|20279447|gb|AAM18727.1 putative TNP... 284 1e-76
AI974280 weakly similar to GP|20279447|g putative TNP2-like tran... 145 1e-34
BF650639 weakly similar to GP|22748425|g putative transposon pro... 128 1e-29
BG452205 weakly similar to GP|1345502|emb TNP2 {Antirrhinum maju... 125 7e-29
AW257182 weakly similar to GP|20279447|g putative TNP2-like tran... 105 9e-23
BG586318 similar to GP|6069576|dbj| transposon-like ORF {Brassic... 65 1e-10
TC91682 62 1e-09
BF631908 weakly similar to GP|21450441|gb| putative transposon p... 60 5e-09
TC89338 57 3e-08
BQ149799 50 5e-06
BQ752332 weakly similar to GP|8778366|gb| F15O4.30 {Arabidopsis ... 41 0.003
AJ548089 37 0.041
TC77891 weakly similar to SP|P93147|C81E_GLYEC Cytochrome P450 8... 36 0.070
TC82803 similar to PIR|D85049|D85049 probable transposon protein... 34 0.27
BG586270 30 5.0
TC76759 similar to GP|7677046|gb|AAF67003.1| putative Hs1pro-1 h... 30 5.0
TC86534 homologue to SP|P12227|RPOD_PEA DNA-directed RNA polymer... 30 5.0
TC76429 similar to GP|4056425|gb|AAC97999.1| ESTs gb|H36249 gb|... 30 6.6
BI266544 weakly similar to PIR|T01969|T019 potassium transport p... 29 8.6
BQ148138 weakly similar to GP|8778537|gb| F22G5.5 {Arabidopsis t... 29 8.6
>TC87958 weakly similar to GP|20279447|gb|AAM18727.1 putative TNP2-like
transposon protein {Oryza sativa (japonica
cultivar-group)}, partial (10%)
Length = 2433
Score = 284 bits (727), Expect = 1e-76
Identities = 181/553 (32%), Positives = 277/553 (49%), Gaps = 24/553 (4%)
Frame = +1
Query: 508 ELPYWKSLYVRHFLDVMHIEKNVFDSVVGTLLNIPGKSKDGINARLDMQSMGLRNELRPV 567
ELPYW S + H LDVMHIEKN+FD+++GTLL+IPGK+KD + R D++ MG+R L P
Sbjct: 1 ELPYWASNTLHHNLDVMHIEKNIFDNIIGTLLDIPGKTKDHVRGRYDLEEMGIRKNLHPR 180
Query: 568 KKDGKRTFLPPAAHTLSRKEKKIFCKFLHNVKVPEGYSSNIKSLVSMKDLKLKGLKSHDG 627
G R + + +++ +EK +FC L K+P+G +SNI V + + K+ G KSHD
Sbjct: 181 DVGGGRVAIANSCFSMTSEEKSMFCGVLKGAKLPDGSASNISRCVKVSEKKICGYKSHDA 360
Query: 628 HVLMGNFLPIAIRSILPENVRWTITKLCFFFKAISSKVI-DPEKLPTLQNEIVVTLCELE 686
H ++ L I IRS +P+ V + +L FF+++ K +P TL
Sbjct: 361 HFMLHYLLQIPIRSTMPKAVAQPLIRLSRFFRSLCQKGYPNP*SGQLTGRNC*STLFN*R 540
Query: 687 MYFPPSFFDIMVHLTVHLIMETQYCGPAYMRWMYPIERYMKILKGYVKNQSRPEGCIVER 746
+F F + M + RWMYP+ERY+ LKGYV+N+SRPEG I E
