

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC137081.4 - phase: 0
(325 letters)
Database: MTGI
36,976 sequences; 27,044,181 total letters
Searching..................................................done
Score E
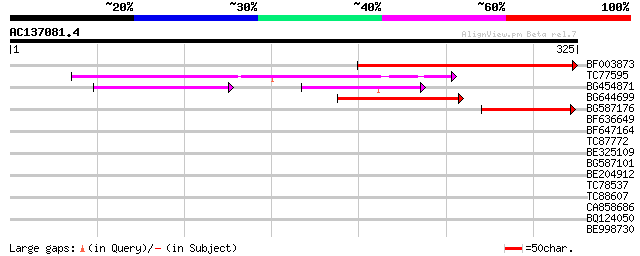
Sequences producing significant alignments: (bits) Value
BF003873 similar to GP|14715222|em putative polyprotein {Cicer a... 224 3e-59
TC77595 weakly similar to PIR|T18350|T18350 probable pol polypro... 87 9e-18
BG454871 weakly similar to GP|10140673|g putative gag-pol polypr... 64 8e-15
BG644699 similar to PIR|T07863|T078 probable polyprotein - pinea... 64 8e-11
BG587176 weakly similar to PIR|G84493|G84 probable retroelement ... 55 5e-08
BF636649 similar to GP|21628724|emb OSJNBa0033H08.7 {Oryza sativ... 35 0.051
BF647164 weakly similar to GP|6691191|gb F7F22.15 {Arabidopsis t... 30 0.97
TC87772 weakly similar to GP|8574455|gb|AAF77578.1| pepper ester... 29 2.2
BE325109 similar to PIR|T04011|T040 hypothetical protein T5L19.2... 29 2.2
BG587101 similar to GP|6691191|gb F7F22.15 {Arabidopsis thaliana... 28 3.7
BE204912 similar to PIR|T02686|T026 probable Ser/Thr protein kin... 27 8.2
TC78537 similar to GP|10178176|dbj|BAB11650. trehalose-6-phospha... 27 8.2
TC88607 similar to GP|8777331|dbj|BAA96921.1 receptor-like prote... 27 8.2
CA858686 homologue to GP|8489871|gb| DEAD box protein P68 {Pisum... 27 8.2
BQ124050 similar to PIR|T50014|T50 trehalose-6-phosphate phospha... 27 8.2
BE998730 27 8.2
>BF003873 similar to GP|14715222|em putative polyprotein {Cicer arietinum},
partial (82%)
Length = 559
Score = 224 bits (572), Expect = 3e-59
Identities = 111/126 (88%), Positives = 117/126 (92%)
Frame = +2
Query: 200 GVGRALKSRKLTPKFIGPYQISERVGTVAYRVGLPPHLSNLHDVFHVSQLRKYVADPSHV 259
GVGRALKS+KLT +FIGPYQISERVGTVAYRVGLPPHL NLHDVFHVSQLRKYV DPSHV
Sbjct: 2 GVGRALKSKKLTVRFIGPYQISERVGTVAYRVGLPPHLLNLHDVFHVSQLRKYVPDPSHV 181
Query: 260 IPRDDVQVRDNLTVETMPLRIDNRKVKSLRGKEIPLVRVVWGGATGESLTWELESKMRES 319
I DDVQVRDNLTVET+P+RID+RKVK+LRGKEIPLVRVVW A GESLTWELESKM ES
Sbjct: 182 IQSDDVQVRDNLTVETLPVRIDDRKVKTLRGKEIPLVRVVWDRANGESLTWELESKMVES 361
Query: 320 YPELFA 325
YPELFA
Sbjct: 362 YPELFA 379
>TC77595 weakly similar to PIR|T18350|T18350 probable pol polyprotein - rice
blast fungus gypsy retroelement (fragment), partial
(14%)
Length = 1708
Score = 87.0 bits (214), Expect = 9e-18
Identities = 68/227 (29%), Positives = 106/227 (45%), Gaps = 6/227 (2%)
Frame = +2
Query: 36 PSSIVSDRDPRFTSRFWKSLQDALGSKLRLSSAYHPQTDGQSERTIQSLEDLLRVCVLEQ 95
P SIVSDR + RFW+ G LS++YHPQTDG +ER Q ++ +LR V
Sbjct: 569 PQSIVSDRGSNWVGRFWREFCRLTGVTQLLSTSYHPQTDGGTERWNQEIQAVLRAYVCWS 748
