

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
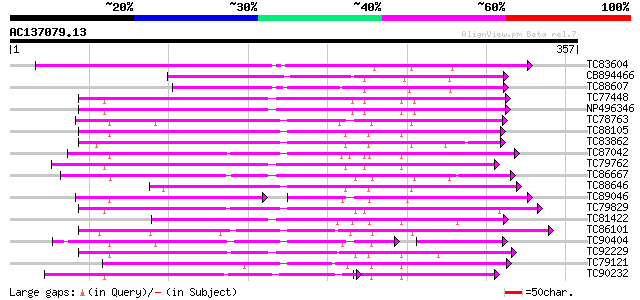
Query= AC137079.13 + phase: 0
(357 letters)
Database: MTGI
36,976 sequences; 27,044,181 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
TC83604 similar to PIR|T05606|T05606 protein kinase homolog F9D1... 189 2e-48
CB894466 similar to PIR|B84782|B84 probable receptor-like protei... 161 3e-40
TC88607 similar to GP|8777331|dbj|BAA96921.1 receptor-like prote... 142 3e-34
TC77448 SYMRK; MtSYMRK [Medicago truncatula]; nod region linked ... 134 7e-32
NP496346 NP496346|AJ418371.1|CAD10810.1 nod region linked recept... 134 7e-32
TC78763 somatic embryogenesis receptor kinase 1 [Medicago trunca... 132 2e-31
TC88105 similar to GP|21593085|gb|AAM65034.1 Putative protein ki... 131 3e-31
TC83862 similar to PIR|T05270|T05270 probable serine/threonine-s... 129 1e-30
TC87042 similar to GP|13605827|gb|AAK32899.1 AT5g48380/MJE7_1 {A... 129 2e-30
TC79762 similar to PIR|H96731|H96731 hypothetical protein F5A18.... 128 3e-30
TC86667 similar to GP|8778873|gb|AAF79872.1| T7N9.25 {Arabidopsi... 126 1e-29
TC88646 similar to GP|14194119|gb|AAK56254.1 At1g01540/F22L4_6 {... 125 3e-29
TC89046 weakly similar to GP|21740816|emb|CAD41006. OSJNBa0042L1... 76 1e-28
TC79829 similar to GP|20146232|dbj|BAB89014. similar to receptor... 121 4e-28
TC81422 similar to PIR|T02154|T02154 protein kinase homolog T1F1... 121 5e-28
TC86101 similar to GP|15217283|gb|AAK92627.1 Putative leucine-ri... 120 6e-28
TC90404 similar to GP|21593619|gb|AAM65586.1 receptor protein ki... 108 9e-28
TC92229 similar to PIR|B86369|B86369 hypothetical protein AAC980... 118 3e-27
TC79121 similar to GP|9651971|gb|AAF91337.1| Pti1 kinase-like pr... 118 4e-27
TC90232 weakly similar to PIR|T45697|T45697 hypothetical protein... 87 4e-27
>TC83604 similar to PIR|T05606|T05606 protein kinase homolog F9D16.210 -
Arabidopsis thaliana, partial (48%)
Length = 1142
Score = 189 bits (479), Expect = 2e-48
Identities = 122/325 (37%), Positives = 178/325 (54%), Gaps = 12/325 (3%)
Frame = +1
Query: 17 KATSEGRLKGGDNNNSELVFFVEDHERFKLEDLLRATADLRSENFWSSLFKVKFENNVEY 76
K + E + + N++L FF + F LEDLLRA+A++ + + + +K E+
Sbjct: 34 KMSPEKAVSRNMDANNKLTFFEGCNYAFDLEDLLRASAEVLGKGTFGTAYKAILEDATAV 213
Query: 77 AVKRLKNLQVSCDEFREILKQISKVKHQNILSLVGYRSTKEEKLIIYKYQSNGSVLNLLN 136
VKRLK + +F + ++ + +KH+N++ L Y +K+EKL++Y Y S GSV +LL+
Sbjct: 214 VVKRLKEVAFGKKDFEQYMEIVGSLKHENVVELKAYYYSKDEKLMVYDYYSRGSVSSLLH 393
Query: 137 DYIARRK-DFPWKLRLNIACGIARGLAFIYKKLEEGEVNSIPHGNLKLSNILLDDKNEAL 195
K W RL IA G ARG+A I+ +E G + HGN+K SNI L+ K
Sbjct: 394 GKRGEDKVPLDWDTRLRIALGAARGIAQIH--VENG--GKLVHGNIKSSNIFLNTKQYGC 561
Query: 196 ISEHGLSKFFEPDRGTFFSSHGYTAPEKSLTEK----GDVYSFGVILLELLTGQS-IEVS 250
+S+ GL+ + GY APE + T K DVYSFGV+LLELLTG+S I +
Sbjct: 562 VSDLGLATISTSLALPISRAAGYRAPEVTDTRKAAQPSDVYSFGVVLLELLTGKSPIHTT 741
Query: 251 R----IDLVRWVRSMVREEWTGEVFDKEVRE--NDHQGAFSLLNIALMCVSRSQENRPNF 304
I LVRWV S+VREEWT EVFD E+ N + +L IA+ CV R + RP
Sbjct: 742 GGDEIIHLVRWVHSVVREEWTAEVFDLELMRYPNIEEEMVEMLQIAMSCVVRMPDQRPKM 921
Query: 305 GEILETIEGVMNAHDQQQMELSASK 329
E+++ IE V + Q S ++
Sbjct: 922 SEVVKMIENVRQIDNTQTQPSSENQ 996
>CB894466 similar to PIR|B84782|B84 probable receptor-like protein kinase
[imported] - Arabidopsis thaliana, partial (36%)
Length = 871
Score = 161 bits (408), Expect = 3e-40
Identities = 99/238 (41%), Positives = 139/238 (57%), Gaps = 23/238 (9%)
Frame = +3