Sbjct: 541 WFFLLVFLMLWFT*LFIFRMRLDWVAQFNFRWMYPVERYLCKLKGYVRNRSRPEGSIAEG 720
Query: 747 YIVEEAVEFFTDFLSN------------------VESIGLPMSRHSGRTTGEGIGASKLV 788
Y+ EE + + ++ +E + GR G G K
Sbjct: 721 YVAEEGITICSRYMHKGVDTRSNRKNQNYDDSDLLEVDEIDYFSSIGRPLG-GKKNGKPF 897
Query: 789 TISDIELEQAHLYVLHNADEVEPYVKKHKEILQSLNPNRSENWLTREHNRRFISWLKEHI 848
+ L QAH Y+L N E+E Y+++H+ I+ S R + + F W K
Sbjct: 898 YLDFDSLAQAHRYLLFNCGEIEVYIREHEGIVNSHTKKRKWD-KAKAQCLNFSDWFKTR- 1071
Query: 849 FSEIAKNPDSISERLRWLANGPNANVLSYSSYVINKYTFYTKEHDQQSTMQNSGVTLVAQ 908
AK D + +LR + GPN +S Y++N Y F+T + D + QNSGVTLV+
Sbjct: 1072 ----AKR-DDVPPQLREFSRGPNNVAKRFSGYLVNGYRFHTMQ*DARRKTQNSGVTLVSL 1236
Query: 909 SLHISSAKDRNPAFAKLSYFGVIEHIWELD-YSSFQVLVFGCKWVDNNNGVQFDHGFMKV 967
+ +S+KD NP ++++G I I EL+ Y + ++F C W + + ++G V
Sbjct: 1237 TASFASSKDENPRTEPVNFYGAINDIIELNYYGHCKFVLFRCDWYEVE---EDEYGLTCV 1407
Query: 968 DLNREGYRDEPFILASQAHQVFYVTDPADEKWSIVLLSNKINDHHDQECENRDV----ED 1023
N++ Y+D+ F++ASQ Q FYV DP + +N H+ + + RD+
Sbjct: 1408 YFNKKCYQDDAFVIASQVQQCFYVQDPLN-----------VNRHYALKTDPRDLFKMGNQ 1554
Query: 1024 DPFFNTSHSLEDE 1036
F TS + DE
Sbjct: 1555 SEFEATSVNAADE 1593
>AI974280 weakly similar to GP|20279447|g putative TNP2-like transposon
protein {Oryza sativa (japonica cultivar-group)},
partial (8%)
Length = 457
Score = 145 bits (365), Expect = 1e-34
Identities = 71/150 (47%), Positives = 96/150 (63%)
Frame = +1
Query: 340 KFMMLTMLISGPTQPGNDIDIYLAPLIEDLKHMWETGVEVYDEYKKESFILRAMLFGTIN 399
+ ML++LI GP+ P +ID+YL PLI++LK +WE G EVYD K+ F A L TI+
Sbjct: 4 RHQMLSLLIPGPSSPKRNIDVYLEPLIDELKELWERGFEVYDASSKQLFEAHAALLWTIS 183
Query: 400 DFPAYGNLSGYSVKGKCACPVCEDHTNWIRLEHCQKNVFLGHRRFLNTKHHYRKWKKAFN 459
D PAY LSG+S GK ACPVC T + L+H +K ++ HRRFL+ KH +RK KK F+
Sbjct: 184 DLPAYTMLSGWSTSGKLACPVCSYDTCSMYLKHSRKTCYMSHRRFLDPKHRWRKDKKGFD 363
Query: 460 GESEEDRAPSPLTGDQLYEKVKLLSSKFGK 489
G+ E AP PL+G + K+ + FGK
Sbjct: 364 GKKEIRVAPFPLSGSAILRKLGNRQNLFGK 453
>BF650639 weakly similar to GP|22748425|g putative transposon protein {Oryza
sativa (japonica cultivar-group)}, partial (6%)
Length = 568
Score = 128 bits (322), Expect = 1e-29
Identities = 61/162 (37%), Positives = 91/162 (55%), Gaps = 13/162 (8%)
Frame = +2
Query: 199 KIHACPNDCILFQNEYASLNMCPKCSASRY-------------KKKETGPSKVLWYFPII 245
KI AC NDC+L+ ++ + C C ASR KK+ +K+L +FP+I
