

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
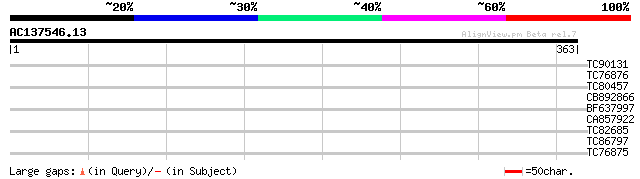
Query= AC137546.13 - phase: 0
(363 letters)
Database: MTGI
36,976 sequences; 27,044,181 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
TC90131 weakly similar to GP|21536774|gb|AAM61106.1 unknown {Ara... 28 4.3
TC76876 similar to GP|20453070|gb|AAM19780.1 AT5g66920/MUD21_18 ... 28 4.3
TC80457 similar to GP|6682258|gb|AAF23310.1| putative RNA helica... 28 5.6
CB892866 similar to GP|12325096|gb unknown protein; 30607-27264 ... 28 7.3
BF637997 similar to PIR|C84514|C845 hypothetical protein At2g141... 27 9.5
CA857922 similar to PIR|T47793|T47 receptor-like protein kinase ... 27 9.5
TC82685 similar to GP|23237938|dbj|BAC16511. contains ESTs C2654... 27 9.5
TC86797 similar to GP|1311386|pdb|1CBG| Cyanogenic Beta-Glucosid... 27 9.5
TC76875 weakly similar to GP|21554666|gb|AAM63650.1 putative ace... 27 9.5
>TC90131 weakly similar to GP|21536774|gb|AAM61106.1 unknown {Arabidopsis
thaliana}, partial (29%)
Length = 1330
Score = 28.5 bits (62), Expect = 4.3
Identities = 30/132 (22%), Positives = 47/132 (34%), Gaps = 14/132 (10%)
Frame = +2
Query: 10 VLKTLRGSSDDIGWLQHAPGMPPVHDGSSRFLELLSDIRNGKDSIPSSFVYLLIPGL--- 66
+ K+L SS + W ++ DG+ S I SIP + VY L +
Sbjct: 560 ISKSLFASSTNDDWDENFKNTLEPGDGNEDGPSYWSSIGQSDPSIPEALVYKLCSNICLV 739
Query: 67 -----------FSNHGPLYFVATKRFFSKMGLACHIAKVHSEASVEHNAMEIKQYIEEIY 115
F + P+Y RF ++G H +V S V + + E+
Sbjct: 740 SEIHIQPYQDYFEDDCPIYSAKAVRF--RLGCERHDMEVESNTVVPDSL----TFNEDFK 901
Query: 116 WGSGKPVMLLGH 127
W PV + H
Sbjct: 902 WTYTSPVFPMSH 937
>TC76876 similar to GP|20453070|gb|AAM19780.1 AT5g66920/MUD21_18 {Arabidopsis
thaliana}, partial (86%)
Length = 3027
Score = 28.5 bits (62), Expect = 4.3
Identities = 18/69 (26%), Positives = 34/69 (49%)
Frame = -3
Query: 90 HIAKVHSEASVEHNAMEIKQYIEEIYWGSGKPVMLLGHSKGGIDAAAALSLYWSDLKGKV 149
H +VH SV + + +++ + + V+L+GHS GG+ + A+ L+ + V
Sbjct: 2890 HPKQVHELDSVTYYYEPLIEFLRSLR--QDQRVILVGHSLGGMCISVAMELFPKKIAAAV 2717
Query: 150 AGLALVQSP 158
A + SP
Sbjct: 2716 FVTAFMPSP 2690
>TC80457 similar to GP|6682258|gb|AAF23310.1| putative RNA helicase
{Arabidopsis thaliana}, partial (28%)
Length = 1425
Score = 28.1 bits (61), Expect = 5.6
Identities = 23/66 (34%), Positives = 33/66 (49%), Gaps = 3/66 (4%)
Frame = +1
Query: 54 IPS-SFVYLLIPGLFSNHGPLYFVATKRFFSKMGLACHIAKVHSEASVEH--NAMEIKQY 110
IPS S V +I + +N G AT+ + M L +IAK+ SEA EH +EI +
Sbjct: 541 IPSVSTVLPVIGSIATNDG-----ATRAALAAMNLQQNIAKIQSEALPEHYEAELEINDF 705
Query: 111 IEEIYW 116
+ W
Sbjct: 706 PQNARW 723
>CB892866 similar to GP|12325096|gb unknown protein; 30607-27264 {Arabidopsis
thaliana}, partial (25%)
Length = 395
Score = 27.7 bits (60), Expect = 7.3
Identities = 11/24 (45%), Positives = 15/24 (61%)
Frame = -3
Query: 25 QHAPGMPPVHDGSSRFLELLSDIR 48