Query: 100 KVKHQNILSLVGYRSTKEEKLIIYKYQSNGSVLNLLNDYIAR-RKDFPWKLRLNIACGIA 158
K+KH NI+ L+ Y KEEKL++Y Y SNGS+ LL+ R W R+++ G A
Sbjct: 3 KLKHPNIVKLLAYYYAKEEKLLVYDYLSNGSLHALLHGNRGPGRIPLDWTTRISLVLGAA 182
Query: 159 RGLAFIYKKLEEGEVNSIPHGNLKLSNILLDDKNEALISEHGLSKFFEPDRGTFFSSHGY 218
RGLA I+ + +V PHGN+K SN+LLD A IS+ GLS P T GY
Sbjct: 183 RGLARIHTEYSAAKV---PHGNVKSSNVLLDKNGVACISDFGLSLLLNPVHATARLG-GY 350
Query: 219 TAPE----KSLTEKGDVYSFGVILLELLTGQSI----------------EVSRIDLVRWV 258
APE K L+++ DVYSFGV+LLE+LTG++ E + +DL +WV
Sbjct: 351 RAPEQTEQKRLSQQADVYSFGVLLLEVLTGKAPSLQYPSPANRPRKVEEEETVVDLPKWV 530
Query: 259 RSMVREEWTGEVFDKEV--RENDHQGAFSLLNIALMCVSRSQENRPNFGEILETIEGV 314
RS+VREEWTGEVFD+E+ +N + S+L++ L CV + E RP ++++ IE +
Sbjct: 531 RSVVREEWTGEVFDQELLRYKNIEEELVSMLHVGLACVVQQPEKRPTMVDVVKMIEDI 704
>TC88607 similar to GP|8777331|dbj|BAA96921.1 receptor-like protein kinase
{Arabidopsis thaliana}, partial (35%)
Length = 899
Score = 142 bits (357), Expect = 3e-34
Identities = 89/224 (39%), Positives = 129/224 (56%), Gaps = 12/224 (5%)
Frame = +3
Query: 103 HQNILSLVGYRSTKEEKLIIYKYQSNGSVLNLLNDY-IARRKDFPWKLRLNIACGIARGL 161
H N++ L Y +K+EKL++ Y NG++ LL+ R W R+ I+ GIARG+
Sbjct: 27 HPNVVPLRAYYYSKDEKLLVCDYFPNGNLSILLHGTRTGGRTTLDWNTRVKISLGIARGI 206
Query: 162 AFIYKKLEEGEVNSIPHGNLKLSNILLDDKNEALISEHGLSKFFEPDRGTFFSSHGYTAP 221
A ++ L G HGN+K SN+LL+ N+ IS+ GL+ T + GY AP
Sbjct: 207 AHLH--LVGGP--RFTHGNVKSSNVLLNQDNDGCISDFGLTPLMNIP-ATPSRTMGYRAP 371
Query: 222 E----KSLTEKGDVYSFGVILLELLTGQSIEVS-----RIDLVRWVRSMVREEWTGEVFD 272
E + T K DVYSFGV+LLE+LTG++ + S +DL RWVRS+VREEWT EVFD
Sbjct: 372 EVIETRKHTHKSDVYSFGVLLLEMLTGKAPQQSPVRDDMVDLPRWVRSVVREEWTAEVFD 551
Query: 273 KEVR--ENDHQGAFSLLNIALMCVSRSQENRPNFGEILETIEGV 314
E+ +N + +L I + CV++ + RPN E++ IE +
Sbjct: 552 VELMRYQNIEEEMVQMLQIGMTCVAKVPDMRPNMEEVVRMIEEI 683
>TC77448 SYMRK; MtSYMRK [Medicago truncatula]; nod region linked receptor
kinase [Medicago truncatula]
Length = 3568
Score = 134 bits (336), Expect = 7e-32
Identities = 90/291 (30%), Positives = 158/291 (53%), Gaps = 19/291 (6%)
Frame = +1
Query: 44 FKLEDLLRATADLRS---ENFWSSLFKVKFENNVEYAVK-RLKNLQVSCDEFREILKQIS 99
F LE + +AT ++ E + S+++ ++ E AVK R EF L +S
Sbjct: 2128 FTLEYIEQATEQYKTLIGEGGFGSVYRGTLDDGQEVAVKVRSSTSTQGTREFDNELNLLS 2307
Query: 100 KVKHQNILSLVGYRSTKEEKLIIYKYQSNGSVLNLLNDYIARRKDFPWKLRLNIACGIAR 159
++H+N++ L+GY + ++++++Y + SNGS+L+ L ++RK W RL+IA G AR
Sbjct: 2308 AIQHENLVPLLGYCNEYDQQILVYPFMSNGSLLDRLYGEASKRKILDWPTRLSIALGAAR 2487
Query: 160 GLAFIYKKLEEGEVNSIPHGNLKLSNILLDDKNEALISEHGLSKFFEPDRGTFFS----- 214
GLA+ L S+ H ++K SNILLD A +++ G SK+ + ++ S
Sbjct: 2488 GLAY----LHTFPGRSVIHRDVKSSNILLDQSMCAKVADFGFSKYAPQEGDSYVSLEVRG 2655
Query: 215 SHGYTAPE----KSLTEKGDVYSFGVILLELLTGQ---SIEVSRID--LVRWVRSMVREE 265
+ GY PE + L+EK DV+SFGV+LLE+++G+ +I+ RI+ LV W + +R
Sbjct: 2656 TAGYLDPEYYKTQQLSEKSDVFSFGVVLLEIVSGREPLNIKRPRIEWSLVEWAKPYIRAS 2835
Query: 266 WTGEVFDKEVRENDH-QGAFSLLNIALMCVSRSQENRPNFGEILETIEGVM 315
E+ D ++ H + + ++ +AL C+ RP +I+ +E +
Sbjct: 2836 KVDEIVDPGIKGGYHAEALWRVVEVALQCLEPYSTYRPCMVDIVRELEDAL 2988
>NP496346 NP496346|AJ418371.1|CAD10810.1 nod region linked receptor kinase
[Medicago truncatula]
Length = 2706
Score = 134 bits (336), Expect = 7e-32
Identities = 90/291 (30%), Positives = 158/291 (53%), Gaps = 19/291 (6%)
Frame = +1
Query: 44 FKLEDLLRATADLRS---ENFWSSLFKVKFENNVEYAVK-RLKNLQVSCDEFREILKQIS 99
F LE + +AT ++ E + S+++ ++ E AVK R EF L +S
Sbjct: 1684 FTLEYIEQATEQYKTLIGEGGFGSVYRGTLDDGQEVAVKVRSSTSTQGTREFDNELNLLS 1863