Sbjct: 71 KIDACXNDCMLYWKDHKNDISCHVCGASRXIESTTEXGQIEENKKEHKVSTKILRHFPLI 250
Query: 246 PRFRRMYRNAEDAKNLTWHANERVNDGMLRHPADSPQWAKIDNDYSDFGKETRNLRLALS 305
P +R + +E A +L WH ++ DG LRHPAD W + + +F +++RN+ L L+
Sbjct: 251 PXLQRXFMCSETATSLRWHHDDXSKDGKLRHPADGQAWKDFNGHHPEFARDSRNIXLGLA 430
Query: 306 TDGINPHGLQSGKHTTWPVILLIYNLPPWLCMKRKFMMLTML 347
+DG NP + K++ WP IL+ YN PPWLCMK + ML
Sbjct: 431 SDGFNPFRAMNLKYSAWPXILIPYNFPPWLCMKAXYTNHRML 556
>BG452205 weakly similar to GP|1345502|emb TNP2 {Antirrhinum majus}, partial
(16%)
Length = 619
Score = 125 bits (315), Expect = 7e-29
Identities = 58/165 (35%), Positives = 95/165 (57%), Gaps = 13/165 (7%)
Frame = +1
Query: 179 LPSRTYEAKRLLCSIGMSYEKIHACPNDCILFQNEYASLNMCPKCSASRY---------- 228
+P + + ++ +G+ Y KI ACPNDC+L+ ++ + C C ASR+
Sbjct: 121 VPDSYNKTR*MIRDLGLDYSKIDACPNDCMLYWKDHKNDISCHVCGASRWIESTTESGQI 300
Query: 229 ---KKKETGPSKVLWYFPIIPRFRRMYRNAEDAKNLTWHANERVNDGMLRHPADSPQWAK 285
KK+ +K+L +FP+IPR +R++ +E A +L WH ++ DG LRHPAD W
Sbjct: 301 EENKKEHKVSTKILRHFPLIPRLQRLFMCSETATSLRWHHDDCSKDGKLRHPADGQAWKD 480
Query: 286 IDNDYSDFGKETRNLRLALSTDGINPHGLQSGKHTTWPVILLIYN 330
+ + +F +++RN+RL L++DG NP + K+ WPV L+ YN
Sbjct: 481 FNGHHPEFARDSRNIRLGLASDGFNPFRAMNLKYXAWPVXLIPYN 615
>AW257182 weakly similar to GP|20279447|g putative TNP2-like transposon
protein {Oryza sativa (japonica cultivar-group)},
partial (6%)
Length = 387
Score = 105 bits (262), Expect = 9e-23
Identities = 49/100 (49%), Positives = 68/100 (68%)
Frame = +1
Query: 366 IEDLKHMWETGVEVYDEYKKESFILRAMLFGTINDFPAYGNLSGYSVKGKCACPVCEDHT 425
IE+LK +WE+GVE YD K + F +RA L TI+DFPAY LSG+S KGK ACP C ++T
Sbjct: 7 IEELKVLWESGVETYDASKNQMFRMRAALLWTISDFPAYAMLSGWSTKGKLACPCCNNNT 186
Query: 426 NWIRLEHCQKNVFLGHRRFLNTKHHYRKWKKAFNGESEED 465
+ L+H +K ++ HR FL H +R+ +K+FNG+ D
Sbjct: 187 SSTYLKHSRKMCYMDHRTFLPMDHAWRENRKSFNGKKNLD 306
>BG586318 similar to GP|6069576|dbj| transposon-like ORF {Brassica rapa},
partial (11%)
Length = 267
Score = 65.1 bits (157), Expect = 1e-10
Identities = 30/64 (46%), Positives = 46/64 (71%)
Frame = +3
Query: 512 WKSLYVRHFLDVMHIEKNVFDSVVGTLLNIPGKSKDGINARLDMQSMGLRNELRPVKKDG 571
W+ +RH LDVMHIEKN FD+++ T+LN+ GK+KD + +RLD+ + R+EL V ++G
Sbjct: 3 WEDHLLRHNLDVMHIEKNFFDNLMNTILNVQGKTKDNLKSRLDLVDICARSELH-VDENG 179
