

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC136973.11 - phase: 2 /pseudo
(192 letters)
Database: MTGI
36,976 sequences; 27,044,181 total letters
Searching..................................................done
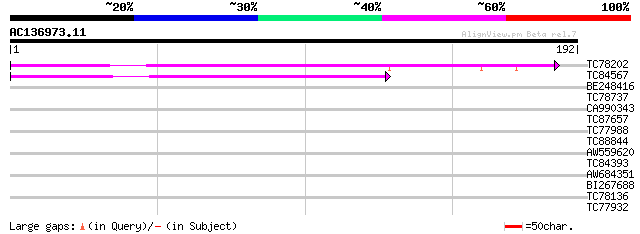
Score E
Sequences producing significant alignments: (bits) Value
TC78202 weakly similar to GP|16974602|gb|AAL31204.1 AT3g27760/MG... 84 4e-17
TC84567 similar to PIR|A86240|A86240 protein F20B24.10 [imported... 75 2e-14
BE248416 weakly similar to GP|15528715|dbj hypothetical protein~... 34 0.031
TC78737 similar to GP|1514443|emb|CAA67508.1 blue light receptor... 29 0.99
CA990343 similar to PIR|G96798|G967 hypothetical protein F22K20.... 28 1.7
TC87657 similar to GP|13430796|gb|AAK26020.1 unknown protein {Ar... 28 1.7
TC77988 homologue to GP|18375499|gb|AAK09427.2 sucrose-phosphate... 27 3.7
TC88844 similar to GP|20147219|gb|AAM10325.1 At1g59820/F23H11_14... 27 4.9
AW559620 weakly similar to GP|10727904|gb| CG5151 gene product {... 27 6.4
TC84393 homologue to GP|19352045|dbj|BAB85916. auxin response fa... 27 6.4
AW684351 weakly similar to GP|6016719|gb|A putative cell divisio... 27 6.4
BI267688 26 8.3
TC78136 similar to GP|22652297|gb|AAN03675.1 putative nucleopori... 26 8.3
TC77932 similar to GP|15289779|dbj|BAB63479. putative t-SNARE SE... 26 8.3
>TC78202 weakly similar to GP|16974602|gb|AAL31204.1 AT3g27760/MGF10_16
{Arabidopsis thaliana}, partial (33%)
Length = 1541
Score = 83.6 bits (205), Expect = 4e-17
Identities = 56/190 (29%), Positives = 90/190 (46%), Gaps = 4/190 (2%)
Frame = +1
Query: 1 WYDFVCFAIVGVSILGALWVLWTNEGSSTSQSDSNIFVESLLVANSPSSDNRVAIGHVST 60
W F+C + +++ +++W E + S+++ R +G +
Sbjct: 217 WRFFICAIFLLLTMGLGSFLIWKYEEFNKSRNER------------VEEGRRETVGLLYE 360
Query: 61 SQLWTSCWRGVHPLVLLTTRLFSFVSLAMLLYLDIHEYDASIFYYYTEWTFTLVMIYFAL 120
+ W +C +G+HP LL+ R+ SF L LL ++ IFY+YT+WTFTLV IYFAL
Sbjct: 361 DEAWNTCVKGIHPNWLLSYRIISFFVLLGLLTANVVVDGGGIFYFYTQWTFTLVTIYFAL 540
Query: 121 GTTVSAY-GCWKVLNKPPPLQNGEMTEFLRRDLETKGSI-FTFQSRYAEEEF--EQTAGF 176
+ S Y C+ + E ++ L+ I +S Y+ E TAG
Sbjct: 541 ASCFSFYRSCFNHNEFEGNTLDRERGTYVAPTLDGISDIPVLSKSSYSNRESLNRNTAGV 720
Query: 177 WGYLMQITFQ 186
WGY++QI FQ
Sbjct: 721 WGYIIQILFQ 750
>TC84567 similar to PIR|A86240|A86240 protein F20B24.10 [imported] -
Arabidopsis thaliana, partial (16%)
Length = 1305
Score = 74.7 bits (182), Expect = 2e-14
Identities = 41/129 (31%), Positives = 67/129 (51%)
Frame = +1
Query: 1 WYDFVCFAIVGVSILGALWVLWTNEGSSTSQSDSNIFVESLLVANSPSSDNRVAIGHVST 60
W F C + +S++ A ++++ EG + +S +++ G +
Sbjct: 208 WRFFFCALWIFISMILASYLIFKYEGFNKERSSER------------DENHQEEDGLLYE 351
