

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC137553.4 - phase: 0 /pseudo
(921 letters)
Database: MTGI
36,976 sequences; 27,044,181 total letters
Searching..................................................done
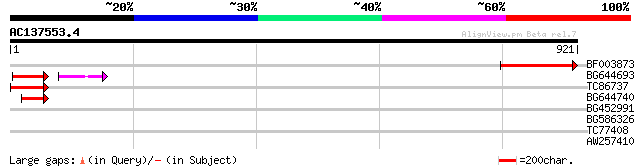
Score E
Sequences producing significant alignments: (bits) Value
BF003873 similar to GP|14715222|em putative polyprotein {Cicer a... 195 8e-50
BG644693 weakly similar to GP|18767374|g Putative 22 kDa kafirin... 62 6e-13
TC86737 weakly similar to GP|6683624|dbj|BAA89272.1 Pol {Alterna... 62 1e-09
BG644740 similar to PIR|A84460|A84 probable retroelement pol pol... 58 2e-08
BG452991 PIR|A25875|A25 histone H4 - Tetrahymena thermophila, pa... 41 0.002
BG586326 similar to PIR|G84493|G8 probable retroelement pol poly... 33 0.003
TC77408 similar to GP|12006855|gb|AAG44951.1 POZ/BTB containing-... 30 4.2
AW257410 weakly similar to GP|10177533|db selenium-binding prote... 29 7.2
>BF003873 similar to GP|14715222|em putative polyprotein {Cicer arietinum},
partial (82%)
Length = 559
Score = 195 bits (495), Expect = 8e-50
Identities = 100/124 (80%), Positives = 109/124 (87%)
Frame = +1
Query: 798 WTCFEVKEVDSEIYWSVSDIGKSWNGGVSSWITTASFEFARYITCVATSKVCTGSISCDT 857
WTCFEVKEVD EI+WSVSDI KSWNGGVSS ITTASFEFA ++CVATS+VC+GSISCD
Sbjct: 7 WTCFEVKEVDCEIHWSVSDIRKSWNGGVSSGITTASFEFA*CLSCVATSEVCSGSISCDP 186
Query: 858 E**CAS*RQPYGSD*TGKD*RSKSEDVERQENTPC*SRLGWSDW*KLDVGA*E*DAGVLS 917
E**CAS RQPYG D TG+D* SKSED+ERQ +T C SRLG S+W*KLDVGA*E*D GVLS
Sbjct: 187 E**CASQRQPYGRDFTGED**SKSEDIERQGDTSCESRLGQSEW*KLDVGA*E*DGGVLS 366
Query: 918 RVVC 921
RVVC
Sbjct: 367 RVVC 378
>BG644693 weakly similar to GP|18767374|g Putative 22 kDa kafirin cluster;
Ty3-Gypsy type {Oryza sativa}, partial (15%)
Length = 716
Score = 62.4 bits (150), Expect(2) = 6e-13
Identities = 32/59 (54%), Positives = 41/59 (69%)
Frame = +2
Query: 5 ELAELKKQLEDLVEMKFFRPSVSPWGAPVLLVKKKDGSMRLRIDYRQLNKVTIKNRYLL 63
+L LK QL+DL+E F +PS+ P G VL +KKKDG +R+ IDY QLN V IK +Y L
Sbjct: 107 KLKVLKLQLKDLLEKGFIQPSIYP*GVVVLFLKKKDGFLRMSIDYPQLNNVNIKIKYPL 283
Score = 30.4 bits (67), Expect(2) = 6e-13
Identities = 31/83 (37%), Positives = 43/83 (51%), Gaps = 3/83 (3%)
Frame = +3
Query: 80 ARLI*GKVITRLK*KMKICRRQLSERGMVTMN---IMLCRSV*PMHLECFWSI*SASSMH 136
+RL *G T + * +K+ + LSE MVTM +L + + HL W+ * S
Sbjct: 333 SRLT*GWDNTNIG**VKMFLKLLSELDMVTMKS**CLLGKQIPRWHL---WN**IEFSKI 503
Query: 137 FWIGSWLFSSMIF*FTPRLKKNM 159
LF MIF*+TPR+K +M
Sbjct: 504 TSTH**LFLVMIF*YTPRMKMSM 572
>TC86737 weakly similar to GP|6683624|dbj|BAA89272.1 Pol {Alternaria
alternata}, partial (21%)
Length = 1540
Score = 61.6 bits (148), Expect = 1e-09
Identities = 33/63 (52%), Positives = 41/63 (64%)
Frame = +1
Query: 1 MSVSELAELKKQLEDLVEMKFFRPSVSPWGAPVLLVKKKDGSMRLRIDYRQLNKVTIKNR 60
MS EL LKK LEDL++ F + S S GAPVL V+K G +R +DYR LN +T K+R
Sbjct: 487 MSRQELLVLKKTLEDLLDKGFIKASGSAAGAPVLFVRKPGGGIRFCVDYRALNAITKKDR 666
Query: 61 YLL 63
Y L
Sbjct: 667 YPL 675
>BG644740 similar to PIR|A84460|A84 probable retroelement pol polyprotein
[imported] - Arabidopsis thaliana, partial (4%)