Query: 572 KRTF 575
+ F
Sbjct: 180 RAPF 191
>TC91682
Length = 1093
Score = 62.0 bits (149), Expect = 1e-09
Identities = 30/56 (53%), Positives = 38/56 (67%)
Frame = +2
Query: 123 RVVIDAETPLYEGCTKFTRLSTILKLYKLKAGNGWSDRSFTDLLTLLKDILPKNNV 178
R+V D++ PLY+GC KFTRLST+LKLY LKA NGWS L +K +P +V
Sbjct: 2 RLVRDSKIPLYDGCAKFTRLSTVLKLYNLKARNGWS**ELYRLANSIKRYMPGTSV 169
>BF631908 weakly similar to GP|21450441|gb| putative transposon protein
5'-partial {Oryza sativa (japonica cultivar-group)},
partial (17%)
Length = 681
Score = 60.1 bits (144), Expect = 5e-09
Identities = 29/77 (37%), Positives = 46/77 (59%), Gaps = 1/77 (1%)
Frame = +3
Query: 707 ETQYCGPAYMRWMYPIERYMKILKGYVKNQSRPEGCIVERYIVEEAVEFFTDFLSN-VES 765
E + GP + RWMYPIER++ LK YV+N++ PEG I E Y+ E + + ++S+ V++
Sbjct: 435 EARIAGPVHYRWMYPIERFILTLKNYVRNRNYPEGSIAEGYVANECLTHCSRYMSDGVQT 614
Query: 766 IGLPMSRHSGRTTGEGI 782
R+ GEG+
Sbjct: 615 KFNKPPRNLDGPVGEGV 665
>TC89338
Length = 1088
Score = 57.4 bits (137), Expect = 3e-08
Identities = 29/75 (38%), Positives = 46/75 (60%), Gaps = 3/75 (4%)
Frame = +3
Query: 3 RKWMSANRASKEYEDGVQEFVRFAIANAEDTSKII-CPCLECCYTN--VSANDLQDHLIC 59
++WM A R S EY++GV EF++FA N +T + CPC+ C + +S +++ HL+
Sbjct: 222 KRWMKAYRLSAEYDNGVTEFLQFAEKNLPNTEGLFPCPCVSCGNRDPKLSKEEIKGHLVS 401
Query: 60 NGIDKSYTCWTMHGE 74
GI ++YT HGE
Sbjct: 402 VGICQNYTQ*IWHGE 446
Score = 50.4 bits (119), Expect = 4e-06
Identities = 24/64 (37%), Positives = 40/64 (62%), Gaps = 3/64 (4%)
Frame = +3
Query: 14 EYEDGVQEFVRFAIANAEDTSKII-CPCLECCYTN--VSANDLQDHLICNGIDKSYTCWT 70
EY++G+ EF++FA N +T + CPC+ C + +S +++ HL+ GI ++YT W
Sbjct: 498 EYDNGMTEFLQFAEKNLPNTEGLFPCPCISCGNWDPKLSKEEIRGHLVSVGICQNYTEWI 677
Query: 71 MHGE 74
HGE
Sbjct: 678 WHGE 689
>BQ149799
Length = 437
Score = 50.1 bits (118), Expect = 5e-06
Identities = 22/44 (50%), Positives = 30/44 (68%), Gaps = 1/44 (2%)
Frame = +1
Query: 1 MDRKWMSANRASKEYEDGVQEFVRFAIANAEDTSKII-CPCLEC 43
MDR+WM A R S EY++GV EF++FA N +T + CPC+ C
Sbjct: 259 MDRRWMKAYRLSVEYDNGVTEFLQFAEKNLPNTEGLFPCPCVSC 390
>BQ752332 weakly similar to GP|8778366|gb| F15O4.30 {Arabidopsis thaliana},
partial (6%)
Length = 268
Score = 40.8 bits (94), Expect = 0.003
Identities = 25/74 (33%), Positives = 42/74 (55%), Gaps = 6/74 (8%)
Frame = -2