Query: 61 SQLWTSCWRGVHPLVLLTTRLFSFVSLAMLLYLDIHEYDASIFYYYTEWTFTLVMIYFAL 120
+ W +C +G+ P+ LL R+ SFV L L+ ++ ASI YYT+ TFTLV IYF L
Sbjct: 352 DEAWNTCVKGIDPIWLLVYRIISFVILLALIIANVAVSGASILAYYTQLTFTLVTIYFGL 531
Query: 121 GTTVSAYGC 129
G++ S YGC
Sbjct: 532 GSSFSIYGC 558
>BE248416 weakly similar to GP|15528715|dbj hypothetical protein~similar to
Arabidopsis thaliana chromosome 5 MJH22.1, partial (5%)
Length = 335
Score = 34.3 bits (77), Expect = 0.031
Identities = 23/95 (24%), Positives = 37/95 (38%)
Frame = +3
Query: 1 WYDFVCFAIVGVSILGALWVLWTNEGSSTSQSDSNIFVESLLVANSPSSDNRVAIGHVST 60
W ++C V +S++ A +W E S + + DN +
Sbjct: 102 WRVYICVVSVLLSMILACLTIWKRESS-----------------RNLTFDNGENQSTLCG 230
Query: 61 SQLWTSCWRGVHPLVLLTTRLFSFVSLAMLLYLDI 95
+ W C + +HP+ LL R+ SF SL L I
Sbjct: 231 DEAWKPCLKDIHPVCLLVFRVISFSSLLASLIAKI 335
>TC78737 similar to GP|1514443|emb|CAA67508.1 blue light receptor {Arabidopsis
thaliana}, partial (62%)
Length = 1721
Score = 29.3 bits (64), Expect = 0.99
Identities = 14/48 (29%), Positives = 24/48 (49%), Gaps = 8/48 (16%)
Frame = +3
Query: 20 VLWTNEGSSTSQSDSNIFVESL--------LVANSPSSDNRVAIGHVS 59
+LW NEG+S + +F+ ++ L N P + R +GH+S
Sbjct: 1245 ILWGNEGNSVGEESVALFLRAIGLREYSRYLCFNFPFTHERALLGHLS 1388
>CA990343 similar to PIR|G96798|G967 hypothetical protein F22K20.8 [imported]
- Arabidopsis thaliana, partial (5%)
Length = 617
Score = 28.5 bits (62), Expect = 1.7
Identities = 16/53 (30%), Positives = 28/53 (52%)
Frame = -2
Query: 41 LLVANSPSSDNRVAIGHVSTSQLWTSCWRGVHPLVLLTTRLFSFVSLAMLLYL 93
LL NS SS+ + I H + ++W +G H LVL++ L + +++ L
Sbjct: 580 LLTINSRSSNV*LLIHHYFSDRIWDLLMKGQHLLVLISCLLLLLLLQLLVVLL 422
>TC87657 similar to GP|13430796|gb|AAK26020.1 unknown protein {Arabidopsis
thaliana}, partial (78%)
Length = 1738
Score = 28.5 bits (62), Expect = 1.7
Identities = 12/38 (31%), Positives = 19/38 (49%)
Frame = +2
Query: 81 LFSFVSLAMLLYLDIHEYDASIFYYYTEWTFTLVMIYF 118
LFSFV ++L+ + +F+Y W FT + F
Sbjct: 1547 LFSFVDFSLLVIITYRSIRYRVFFYCFHWFFTFSVFGF 1660
>TC77988 homologue to GP|18375499|gb|AAK09427.2 sucrose-phosphate synthase
{Medicago sativa}, partial (62%)
Length = 2248
Score = 27.3 bits (59), Expect = 3.7
Identities = 24/71 (33%), Positives = 33/71 (45%), Gaps = 3/71 (4%)
Frame = +3
Query: 29 TSQSDSNIFVESLLVANSPSSDNRVAIGHVSTSQLWTSCW-RGVHPL--VLLTTRLFSFV 85
TS+ +S + + A P+SDN I V S SC+ H L L +F V
Sbjct: 1998 TSRDESYLSLHVFCEALKPTSDNLTFIHIVHKSIFLPSCYFYNNHSLHHNLAP*SIFRLV 2177
Query: 86 SLAMLLYLDIH 96
SL LL+L +H
Sbjct: 2178 SLICLLFLYLH 2210
>TC88844 similar to GP|20147219|gb|AAM10325.1 At1g59820/F23H11_14
{Arabidopsis thaliana}, partial (44%)