Q AP PP+HD S +L+ D+R
Sbjct: 291 QDAPHCPPLHDRQSSYLQFQIDLR 220
>BF637997 similar to PIR|C84514|C845 hypothetical protein At2g14110
[imported] - Arabidopsis thaliana, partial (60%)
Length = 655
Score = 27.3 bits (59), Expect = 9.5
Identities = 11/33 (33%), Positives = 19/33 (57%)
Frame = +1
Query: 73 LYFVATKRFFSKMGLACHIAKVHSEASVEHNAM 105
L+F+ + F+S H K+HS+ V +N+M
Sbjct: 514 LFFIV*EIFYSSTHKTEHFQKIHSKTGVPYNSM 612
>CA857922 similar to PIR|T47793|T47 receptor-like protein kinase -
Arabidopsis thaliana, partial (47%)
Length = 810
Score = 27.3 bits (59), Expect = 9.5
Identities = 16/37 (43%), Positives = 19/37 (51%)
Frame = -1
Query: 44 LSDIRNGKDSIPSSFVYLLIPGLFSNHGPLYFVATKR 80
L RNG S S LI GLFS G ++ AT+R
Sbjct: 699 LRSSRNGYSSASSILTT*LILGLFSASGSIHLSATRR 589
>TC82685 similar to GP|23237938|dbj|BAC16511. contains ESTs C26549(C12568)
AU092341(C12568)~unknown protein, partial (9%)
Length = 1373
Score = 27.3 bits (59), Expect = 9.5
Identities = 12/33 (36%), Positives = 19/33 (57%)
Frame = -2
Query: 226 SEASITPSVLATMTQIAHAELPRLILPKFGSKV 258
S S +PS + T Q A +PR+ LP F +++
Sbjct: 733 SSISSSPSEVVTEIQSAKVSVPRVFLPNFSNRL 635
>TC86797 similar to GP|1311386|pdb|1CBG| Cyanogenic Beta-Glucosidase Mol_id:
1; Molecule: Cyanogenic Beta-Glucosidase; Chain: Null;
Ec: 3.2., partial (97%)
Length = 1517
Score = 27.3 bits (59), Expect = 9.5
Identities = 10/17 (58%), Positives = 12/17 (69%)
Frame = +1
Query: 319 HAWMVYSSNSKKKKSSE 335
H WMVY SN K+K S +
Sbjct: 856 HVWMVYGSNHKRKLSKK 906
>TC76875 weakly similar to GP|21554666|gb|AAM63650.1 putative
acetone-cyanohydrin lyase {Arabidopsis thaliana},
partial (62%)
Length = 1020
Score = 27.3 bits (59), Expect = 9.5
Identities = 18/69 (26%), Positives = 33/69 (47%)
Frame = +1
Query: 90 HIAKVHSEASVEHNAMEIKQYIEEIYWGSGKPVMLLGHSKGGIDAAAALSLYWSDLKGKV 149
H +VH SV + +++ + + V+L+GHS GG+ + A+ L+ + V
Sbjct: 229 HPKQVHELDSVTDYYEPLIEFLRSL--PQDQRVILVGHSLGGMCISVAMELFPKKIAAAV 402
Query: 150 AGLALVQSP 158
A + SP
Sbjct: 403 FVTAFMPSP 429
Database: MTGI
Posted date: Oct 22, 2004 3:39 PM
Number of letters in database: 27,044,181
Number of sequences in database: 36,976
Lambda K H
0.320 0.136 0.396
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 10,063,656
Number of Sequences: 36976
Number of extensions: 132025
Number of successful extensions: 636
Number of sequences better than 10.0: 18
Number of HSP's better than 10.0 without gapping: 634
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 636
length of query: 363
length of database: 9,014,727
effective HSP length: 97
effective length of query: 266
effective length of database: 5,428,055
effective search space: 1443862630
effective search space used: 1443862630
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.8 bits)
S2: 59 (27.3 bits)
Medicago: description of AC137546.13