Query: 927 YFGVIEHIWELDYSSF---QVLVFGCKWVDN--NNGVQ-FDHGFMKVDLNREGYRDEPFI 980
++G++ I E+++ + ++F C+W D N GV+ G + V+ R + EPFI
Sbjct: 264 FYGILTEIIEVEFPGILKLKCVLFKCEWFDPVVNRGVRSIKFGVVDVNGGRRYNKFEPFI 85
Query: 981 LASQAHQVFYVTDP 994
LASQA QV ++ P
Sbjct: 84 LASQADQVSFLPYP 43
>AJ548089
Length = 560
Score = 37.0 bits (84), Expect = 0.041
Identities = 18/55 (32%), Positives = 32/55 (57%), Gaps = 3/55 (5%)
Frame = +1
Query: 23 VRFAIANAEDTSKII-CPCLECCYTN--VSANDLQDHLICNGIDKSYTCWTMHGE 74
++FA N +T + CPC+ C + +S +++ HL+ GI ++YT W +GE
Sbjct: 193 IKFAEKNLPNTEGLFPCPCISCGNWDPKLSKEEIRGHLVSVGICQNYTEWIWYGE 357
>TC77891 weakly similar to SP|P93147|C81E_GLYEC Cytochrome P450 81E1 (EC
1.14.-.-) (Isoflavone 2'-hydroxylase) (P450 91A4) (CYP
GE-3). [Licorice], partial (91%)
Length = 1707
Score = 36.2 bits (82), Expect = 0.070
Identities = 15/26 (57%), Positives = 20/26 (76%)
Frame = -1
Query: 327 LIYNLPPWLCMKRKFMMLTMLISGPT 352
+IYNL WLC+KRK +ML+M+I T
Sbjct: 1707 VIYNLSSWLCIKRKHVMLSMIIQTNT 1630
>TC82803 similar to PIR|D85049|D85049 probable transposon protein [imported]
- Arabidopsis thaliana, partial (5%)
Length = 755
Score = 34.3 bits (77), Expect = 0.27
Identities = 17/40 (42%), Positives = 23/40 (57%)
Frame = +3
Query: 694 FDIMVHLTVHLIMETQYCGPAYMRWMYPIERYMKILKGYV 733
FD M HL++HL E + P R MYP ER I++ Y+
Sbjct: 288 FDSMEHLSIHLAYEARVGNPVQYR*MYPFERL--IIQPYI 401
>BG586270
Length = 822
Score = 30.0 bits (66), Expect = 5.0
Identities = 15/40 (37%), Positives = 20/40 (49%), Gaps = 2/40 (5%)
Frame = -2
Query: 426 NWIRLEHCQKNVFLGHRRFLNTKHHYRKW--KKAFNGESE 463
+W +HC +F RRF HY +W KKA E+E
Sbjct: 620 SWHGRKHCSHVIFYYRRRFQRPDTHYLEWAPKKAAEYEAE 501
>TC76759 similar to GP|7677046|gb|AAF67003.1| putative Hs1pro-1 homolog
{Pisum sativum}, partial (95%)
Length = 1800
Score = 30.0 bits (66), Expect = 5.0
Identities = 15/37 (40%), Positives = 21/37 (56%)
Frame = +1
Query: 299 NLRLALSTDGINPHGLQSGKHTTWPVILLIYNLPPWL 335
+L L++ST N H S +H T IL++ NL WL
Sbjct: 13 SLPLSISTQQHNKHNSISTQHKTNTTILIVLNLQRWL 123
>TC86534 homologue to SP|P12227|RPOD_PEA DNA-directed RNA polymerase beta'
chain (EC 2.7.7.6) (Fragment). [Garden pea] {Pisum
sativum}, partial (61%)
Length = 2441
Score = 30.0 bits (66), Expect = 5.0
Identities = 16/57 (28%), Positives = 28/57 (49%)
Frame = -3