Length = 754
Score = 57.8 bits (138), Expect = 2e-08
Identities = 27/44 (61%), Positives = 34/44 (76%)
Frame = -1
Query: 20 KFFRPSVSPWGAPVLLVKKKDGSMRLRIDYRQLNKVTIKNRYLL 63
+F +PS+SP GA +L V+KKDG R+ IDYRQ NKVT KN+Y L
Sbjct: 271 RFQQPSISP*GAALLFVRKKDGYFRMCIDYRQFNKVTTKNKYPL 140
>BG452991 PIR|A25875|A25 histone H4 - Tetrahymena thermophila, partial (33%)
Length = 560
Score = 41.2 bits (95), Expect = 0.002
Identities = 28/58 (48%), Positives = 35/58 (60%)
Frame = +2
Query: 421 TV*RYEFGV*TAPSQCETGNAED***ISEEY*RGTES*CQVCRLVGC**LD*RW*L*S 478
TV R EFG+*+ ++CE G+ ED I Y RG+E C+V R * D*RW* *S
Sbjct: 74 TVQRLEFGL*SVATECEIGDVEDQQRIFG*YQRGSEGGCEVGRFNVWE*SD*RW*F*S 247
>BG586326 similar to PIR|G84493|G8 probable retroelement pol polyprotein
[imported] - Arabidopsis thaliana, partial (13%)
Length = 736
Score = 33.1 bits (74), Expect(2) = 0.003
Identities = 12/23 (52%), Positives = 19/23 (82%)
Frame = +2
Query: 350 RFEVFSDHKTLKYLFDQKELNVR 372
+ ++ +DHK+LKY+F Q ELN+R
Sbjct: 230 KVQIHTDHKSLKYIFTQPELNLR 298
Score = 26.2 bits (56), Expect(2) = 0.003
Identities = 19/37 (51%), Positives = 21/37 (56%)
Frame = +3
Query: 315 QDS*QFMRRITRRTI*S*LL*SLY*RYGDITCMVRRF 351
Q S*+ MR T I* L * *R+G TCMV RF
Sbjct: 126 QGS*ENMRETTPPMI*KWLR*YSP*RFGAHTCMVPRF 236
>TC77408 similar to GP|12006855|gb|AAG44951.1 POZ/BTB containing-protein
AtPOB1 {Arabidopsis thaliana}, partial (85%)
Length = 2287
Score = 30.0 bits (66), Expect = 4.2
Identities = 20/61 (32%), Positives = 34/61 (54%)
Frame = +3
Query: 512 EYSPGSYEDVSRSKEEILVVWIEARCGTVCVLLFDLLEVEN*ASETCWYDGVLRCARVEM 571
+Y Y+D+++ +EE++ + + G +L D L+V ASE YD VL+ AR +
Sbjct: 918 QYLASRYKDITKFQEEVMNLPL---AGVEAILSSDDLQV---ASEDAVYDFVLKWARHQY 1079
Query: 572 G 572
G
Sbjct: 1080G 1082
>AW257410 weakly similar to GP|10177533|db selenium-binding protein-like
{Arabidopsis thaliana}, partial (21%)
Length = 724
Score = 29.3 bits (64), Expect = 7.2
Identities = 14/33 (42%), Positives = 18/33 (54%)
Frame = +1
Query: 543 LLFDLLEVEN*ASETCWYDGVLRCARVEMGYYI 575
+LFDLL + S DG ++C VEM Y I
Sbjct: 445 VLFDLLPSRDIVSWNTMIDGYIKCGNVEMAYKI 543
Database: MTGI
Posted date: Oct 22, 2004 3:39 PM
Number of letters in database: 27,044,181
Number of sequences in database: 36,976
Lambda K H
0.366 0.163 0.618
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 27,731,151
Number of Sequences: 36976
Number of extensions: 391422
Number of successful extensions: 5084
Number of sequences better than 10.0: 16
Number of HSP's better than 10.0 without gapping: 1051
Number of HSP's successfully gapped in prelim test: 128
Number of HSP's that attempted gapping in prelim test: 3959
Number of HSP's gapped (non-prelim): 1323
length of query: 921
length of database: 9,014,727
effective HSP length: 105
effective length of query: 816
effective length of database: 5,132,247
effective search space: 4187913552
effective search space used: 4187913552
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 14 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 36 (21.6 bits)
S2: 63 (28.9 bits)
Medicago: description of AC137553.4