Query: 96 GGASDSHLPLIEFTYNNSYHSSIGMAPFEALYGWRCRTPLCWFESGESVVLGPD-----L 150
LP ++ N ++SSIG PF +G+ P+ E VV + L
Sbjct: 749 QDNWGDLLPTVQLALRNRHNSSIGATPFFVEHGYHV-DPIPTVEDTGGVVSEGEAAAQLL 925
Query: 151 VHETTEKVRMIREKMKASQSRQKSYHDKRRKDLE-FQEGDHVFLRVTPLTGVGRALKSRK 209
V + I+ ++ A+Q R ++ +KRR + +Q GD V+L V+ + K
Sbjct: 926 VKRMKDVTGFIQAEIVAAQQRSEASANKRRCPADRYQVGDKVWLNVSNYKSPRPSKKLDW 1105
Query: 210 LTPKFIGPYQISERVGTVAYRVGLPPHLSNLHDVFHVSQLRKYVADP 256
L K Y+++ V + +P ++ FHV LR+ +DP
Sbjct: 1106LHHK----YEVTRFVTPHVVELNVP---GTVYPKFHVDLLRRAASDP 1225
>BG454871 weakly similar to GP|10140673|g putative gag-pol polyprotein {Oryza
sativa (japonica cultivar-group)}, partial (7%)
Length = 674
Score = 64.3 bits (155), Expect(2) = 8e-15
Identities = 35/80 (43%), Positives = 43/80 (53%)
Frame = +2
Query: 49 SRFWKSLQDALGSKLRLSSAYHPQTDGQSERTIQSLEDLLRVCVLEQGGASDSHLPLIEF 108
S FWK L G+ L +SSAYHP +DGQSE + E LR + P E+
Sbjct: 32 SNFWKQLFKLHGTILTMSSAYHP*SDGQSEALNKGXEMYLRCLMFTDPLKWSKAFPWAEY 211
Query: 109 TYNNSYHSSIGMAPFEALYG 128
YN SY+ S M PF+ALYG
Sbjct: 212 WYNTSYNISAAMTPFKALYG 271
Score = 33.1 bits (74), Expect(2) = 8e-15
Identities = 21/73 (28%), Positives = 34/73 (45%), Gaps = 2/73 (2%)
Frame = +1
Query: 168 SQSRQKSYHDKRRKDLEFQEGDHVFLRVTPLTGVGRALKSRKL--TPKFIGPYQISERVG 225
+Q K DK+R+ EFQ G+HV +++ P AL+ + +P F +
Sbjct: 388 AQQTMKHQADKKRRHFEFQLGEHVLVKLQPYQQSSVALRKYQKFGSPNFGSLLTVCSL*V 567
Query: 226 TVAYRVGLPPHLS 238
A+ PP+LS
Sbjct: 568 ESAFHCKSPPYLS 606
>BG644699 similar to PIR|T07863|T078 probable polyprotein - pineapple
retrotransposon dea1 (fragment), partial (5%)
Length = 231
Score = 63.9 bits (154), Expect = 8e-11
Identities = 31/73 (42%), Positives = 48/73 (65%), Gaps = 1/73 (1%)
Frame = +2
Query: 189 DHVFLRVTPLT-GVGRALKSRKLTPKFIGPYQISERVGTVAYRVGLPPHLSNLHDVFHVS 247
+ V L+V P G R K KL+ ++IGP+++ +R+G VAY + LPP LS +H VFHVS
Sbjct: 2 EQVLLKVLPTERGDCRFGKRGKLSLRYIGPFEVIKRIGEVAYELALPPGLSGVHPVFHVS 181
Query: 248 QLRKYVADPSHVI 260
++Y D +++I
Sbjct: 182 MFKRYHGDGNYII 220
>BG587176 weakly similar to PIR|G84493|G84 probable retroelement pol
polyprotein [imported] - Arabidopsis thaliana, partial
(1%)
Length = 729
Score = 54.7 bits (130), Expect = 5e-08
Identities = 24/54 (44%), Positives = 37/54 (68%)
Frame = -1
Query: 271 LTVETMPLRIDNRKVKSLRGKEIPLVRVVWGGATGESLTWELESKMRESYPELF 324
L +ET P+RI +R K++R K I +V++VW + E +TWE E++M+ YPE F
Sbjct: 717 LDLETRPVRILDRMEKAMRKKPIQMVKIVWDCSGREEITWETEARMKADYPEWF 556
>BF636649 similar to GP|21628724|emb OSJNBa0033H08.7 {Oryza sativa}, partial
(4%)
Length = 653
Score = 34.7 bits (78), Expect = 0.051
Identities = 36/166 (21%), Positives = 66/166 (39%), Gaps = 4/166 (2%)
Frame = +1