Query: 100 KVKHQNILSLVGYRSTKEEKLIIYKYQSNGSVLNLLNDYIARRKDFPWKLRLNIACGIAR 159
++H+N++ L+GY + ++++++Y + SNGS+L+ L ++RK W RL+IA G AR
Sbjct: 1864 AIQHENLVPLLGYCNEYDQQILVYPFMSNGSLLDRLYGEASKRKILDWPTRLSIALGAAR 2043
Query: 160 GLAFIYKKLEEGEVNSIPHGNLKLSNILLDDKNEALISEHGLSKFFEPDRGTFFS----- 214
GLA+ L S+ H ++K SNILLD A +++ G SK+ + ++ S
Sbjct: 2044 GLAY----LHTFPGRSVIHRDVKSSNILLDQSMCAKVADFGFSKYAPQEGDSYVSLEVRG 2211
Query: 215 SHGYTAPE----KSLTEKGDVYSFGVILLELLTGQ---SIEVSRID--LVRWVRSMVREE 265
+ GY PE + L+EK DV+SFGV+LLE+++G+ +I+ RI+ LV W + +R
Sbjct: 2212 TAGYLDPEYYKTQQLSEKSDVFSFGVVLLEIVSGREPLNIKRPRIEWSLVEWAKPYIRAS 2391
Query: 266 WTGEVFDKEVRENDH-QGAFSLLNIALMCVSRSQENRPNFGEILETIEGVM 315
E+ D ++ H + + ++ +AL C+ RP +I+ +E +
Sbjct: 2392 KVDEIVDPGIKGGYHAEALWRVVEVALQCLEPYSTYRPCMVDIVRELEDAL 2544
>TC78763 somatic embryogenesis receptor kinase 1 [Medicago truncatula]
Length = 2737
Score = 132 bits (333), Expect = 2e-31
Identities = 91/299 (30%), Positives = 160/299 (53%), Gaps = 27/299 (9%)
Frame = +3
Query: 42 ERFKLEDLLRATADLRSENF-----WSSLFKVKFENNVEYAVKRLKNLQVSCDE--FREI 94
+RF L +L AT ++N + ++K + + AVKRLK + E F+
Sbjct: 1290 KRFSLRELQVATDTFSNKNILGRGGFGKVYKGRLADGSLVAVKRLKEERTPGGELQFQTE 1469
Query: 95 LKQISKVKHQNILSLVGYRSTKEEKLIIYKYQSNGSVLNLLNDYIARRKDFPWKLRLNIA 154
++ IS H+N+L L G+ T E+L++Y Y +NGSV + L + ++ W R IA
Sbjct: 1470 VEMISMAVHRNLLRLRGFCMTPTERLLVYPYMANGSVASCLRERPPHQEPLDWPTRKRIA 1649
Query: 155 CGIARGLAFIYKKLEEGEVNSIPHGNLKLSNILLDDKNEALISEHGLSKFFE-------- 206
G ARGL++++ + I H ++K +NILLD++ EA++ + GL+K +
Sbjct: 1650 LGSARGLSYLHDHCDP----KIIHRDVKAANILLDEEFEAVVGDFGLAKLMDYKDTHVTT 1817
Query: 207 PDRGTFFSSHGYTAPEKSLT----EKGDVYSFGVILLELLTGQ-SIEVSR------IDLV 255
RGT G+ APE T EK DV+ +G++LLEL+TGQ + +++R + L+
Sbjct: 1818 AVRGTI----GHIAPEYLSTGKSSEKTDVFGYGIMLLELITGQRAFDLARLANDDDVMLL 1985
Query: 256 RWVRSMVREEWTGEVFDKEVRENDHQGAF-SLLNIALMCVSRSQENRPNFGEILETIEG 313
WV+ +++E+ + D +++ N + L+ +AL+C S +RP +++ +EG
Sbjct: 1986 DWVKGLLKEKKLEMLVDPDLKTNYIEAEVEQLIQVALLCTQGSPMDRPKMSDVVRMLEG 2162
>TC88105 similar to GP|21593085|gb|AAM65034.1 Putative protein kinase
{Arabidopsis thaliana}, partial (72%)
Length = 2379
Score = 131 bits (330), Expect = 3e-31
Identities = 89/289 (30%), Positives = 146/289 (49%), Gaps = 20/289 (6%)
Frame = +2
Query: 44 FKLEDLLRATADLRSENF-----WSSLFKVKFENNVEYAVKRL-KNLQVSCDEFREILKQ 97
F L DL AT +N + +++ + N A+K+L NL + EFR ++
Sbjct: 689 FTLRDLELATNKFSKDNIIGEGGYGVVYQGQLINGNPVAIKKLLNNLGQAEKEFRVEVEA 868
Query: 98 ISKVKHQNILSLVGYRSTKEEKLIIYKYQSNGSVLNLLNDYIARRKDFPWKLRLNIACGI 157
I V+H+N++ L+G+ +L+IY+Y +NG++ L+ + + W R+ I G
Sbjct: 869 IGHVRHKNLVRLLGFCIEGTHRLLIYEYVNNGNLEQWLHGAMRQYGYLTWDARIKILLGT 1048
Query: 158 ARGLAFIYKKLEEGEVNSIPHGNLKLSNILLDDKNEALISEHGLSKFFEPDRG----TFF 213
A+ LA++++ +E + H ++K SNIL+DD A IS+ GL+K +
Sbjct: 1049AKALAYLHEAIEP----KVVHRDIKSSNILIDDDFNAKISDFGLAKLLGAGKSHITTRVM 1216
Query: 214 SSHGYTAPEKS----LTEKGDVYSFGVILLELLTGQ-----SIEVSRIDLVRWVRSMVRE 264
+ GY APE + L EK DVYSFGV+LLE +TG+ + + ++LV W++ MV
Sbjct: 1217GTFGYVAPEYANSGLLNEKSDVYSFGVLLLEAITGRDPVDYNRSAAEVNLVDWLKMMVGN 1396
Query: 265 EWTGEVFDKEVRENDHQGAFS-LLNIALMCVSRSQENRPNFGEILETIE 312
EV D + A +L AL CV E RP +++ +E
Sbjct: 1397RHAEEVVDPNIETRPSTSALKRVLLTALRCVDPDSEKRPKMSQVVRMLE 1543
>TC83862 similar to PIR|T05270|T05270 probable serine/threonine-specific
protein kinase (EC 2.7.1.-) T4L20.80 - Arabidopsis
thaliana, partial (60%)
Length = 948
Score = 129 bits (325), Expect = 1e-30
Identities = 89/290 (30%), Positives = 155/290 (52%), Gaps = 21/290 (7%)