Query: 477 YEKVKLLSSKFGKPFSGELVTGGWKKKSIFFELPYWKSLYVRHFLDVMHIEKNVFDS 533
Y ++ +G+ F + G KKKSI+F+L +W + L+ + KN +S
Sbjct: 1731 YSNETIIPYHYGRRFG*NMHFFGNKKKSIYFDLVFWL*FFDSSILNEIFFLKNGMNS 1561
>TC76429 similar to GP|4056425|gb|AAC97999.1| ESTs gb|H36249 gb|AA59732 and
gb|AA651219 come from this gene. {Arabidopsis thaliana},
partial (55%)
Length = 666
Score = 29.6 bits (65), Expect = 6.6
Identities = 21/82 (25%), Positives = 37/82 (44%), Gaps = 3/82 (3%)
Frame = +3
Query: 878 SSYVINKYTFYTKEHDQQSTMQNSGVTLVAQSLHISSAKDRNPAFAKLSYFGVIEHIWEL 937
S + +++ FYT + +Q V S+ I+ +R E ++ L
Sbjct: 3 SIFFLSRLNFYTLQSILSKILQTFFSLTVPSSITITDPSNRRSH----------ESLFAL 152
Query: 938 DYSSFQVLVFGCKWV---DNNN 956
SSF +L+FG K++ DN+N
Sbjct: 153 TLSSFTLLIFGAKFIMKLDNSN 218
>BI266544 weakly similar to PIR|T01969|T019 potassium transport protein homolog
T9A4.5 - Arabidopsis thaliana, partial (12%)
Length = 373
Score = 29.3 bits (64), Expect = 8.6
Identities = 12/23 (52%), Positives = 17/23 (73%)
Frame = -2
Query: 1054 WINPSFWVRKKQKTRTRITRKRK 1076
W N SFW+RK + T++T+KRK
Sbjct: 264 WFNRSFWLRKLE---TQVTKKRK 205
>BQ148138 weakly similar to GP|8778537|gb| F22G5.5 {Arabidopsis thaliana},
partial (25%)
Length = 566
Score = 29.3 bits (64), Expect = 8.6
Identities = 11/43 (25%), Positives = 20/43 (45%)
Frame = +3
Query: 891 EHDQQSTMQNSGVTLVAQSLHISSAKDRNPAFAKLSYFGVIEH 933
EH + +G+ + + LH K N A+++Y G + H
Sbjct: 378 EHSSSAAKPGTGIAVAVKRLHQDCFKGHNKFMAEVNYLGQLSH 506
Database: MTGI
Posted date: Oct 22, 2004 3:39 PM
Number of letters in database: 27,044,181
Number of sequences in database: 36,976
Lambda K H
0.320 0.137 0.425
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 38,188,674
Number of Sequences: 36976
Number of extensions: 598448
Number of successful extensions: 3037
Number of sequences better than 10.0: 43
Number of HSP's better than 10.0 without gapping: 2988
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 3030
length of query: 1079
length of database: 9,014,727
effective HSP length: 106
effective length of query: 973
effective length of database: 5,095,271
effective search space: 4957698683
effective search space used: 4957698683
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.8 bits)
S2: 64 (29.3 bits)
Medicago: description of AC137666.7