Query: 73 TDGQSERTIQSLEDLLRVCVLEQGGASDSHLPLIEFTYNNSYHSSIGMAPFEALYGWRCR 132
+D Q+ +LE L EQ G + L E YN ++H + G PF+ +Y
Sbjct: 7 SDEQAGLLNHTLETHLLYFTSEQQGV*NFFLTWAECLYNTNFHRTAGCTPFKVVY---VV 177
Query: 133 TPLCWFESGESVVLGPDLVHETTEKVRMIREKMKASQSRQKSY----HDKRRKDLEFQEG 188
L F ++ + +H+++ R + +Y H +R + +
Sbjct: 178 AHLQKFVVARDLIYRNEGLHKSST*TSFGRGTRAYEALSRPAYETC*HPRRPLSIVYTRD 357
Query: 189 DHVFLRVTPLTGVGRALKSRKLTPKFIGPYQISERVGTVAYRVGLP 234
+V P K GPYQ+ +++G+VA+++ LP
Sbjct: 358 RTYEWQVLP-----------KYVA*CYGPYQVIKQIGSVAFKL*LP 462
>BF647164 weakly similar to GP|6691191|gb F7F22.15 {Arabidopsis thaliana},
partial (6%)
Length = 469
Score = 30.4 bits (67), Expect = 0.97
Identities = 22/69 (31%), Positives = 33/69 (46%)
Frame = +1
Query: 163 EKMKASQSRQKSYHDKRRKDLEFQEGDHVFLRVTPLTGVGRALKSRKLTPKFIGPYQISE 222
E K + R K +HD+R EF+EG+ V L + L L KL + GP+Q+
Sbjct: 142 ENAKIYKERTKKWHDRRIIRREFREGELVLLFNSRL-----KLFPGKLRSHWSGPFQVKN 306
Query: 223 RVGTVAYRV 231
+ + A V
Sbjct: 307 VMPSGAVEV 333
>TC87772 weakly similar to GP|8574455|gb|AAF77578.1| pepper esterase
{Capsicum annuum}, partial (34%)
Length = 1313
Score = 29.3 bits (64), Expect = 2.2
Identities = 18/52 (34%), Positives = 27/52 (51%), Gaps = 4/52 (7%)
Frame = -2
Query: 236 HLSNLHDVFHVSQLRKYVADPS----HVIPRDDVQVRDNLTVETMPLRIDNR 283
H+SN HDV + LR ++ S H +P D V V+D++ V+ D R
Sbjct: 628 HVSNQHDVLAMDLLRF*SSEESPRSLHRLPEDGVPVQDDIRVKQRRSEFD*R 473
>BE325109 similar to PIR|T04011|T040 hypothetical protein T5L19.200 -
Arabidopsis thaliana, partial (9%)
Length = 430
Score = 29.3 bits (64), Expect = 2.2
Identities = 13/43 (30%), Positives = 25/43 (57%)
Frame = +1
Query: 53 KSLQDALGSKLRLSSAYHPQTDGQSERTIQSLEDLLRVCVLEQ 95
KSLQ G++++L + P+ D ERT+Q D ++ + ++
Sbjct: 79 KSLQTKTGARIQLIPQHLPEGDDSKERTVQVTGDKRQIEIAQE 207
>BG587101 similar to GP|6691191|gb F7F22.15 {Arabidopsis thaliana}, partial
(10%)
Length = 624
Score = 28.5 bits (62), Expect = 3.7
Identities = 12/43 (27%), Positives = 25/43 (57%)
Frame = +2
Query: 36 PSSIVSDRDPRFTSRFWKSLQDALGSKLRLSSAYHPQTDGQSE 78
P ++SD F ++ ++ L G + ++++AYHPQ +S+
Sbjct: 488 PRVVISDGGSHFINKVFEKLLKKNGVRHKVATAYHPQKAERSK 616
>BE204912 similar to PIR|T02686|T026 probable Ser/Thr protein kinase
[imported] - Arabidopsis thaliana, partial (2%)
Length = 603
Score = 27.3 bits (59), Expect = 8.2
Identities = 18/47 (38%), Positives = 26/47 (55%), Gaps = 2/47 (4%)
Frame = +3
Query: 227 VAYRVGLPPHLS--NLHDVFHVSQLRKYVADPSHVIPRDDVQVRDNL 271
VAYR P LS N+ D+F++S LR +A S++ D + D L
Sbjct: 186 VAYRAQSPNFLSLSNISDIFNLSPLR--IAKASNIEAEDKKLIPDQL 320
>TC78537 similar to GP|10178176|dbj|BAB11650. trehalose-6-phosphate
phosphatase {Arabidopsis thaliana}, partial (75%)
Length = 1609