Frame = +1
Query: 44 FKLEDLLRAT-----ADLRSENFWSSLFKVKFENNVEYAVKRLKNLQVSCD-EFREILKQ 97
+ L++L AT + E + +++ ++ AVK L N + + EF+ ++
Sbjct: 13 YSLKELENATDGFAEGSVIGERGYGIVYRGILQDGSIVAVKNLLNNKGQAEKEFKVEVEA 192
Query: 98 ISKVKHQNILSLVGYRSTKEEKLIIYKYQSNGSVLNLLNDYIARRKDFPWKLRLNIACGI 157
I KV+H+N++ LVGY + +++++Y+Y NG++ L+ + W +R+ IA G
Sbjct: 193 IGKVRHKNLVGLVGYCAEGAKRMLVYEYVDNGNLEQWLHGDVGPVSPLTWDIRMKIAVGT 372
Query: 158 ARGLAFIYKKLEEGEVNSIPHGNLKLSNILLDDKNEALISEHGLSKFFEPDRG----TFF 213
A+GLA++++ LE + H ++K SNILLD K A +S+ GL+K +
Sbjct: 373 AKGLAYLHEGLEP----KVVHRDVKSSNILLDKKWHAKVSDFGLAKLLGSGKSYVTTRVM 540
Query: 214 SSHGYTAPEKS----LTEKGDVYSFGVILLELLTGQS-IEVSR----IDLVRWVRSMVRE 264
+ GY +PE + L E DVYSFG++L+EL+TG+S I+ SR ++LV W + MV
Sbjct: 541 GTFGYVSPEYASTGMLNEGSDVYSFGILLMELVTGRSPIDYSRAPAEMNLVDWFKGMVAS 720
Query: 265 EWTGEVFDK--EVRENDHQGAFSLLNIALMCVSRSQENRPNFGEILETIE 312
E+ D E++ + +LL + L C+ RP G+I+ +E
Sbjct: 721 RRGEELVDPLIEIQPSPRSLKRALL-VCLRCIDLDANKRPKMGQIVHMLE 867
>TC87042 similar to GP|13605827|gb|AAK32899.1 AT5g48380/MJE7_1 {Arabidopsis
thaliana}, partial (51%)
Length = 2206
Score = 129 bits (324), Expect = 2e-30
Identities = 93/309 (30%), Positives = 151/309 (48%), Gaps = 24/309 (7%)
Frame = +2
Query: 37 FVEDHERFKLEDLLRATADLRSENF-----WSSLFKVKFENNVEYAVKRLKNLQVSCDEF 91
F + + KL DL++AT + + N +++K E+ + VKRL+ Q S EF
Sbjct: 1007 FEKSISKMKLSDLMKATNNFSNINIIGTGRTGTVYKATLEDGTAFMVKRLQESQHSEKEF 1186
Query: 92 REILKQISKVKHQNILSLVGYRSTKEEKLIIYKYQSNGSVLNLLNDYIARRKDFPWKLRL 151
+ + VKH+N++ L+G+ K+E+L+++K NG + + L+ A W RL
Sbjct: 1187 MSEMATLGTVKHRNLVPLLGFCVAKKERLLVFKNMPNGMLHDQLHP-AAGECTLDWPSRL 1363
Query: 152 NIACGIARGLAFIYKKLEEGEVNSIPHGNLKLSNILLDDKNEALISEHGLSKFFEP---D 208
IA G A+G A+++ I H N+ ILLD E IS+ GL++ P
Sbjct: 1364 KIAIGAAKGFAWLHHSCNP----RIIHRNISSKCILLDADFEPKISDFGLARLMNPLDTH 1531
Query: 209 RGTF----FSSHGYTAPE--KSL--TEKGDVYSFGVILLELLTGQ-------SIEVSRID 253
TF F GY APE K+L T KGDV+SFG +LLEL+TG+ + E + +
Sbjct: 1532 LSTFVNGEFGDFGYVAPEYTKTLVATPKGDVFSFGTVLLELVTGERPANVAKAPETFKGN 1711
Query: 254 LVRWVRSMVREEWTGEVFDKE-VRENDHQGAFSLLNIALMCVSRSQENRPNFGEILETIE 312
LV W+ + + D+ + + D F L +A CV+ + RP E+ + +
Sbjct: 1712 LVEWITELSSNSKLHDAIDESLLNKGDDNELFQFLKVACNCVTEVPKERPTMFEVYQFLR 1891
Query: 313 GVMNAHDQQ 321
+ ++ Q
Sbjct: 1892 AIGGKYNFQ 1918
>TC79762 similar to PIR|H96731|H96731 hypothetical protein F5A18.8
[imported] - Arabidopsis thaliana, partial (55%)
Length = 1564
Score = 128 bits (322), Expect = 3e-30
Identities = 90/302 (29%), Positives = 151/302 (49%), Gaps = 20/302 (6%)
Frame = +3
Query: 27 GDNNNSELVFFVEDHERFKLEDLLRATADLRS-----ENFWSSLFKVKFENNVEYAVKRL 81
GDN + + F E LL AT + + E + ++K K + E AVK+L
Sbjct: 297 GDNEAELQKMASREQKIFSYETLLSATKNFNATHKLGEGGFGPVYKGKLSDGREVAVKKL 476
Query: 82 KNLQ-VSCDEFREILKQISKVKHQNILSLVGYRSTKEEKLIIYKYQSNGSVLNLLNDYIA 140
EF K +++V+H+N+++L+GY EK+++Y+Y + S+ L
Sbjct: 477 SQTSNQGKKEFMNEAKLLARVQHKNVVNLLGYCVHGTEKILVYEYVPHESLDKFLFKEAE 656
Query: 141 RRKDFPWKLRLNIACGIARGLAFIYKKLEEGEVNSIPHGNLKLSNILLDDKNEALISEHG 200
+R+ WK R I G+A+GL + L E N I H ++K SNILLDDK A I++ G
Sbjct: 657 KREQLDWKRRFGIITGVAKGLLY----LHEDSHNCIIHRDIKASNILLDDKWTAKIADFG 824
Query: 201 LSKFFEPDRG----TFFSSHGYTAPE----KSLTEKGDVYSFGVILLELLTGQ-----SI 247
+++ F D+ ++GY APE L+ K DV+S+GV++LEL+TGQ ++
Sbjct: 825 MARLFPEDQSQVKTRVAGTNGYMAPEYMMHGRLSVKADVFSYGVLVLELITGQRNSSFNL 1004
Query: 248 EVSRIDLVRWVRSMVREEWTGEVFDKEVRENDHQGAFSL-LNIALMCVSRSQENRPNFGE 306
V +L+ W M ++ + E+ D + + + +AL+C+ + RP
Sbjct: 1005XVEEHNLLDWAYKMYKKGRSLEIVDSALASTVLTEQVDMCIQLALLCIQGDPQLRPTMRR 1184
Query: 307 IL 308
I+
Sbjct: 1185IV 1190
>TC86667 similar to GP|8778873|gb|AAF79872.1| T7N9.25 {Arabidopsis
thaliana}, partial (52%)
Length = 1443
Score = 126 bits (317), Expect = 1e-29
Identities = 97/309 (31%), Positives = 157/309 (50%), Gaps = 23/309 (7%)
Frame = +3
Query: 33 ELVFFVEDHERFKLEDLLRATADLRSENFW-----SSLFKVKFENNVEYAVKRLKNLQVS 87
+L+FF R DL+ AT + +EN + ++ + AVKRL + ++
Sbjct: 90 KLIFFRNRLLR*SXGDLMAATNNFSNENVLITTRTGATYRADLPDGSTLAVKRLSSCKIG 269
Query: 88 CDEFREILKQISKVKHQNILSLVGYRSTKEEKLIIYKYQSNGSVLNLLNDYIARRKDFPW 147
+FR + ++ +V+H N+ L+GY +EEKL++YK+ SNG++ +LL+ W
Sbjct: 270 EKQFRMEMNRLGQVRHPNLAPLLGYCVVEEEKLLVYKHMSNGTLYSLLH---KNSGVLDW 440
Query: 148 KLRLNIACGIARGLAFIYKKLEEGEVNSIPHGNLKLSNILLDDKNEALISEHGLSKFFEP 207
+R I G ARGLA+ L G I N+ + IL+D++ +A I + GL++
Sbjct: 441 LMRFRIGLGAARGLAW----LHHGCHPPIIQQNICSNVILVDEEFDARIMDFGLARLMTS 608
Query: 208 D-RGTFFSSH----GYTAPEKSLTE----KGDVYSFGVILLELLTG-QSIEVSRID---- 253
D G+F + GY APE S T KGDVY FGV+LLEL+TG + +EV+ ID
Sbjct: 609 DANGSFVNGDLGELGYIAPEYSSTMVASLKGDVYGFGVLLLELVTGCKPLEVNNIDEEFK 788
Query: 254 --LVRWVRSMVREEWTGEVFDKEV--RENDHQGAFSLLNIALMCVSRSQENRPNFGEILE 309
LV WV + D+ + + ND + L IA CV ++R + ++
Sbjct: 789 GNLVDWVNMHSSSGRLKDCIDRSISGKGNDEE-ILQFLKIASNCVIARAKDRWSMYQVYN 965
Query: 310 TIEGVMNAH 318
+++G+ H
Sbjct: 966 SLKGISKDH 992
>TC88646 similar to GP|14194119|gb|AAK56254.1 At1g01540/F22L4_6 {Arabidopsis
thaliana}, partial (72%)
Length = 1825
Score = 125 bits (313), Expect = 3e-29
Identities = 73/252 (28%), Positives = 137/252 (53%), Gaps = 18/252 (7%)
Frame = +3
Query: 89 DEFREIL----KQISKVKHQNILSLVGYRSTKEEKLIIYKYQSNGSVLNLLNDYIARRKD 144
D+ +E L K I +V+H+N++ L+GY + ++++Y++ NG++ L+ +
Sbjct: 687 DKLKESLRLRWKPIGRVRHKNLVRLLGYCAEGAHRMLVYEFVDNGNLEQWLHGDVGPCSP 866
Query: 145 FPWKLRLNIACGIARGLAFIYKKLEEGEVNSIPHGNLKLSNILLDDKNEALISEHGLSKF 204
W++R+NI G A+GL ++++ LE + H ++K SNILL + + +S+ GL+K
Sbjct: 867 LTWEIRMNIILGTAKGLTYLHEGLEP----KVVHRDIKSSNILLSKQWNSKVSDFGLAKL 1034
Query: 205 FEPDRG----TFFSSHGYTAPEKS----LTEKGDVYSFGVILLELLTGQS-IEVSR---- 251
P+ + GY APE + L E+ DVYSFG++++E++TG + +E SR
Sbjct: 1035LSPESSYITTRVMGTFGYVAPEYASTGMLNERSDVYSFGILIMEVITGTNPVEYSRPAEE 1214
Query: 252 IDLVRWVRSMVREEWTGEVFDKEVRENDHQGAFS-LLNIALMCVSRSQENRPNFGEILET 310
++LV W++ MV V D ++ E A L +AL C + + RP G ++
Sbjct: 1215VNLVEWLKKMVSNRNPEGVLDPKLPEKPTSRALKRALLVALRCTDPNAQKRPKMGHVIHM 1394
Query: 311 IEGVMNAHDQQQ 322
+E + + +++
Sbjct: 1395LEAEDSPYKEER 1430
>TC89046 weakly similar to GP|21740816|emb|CAD41006. OSJNBa0042L16.4 {Oryza
sativa}, partial (35%)
Length = 1399
Score = 75.9 bits (185), Expect(2) = 1e-28
Identities = 45/127 (35%), Positives = 73/127 (57%), Gaps = 6/127 (4%)
Frame = +3
Query: 42 ERFKLEDLLRATADLRSENF-----WSSLFKVKFENNVEYAVKRLKNLQVSCD-EFREIL 95
ER+ L++LLRAT + +N + S++ + VE AVKRLK + + EF +
Sbjct: 189 ERYTLKELLRATNNFHQDNKIGEGGFGSVYWGQTSKGVEIAVKRLKTMTAKAEMEFAVEV 368
Query: 96 KQISKVKHQNILSLVGYRSTKEEKLIIYKYQSNGSVLNLLNDYIARRKDFPWKLRLNIAC 155
+ + +V+H+N+L L G+ + +E+LI+Y Y SN S+L L+ +A W R++I
Sbjct: 369 EVLGRVRHKNLLGLRGFYAGGDERLIVYDYMSNHSLLTHLHGQLASDCLLDWPRRMSITV 548
Query: 156 GIARGLA 162
G A GLA
Sbjct: 549 GAAEGLA 569
Score = 68.2 bits (165), Expect(2) = 1e-28
Identities = 50/172 (29%), Positives = 93/172 (54%), Gaps = 18/172 (10%)
Frame = +1
Query: 176 IPHGNLKLSNILLDDKNEALISEHGLSKFFEPD--------RGTFFSSHGYTAPEKSL-- 225
I H ++K SN+LLD + +A +++ G +K +GT GY APE ++