Score = 27.3 bits (59), Expect = 8.2
Identities = 14/43 (32%), Positives = 22/43 (50%)
Frame = +1
Query: 154 TTEKVRMIREKMKASQSRQKSYHDKRRKDLEFQEGDHVFLRVT 196
T K+R+ R +S +R+K D R + +G +F RVT
Sbjct: 889 TASKIRVKRVPEASSNTRKKGIGDSSRN*MGQGQGPRIFARVT 1017
>TC88607 similar to GP|8777331|dbj|BAA96921.1 receptor-like protein kinase
{Arabidopsis thaliana}, partial (35%)
Length = 899
Score = 27.3 bits (59), Expect = 8.2
Identities = 14/39 (35%), Positives = 19/39 (47%)
Frame = +3
Query: 148 PDLVHETTEKVRMIREKMKASQSRQKSYHDKRRKDLEFQ 186
PD+ E VRMI E ++ + S D + KDL Q
Sbjct: 633 PDMRPNMEEVVRMIEEIRQSDSDNRPSSDDNKSKDLNVQ 749
>CA858686 homologue to GP|8489871|gb| DEAD box protein P68 {Pisum sativum},
partial (31%)
Length = 598
Score = 27.3 bits (59), Expect = 8.2
Identities = 24/92 (26%), Positives = 36/92 (39%), Gaps = 2/92 (2%)
Frame = +2
Query: 166 KASQSRQKSYHDKRRKDLEFQEGDHVFLRVTPLTGVGRA--LKSRKLTPKFIGPYQISER 223
++ R+ + HD R V R +TGV L K T +I + R
Sbjct: 308 RSQSEREAALHDFRSSSTSILVATDVASRGLDVTGVSHVINLDLPKTTEDYIHRIGRTGR 487
Query: 224 VGTVAYRVGLPPHLSNLHDVFHVSQLRKYVAD 255
G+ G+ D+F VS +RK +AD
Sbjct: 488 AGST----GIATSFYTDRDMFLVSNIRKAIAD 571
>BQ124050 similar to PIR|T50014|T50 trehalose-6-phosphate phosphatase-like
protein - Arabidopsis thaliana, partial (32%)
Length = 758
Score = 27.3 bits (59), Expect = 8.2
Identities = 14/43 (32%), Positives = 22/43 (50%)
Frame = +2
Query: 154 TTEKVRMIREKMKASQSRQKSYHDKRRKDLEFQEGDHVFLRVT 196
T K+R+ R +S +R+K D R + +G +F RVT
Sbjct: 71 TASKIRVKRVPEASSNTRKKGIGDSSRN*MGQGQGPRIFARVT 199
>BE998730
Length = 820
Score = 27.3 bits (59), Expect = 8.2
Identities = 15/44 (34%), Positives = 20/44 (45%), Gaps = 6/44 (13%)
Frame = +1
Query: 286 KSLRGKEIPLVRVV------WGGATGESLTWELESKMRESYPEL 323
K RGK+ P+ + WG SL W + K+ SYP L
Sbjct: 592 KQTRGKQFPVRHTLFNMFKPWGVFVQNSLHWVMAKKLDGSYPWL 723
Database: MTGI
Posted date: Oct 22, 2004 3:39 PM
Number of letters in database: 27,044,181
Number of sequences in database: 36,976
Lambda K H
0.319 0.135 0.404
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 8,805,556
Number of Sequences: 36976
Number of extensions: 113695
Number of successful extensions: 567
Number of sequences better than 10.0: 32
Number of HSP's better than 10.0 without gapping: 564
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 566
length of query: 325
length of database: 9,014,727
effective HSP length: 96
effective length of query: 229
effective length of database: 5,465,031
effective search space: 1251492099
effective search space used: 1251492099
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.7 bits)
S2: 58 (26.9 bits)
Medicago: description of AC137081.4