Sbjct: 601 IIHRDIKASNVLLDTEFQAKVADFGFAKLIPAGVSHLTTRVKGTL----GYLAPEYAMWG 768
Query: 226 --TEKGDVYSFGVILLELLTGQS-IEV----SRIDLVRWVRSMVREEWTGEVFDKEVREN 278
+E DVYSFG++LLE+++ + IE + D+V+WV V++ + D +++ N
Sbjct: 769 KVSESCDVYSFGILLLEIISAKKPIEKLPGGIKRDIVQWVTPYVQKGVFKHIADPKLKGN 948
Query: 279 -DHQGAFSLLNIALMCVSRSQENRPNFGEILETIEGVMNAHDQQQMELSASK 329
D + S++ IA+ C S + RP+ E++E ++ ++ ++ LS +K
Sbjct: 949 FDLEQLKSVIMIAVRCTDSSPDKRPSMIEVVEWLKDGVSKRKKEIPNLSNNK 1104
>TC79829 similar to GP|20146232|dbj|BAB89014. similar to receptor protein
kinase {Oryza sativa (japonica cultivar-group)}, partial
(40%)
Length = 1736
Score = 121 bits (304), Expect = 4e-28
Identities = 91/315 (28%), Positives = 154/315 (48%), Gaps = 23/315 (7%)
Frame = +2
Query: 44 FKLEDLLRATADLRS-----ENFWSSLFKVKFENNVEYAVKRLKNLQVSCD-EFREILKQ 97
F E+L AT + S + + ++K A+KR + + + EF +
Sbjct: 350 FTYEELSSATNNFSSSAQVGQGGYGKVYKGVISGGTAVAIKRAQEGSLQGEKEFLTEISL 529
Query: 98 ISKVKHQNILSLVGYRSTKEEKLIIYKYQSNGSVLNLLNDYIARRKDFPWKLRLNIACGI 157
+S++ H+N++SL+GY + E++++Y+Y NG++ + L+ ++ ++ + +RL IA G
Sbjct: 530 LSRLHHRNLVSLIGYCDEEGEQMLVYEYMPNGTLRDHLS--VSAKEPLTFIMRLKIALGS 703
Query: 158 ARGLAFIYKKLEEGEVNSIPHGNLKLSNILLDDKNEALISEHGLSKFFE-PDRGTFFSSH 216
A+GL +++ + + I H ++K SNILLD K A +++ GLS+ PD H
Sbjct: 704 AKGLMYLHNEADP----PIFHRDVKASNILLDSKLSAKVADFGLSRLAPVPDMEGIVPGH 871
Query: 217 ---------GYTAPE----KSLTEKGDVYSFGVILLELLTGQSIEVSRIDLVRWVRSMVR 263
GY PE LT+K DVYS GV+ LE+LTG ++VR V +
Sbjct: 872 VSTVVKGTPGYLDPEYFLTHKLTDKSDVYSLGVVFLEILTGMHPISHGKNIVREVNLSYQ 1051
Query: 264 EEWTGEVFDKEVRENDHQGAFSLLNIALMCVSRSQENRPNFGEI---LETIEGVMNAHDQ 320
+ D+ + + L +AL CV+ +NRP E+ LE I VM D
Sbjct: 1052SGVIFSIIDERMGSYPSEHVEKFLTLALKCVNDEPDNRPTMAEVVRELENIWNVMPESDT 1231
Query: 321 QQMELSASKCCSNGS 335
++ E S S+ S
Sbjct: 1232RRAESITSGSVSDSS 1276
>TC81422 similar to PIR|T02154|T02154 protein kinase homolog T1F15.2 -
Arabidopsis thaliana, partial (34%)
Length = 1130
Score = 121 bits (303), Expect = 5e-28
Identities = 90/260 (34%), Positives = 127/260 (48%), Gaps = 35/260 (13%)
Frame = +2
Query: 90 EFREILKQISKVKHQNILSLVGYRSTKEEKLIIYKYQSNGSVLNLLNDYIAR-RKDFPWK 148
EF ++ I KVKH NI+ L Y +EKL+I + SNG++ N L + + W
Sbjct: 20 EFATEVQAIGKVKHPNIVKLRAYYWAHDEKLLISDFVSNGNLANALRGRNGQPSPNLSWS 199
Query: 149 LRLNIACGIARGLAFIYKKLEEGEVNSIPHGNLKLSNILLDDKNEALISEHGLSKFF--- 205
+RL IA G ARGLA+ L E HG+LK SNILLD + LIS+ GL++
Sbjct: 200 IRLRIAKGTARGLAY----LHECSPRKFVHGDLKPSNILLDTDFQPLISDFGLNRLISIT 367
Query: 206 --EPDRGTFFS-------------SHGYTAPEKSL-----TEKGDVYSFGVILLELLTGQ 245
P G F ++ Y APE + T+K DVYSFGV+LLELLTG+
Sbjct: 368 GNNPSTGGFMGGALPYMKSSQTERTNNYKAPEAKVPGCRPTQKWDVYSFGVVLLELLTGK 547
Query: 246 --------SIEVSRIDLVRWV-RSMVREEWTGEVFDKEVRENDH--QGAFSLLNIALMCV 294
S V DLVRWV + +E E+ D + + H + ++ ++AL C
Sbjct: 548 SPDSSPGASTSVEVPDLVRWVKKGFEQESPLSEMVDPSLLQEIHAKKEVLAVFHVALSCT 727
Query: 295 SRSQENRPNFGEILETIEGV 314
E RP + + +E +
Sbjct: 728 EGDPEVRPRMKTVSDNLERI 787
>TC86101 similar to GP|15217283|gb|AAK92627.1 Putative leucine-rich repeat
transmembrane protein kinase 1 {Oryza sativa}, partial
(64%)
Length = 2629
Score = 120 bits (302), Expect = 6e-28
Identities = 94/324 (29%), Positives = 160/324 (49%), Gaps = 25/324 (7%)
Frame = +3
Query: 44 FKLEDLLRATAD-----LRSENFWSSLFKVKFENNVEYAVKRLKNLQVS---CDEFREIL 95
+ + DL AT L E + +++ +F++ AVK++ + + D+F EI+
Sbjct: 1476 YSIADLQIATGSFSVDHLLGEGSFGRVYRAQFDDGQVLAVKKIDSSVLPNNLSDDFMEIV 1655
Query: 96 KQISKVKHQNILSLVGYRSTKEEKLIIYKYQSNGSV---LNLLNDYIARRKDFPWKLRLN 152
+S++ H N+ L+GY S + L++Y+Y NGS+ L+L +DYI K W R+
Sbjct: 1656 SNLSRLHHPNVTELIGYCSEHGQHLLVYEYHKNGSLHDFLHLPDDYI---KPLIWNSRVK 1826
Query: 153 IACGIARGLAFIYKKLEEGEVNSIPHGNLKLSNILLDDKNEALISEHGLSKFFEPDRGTF 212
+A GIAR L ++++ S+ H N+K +NILLD +S+ GL+ + P+
Sbjct: 1827 VALGIARALEYLHEICSP----SVVHKNIKAANILLDADLNPHLSDSGLASYI-PNTNQV 1991
Query: 213 F---SSHGYTAPEKSLTE----KGDVYSFGVILLELLTGQS-IEVSRI----DLVRWVRS 260
S GY APE LT K DVYSFGV++LELL+G+ + SR LVRW
Sbjct: 1992 LNNNSGSGYDAPEVGLTGQYTLKSDVYSFGVVMLELLSGRKPFDSSRSRFEQSLVRWATP 2171
Query: 261 MVRE-EWTGEVFDKEVRENDHQGAFS-LLNIALMCVSRSQENRPNFGEILETIEGVMNAH 318
+ + + ++ D + + S ++ +CV E RP E+++ + ++
Sbjct: 2172 QLHDIDALAKMVDPALEGMYPVKSLSRFADVIALCVQSEPEFRPPMSEVVQALVRLVQRT 2351
Query: 319 DQQQMELSASKCCSNGSNQECCSL 342
+ + A + SN + SL
Sbjct: 2352 NMSKRTFGADQGGSNRGGDDQDSL 2423
>TC90404 similar to GP|21593619|gb|AAM65586.1 receptor protein kinase-like
protein {Arabidopsis thaliana}, partial (52%)
Length = 1252
Score = 108 bits (270), Expect(2) = 9e-28
Identities = 79/237 (33%), Positives = 126/237 (52%), Gaps = 19/237 (8%)
Frame = +3
Query: 28 DNNNSELVFFVEDHERFKLEDLLRATADLRSENF-----WSSLFKVKFENNVEYAVKRLK 82
D ++ E+ ++ + +RF +L AT + S+N + +++K + AVKRLK
Sbjct: 9 DRHHEEV--YLGNLKRFSFRELQVATNNFSSKNLVGKGGFGNVYKGVLSDGTVIAVKRLK 182
Query: 83 NLQVSCDE--FREILKQISKVKHQNILSLVGYRSTKEEKLIIYKYQSNGSVLNLLNDYIA 140
+ E F+ ++ IS H+N+L L G+ T E+L++Y Y NGSV + L
Sbjct: 183 DGNAIGGEIQFQTEVEMISLAVHRNLLRLYGFCMTSSERLLVYPYMCNGSVASRLKG--- 353
Query: 141 RRKDFPWKLRLNIACGIARGLAFIYKKLEEGEVNSIPHGNLKLSNILLDDKNEALISEHG 200
+ W R NIA G ARGL +++++ + I H ++K +NILLD+ EA++ + G
Sbjct: 354 -KPVLDWGTRKNIALGAARGLLYLHEQCDP----KIIHRDVKAANILLDNYYEAVVGDFG 518
Query: 201 LSKFFEPD--------RGTFFSSHGYTAPEKSLT----EKGDVYSFGVILLELLTGQ 245
L+K + RGT G+ APE T EK DV+ FG++LLEL+TGQ
Sbjct: 519 LAKLLDHQDSHVTTAVRGTV----GHIAPEYLSTGQSSEKTDVFGFGILLLELITGQ 677
Score = 32.7 bits (73), Expect(2) = 9e-28
Identities = 15/58 (25%), Positives = 31/58 (52%), Gaps = 1/58 (1%)
Frame = +1
Query: 257 WVRSMVREEWTGEVFDKEVREN-DHQGAFSLLNIALMCVSRSQENRPNFGEILETIEG 313
WV+ + +E+ + DK+++ N D ++ +AL+C +RP E++ +EG
Sbjct: 730 WVKKIHQEKKLELLVDKDLKSNYDKIELEEMVQVALLCTQYLPSHRPKMSEVVRMLEG 903
>TC92229 similar to PIR|B86369|B86369 hypothetical protein AAC98010.1
[imported] - Arabidopsis thaliana, partial (44%)
Length = 1347
Score = 118 bits (296), Expect = 3e-27
Identities = 95/301 (31%), Positives = 155/301 (50%), Gaps = 25/301 (8%)
Frame = +2
Query: 44 FKLEDLLRATADLRSENF-----WSSLFKVKFENNVEYAVKRLKNLQVSCD-EFREILKQ 97
F + +L T SEN + ++K + A+K LK + EFR +
Sbjct: 158 FSYDQILEITNGFSSENVIGEGGFGRVYKALMPDGRVGALKLLKAGSGQGEREFRAEVDT 337
Query: 98 ISKVKHQNILSLVGYRSTKEEKLIIYKYQSNGSVLNLLNDYIARRKDFPWKLRLNIACGI 157
IS+V H++++SL+GY ++++++IY++ NG++ L++ ++ W R+ IA G
Sbjct: 338 ISRVHHRHLVSLIGYCIAEQQRVLIYEFVPNGNLDQHLHE--SQWNVLDWPKRMKIAIGA 511
Query: 158 ARGLAFIYKKLEEGEVNSIPHGNLKLSNILLDDKNEALISEHGLSKFFEPDRGTFFSSH- 216
ARGLA+ L EG I H ++K SNILLDD EA +++ GL++ + D T S+
Sbjct: 512 ARGLAY----LHEGCNPKIIHRDIKSSNILLDDSYEAQVADFGLARLTD-DTNTHVSTRV 676
Query: 217 ----GYTAPEKS----LTEKGDVYSFGVILLELLTGQ-----SIEVSRIDLVRWVRS-MV 262
GY APE + LT++ DV+SFGV+LLEL+TG+ + V LV W R ++
Sbjct: 677 MGTFGYMAPEYATSGKLTDRSDVFSFGVVLLELVTGRKPVDPTQPVGDESLVEWARPILL 856
Query: 263 REEWTG---EVFDKEV-RENDHQGAFSLLNIALMCVSRSQENRPNFGEILETIEGVMNAH 318
R TG E+ D + R+ F ++ A C+ S RP +I ++ +
Sbjct: 857 RAIETGDFSELADPRLHRQYIDSEMFRMIEAAAACIRHSAPKRPRMVQIARALDSGDQLY 1036
Query: 319 D 319
D
Sbjct: 1037D 1039
>TC79121 similar to GP|9651971|gb|AAF91337.1| Pti1 kinase-like protein
{Glycine max}, partial (96%)
Length = 1283
Score = 118 bits (295), Expect = 4e-27
Identities = 86/279 (30%), Positives = 139/279 (48%), Gaps = 21/279 (7%)
Frame = +3
Query: 59 ENFWSSLFKVKFENNVEYAVKRLKNLQVSCDEFREILKQISKVKHQNILSLVGYRSTKEE 118
E + +++ +N E A+K+L + + EF + +S++KH+N++ LV Y
Sbjct: 369 EGAYGKVYRATLKNGREVAIKKLDSSKQPDQEFLSQVSIVSRLKHENVVELVNYCVDGPL 548
Query: 119 KLIIYKYQSNGSVLNLLNDYIARRKDFP-----WKLRLNIACGIARGLAFIYKKLEEGEV 173
+ + Y+Y NGS+ ++L+ + P W R+ IA G ARGL ++++K E
Sbjct: 549 RALAYEYAPNGSLHDILHGRKGVKGAEPGQVLSWAERVKIAVGAARGLEYLHEKAEV--- 719
Query: 174 NSIPHGNLKLSNILLDDKNEALISEHGLSKFFEPDRGT------FFSSHGYTAPEKSLT- 226
I H +K SNILL + A I++ LS PD + GY APE ++T
Sbjct: 720 -HIVHRYIKSSNILLFEDGVAKIADFDLSN-QAPDAAARLHSTRVLGTFGYHAPEYAMTG 893
Query: 227 ---EKGDVYSFGVILLELLTGQ-----SIEVSRIDLVRWVRSMVREEWTGEVFDKEVR-E 277
K DVYSFGVILLELLTG+ ++ + LV W + E+ + D ++ E
Sbjct: 894 NLSSKSDVYSFGVILLELLTGRKPVDHTLPRGQQSLVTWATPKLSEDKVKQCVDVRLKGE 1073
Query: 278 NDHQGAFSLLNIALMCVSRSQENRPNFGEILETIEGVMN 316
+ L +A +CV E RPN I++ ++ +MN
Sbjct: 1074YPSKSVAKLAAVAALCVQYEAEFRPNMSIIVKALQPLMN 1190
>TC90232 weakly similar to PIR|T45697|T45697 hypothetical protein F18L15.120
- Arabidopsis thaliana, partial (26%)
Length = 1272
Score = 87.4 bits (215), Expect(2) = 4e-27
Identities = 65/203 (32%), Positives = 101/203 (49%), Gaps = 4/203 (1%)
Frame = +3
Query: 23 RLKGGDNNNSELVFFVEDHERFKLEDLLRATADLRS---ENFWSSLFKVKFENNVEYAVK 79
R K G +N+ + HE F ++L T + ++ E + ++ +N + AVK
Sbjct: 240 RQKVGHSNSKKRGSLESTHEAFSYTEILNITNNFKTTIGEGGFGKVYLGILQNKTQVAVK 419
Query: 80 RLKNLQVS-CDEFREILKQISKVKHQNILSLVGYRSTKEEKLIIYKYQSNGSVLNLLNDY 138
L + EF+ + ++ V H+N++SL+GY E K +IY+Y +NG++ L +
Sbjct: 420 MLSPSSMQGYKEFQSEAQLLAIVHHRNLVSLIGYCDEGEIKALIYEYMANGNLQQHL--F 593
Query: 139 IARRKDFPWKLRLNIACGIARGLAFIYKKLEEGEVNSIPHGNLKLSNILLDDKNEALISE 198
+ W RLNIA A+GL +++ G I H +LK SNILLDD A IS+
Sbjct: 594 VENSNILNWNERLNIAVDAAQGLDYMH----NGCKPPILHRDLKPSNILLDDNMHAKISD 761
Query: 199 HGLSKFFEPDRGTFFSSHGYTAP 221
GLS+ F G SH T P
Sbjct: 762 FGLSRAF----GNDVDSHISTGP 818
Score = 51.6 bits (122), Expect(2) = 4e-27
Identities = 34/101 (33%), Positives = 49/101 (47%), Gaps = 9/101 (8%)
Frame = +1
Query: 217 GYTAPEKSLT----EKGDVYSFGVILLELLTGQ----SIEVSRIDLVRWVRSMVREEWTG 268
GY PE T +K D+YSFG+IL EL+TGQ + ++ WV +V
Sbjct: 832 GYADPEYQRTGNTNKKNDIYSFGIILFELITGQKALTKASGENLHILEWVIPIVEGGDIQ 1011
Query: 269 EVFDKEVR-ENDHQGAFSLLNIALMCVSRSQENRPNFGEIL 308
V D ++ E A+ ++ IA+ C S RP+ EIL
Sbjct: 1012NVVDSRLQGEFSINSAWKVVEIAMSCTSPDVVERPDMSEIL 1134
Database: MTGI
Posted date: Oct 22, 2004 3:39 PM
Number of letters in database: 27,044,181
Number of sequences in database: 36,976
Lambda K H
0.316 0.135 0.389
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 10,834,224
Number of Sequences: 36976
Number of extensions: 142129
Number of successful extensions: 1681
Number of sequences better than 10.0: 484
Number of HSP's better than 10.0 without gapping: 1253
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 1269
length of query: 357
length of database: 9,014,727
effective HSP length: 97
effective length of query: 260
effective length of database: 5,428,055
effective search space: 1411294300
effective search space used: 1411294300
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.3 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.6 bits)
S2: 59 (27.3 bits)
Medicago: description of AC137079.13