Length = 1786
Score = 26.9 bits (58), Expect = 4.9
Identities = 17/68 (25%), Positives = 32/68 (47%)
Frame = +2
Query: 58 VSTSQLWTSCWRGVHPLVLLTTRLFSFVSLAMLLYLDIHEYDASIFYYYTEWTFTLVMIY 117
V S + WRG+ L+++ R +S++ + ++ I+++Y TFTL +
Sbjct: 656 VMASDFAIAQWRGLEDLLVVHGR-WSYLRICKVV----------IYFFYKNLTFTLTQFW 802
Query: 118 FALGTTVS 125
F T S
Sbjct: 803 FTFQTGFS 826
>AW559620 weakly similar to GP|10727904|gb| CG5151 gene product {Drosophila
melanogaster}, partial (4%)
Length = 352
Score = 26.6 bits (57), Expect = 6.4
Identities = 22/75 (29%), Positives = 37/75 (49%)
Frame = +2
Query: 35 NIFVESLLVANSPSSDNRVAIGHVSTSQLWTSCWRGVHPLVLLTTRLFSFVSLAMLLYLD 94
N+F +NS SS R++ + +T++L T C P LL LF ++ YL+
Sbjct: 59 NLFFCFWFPSNSSSSTTRLSTTNTTTTKLTTCCIGIC*PQSLLQLGLFLYL---CPFYLE 229
Query: 95 IHEYDASIFYYYTEW 109
+ Y I +Y +E+
Sbjct: 230 MDNY--IINFYISEF 268
>TC84393 homologue to GP|19352045|dbj|BAB85916. auxin response factor 7a
{Oryza sativa}, partial (1%)
Length = 659
Score = 26.6 bits (57), Expect = 6.4
Identities = 9/21 (42%), Positives = 15/21 (70%)
Frame = +1
Query: 14 ILGALWVLWTNEGSSTSQSDS 34
IL A W+LW N ++T+ +D+
Sbjct: 151 ILQATWILWANPSTNTTGTDT 213
>AW684351 weakly similar to GP|6016719|gb|A putative cell division control
protein {Arabidopsis thaliana}, partial (3%)
Length = 554
Score = 26.6 bits (57), Expect = 6.4
Identities = 10/28 (35%), Positives = 18/28 (63%)
Frame = +1
Query: 24 NEGSSTSQSDSNIFVESLLVANSPSSDN 51
N+ + + SNIFV+ L+ N P+++N
Sbjct: 49 NQSTLPPKKSSNIFVQITLIINEPNTNN 132
>BI267688
Length = 287
Score = 26.2 bits (56), Expect = 8.3
Identities = 12/34 (35%), Positives = 17/34 (49%)
Frame = +1
Query: 78 TTRLFSFVSLAMLLYLDIHEYDASIFYYYTEWTF 111
T L++F + LL H + S+F Y EW F
Sbjct: 22 TPPLYNFPICSSLLTFSSHIHAVSVFVYPQEWLF 123
>TC78136 similar to GP|22652297|gb|AAN03675.1 putative nucleoporin PRECOZ
{Arabidopsis thaliana}, partial (48%)
Length = 2532
Score = 26.2 bits (56), Expect = 8.3
Identities = 16/56 (28%), Positives = 27/56 (47%), Gaps = 14/56 (25%)
Frame = -3
Query: 63 LWTSCWRGVHPL-----VLLTTRLFSFVSLAMLLYLDI---------HEYDASIFY 104
LW SC+ G++ L +L+ TR+ F+S L +L EY ++F+
Sbjct: 2470 LWESCYLGIYHLLSDYTLLIFTRILQFLS*QELWHLQYKG*HATGFGKEYSTTVFF 2303
>TC77932 similar to GP|15289779|dbj|BAB63479. putative t-SNARE SED5 {Oryza
sativa (japonica cultivar-group)}, partial (71%)
Length = 1431
Score = 26.2 bits (56), Expect = 8.3
Identities = 11/21 (52%), Positives = 14/21 (66%)
Frame = -2
Query: 129 CWKVLNKPPPLQNGEMTEFLR 149
CW V N+ P Q GE+ +FLR
Sbjct: 500 CWIVKNRSPFRQFGELGKFLR 438
Database: MTGI
Posted date: Oct 22, 2004 3:39 PM
Number of letters in database: 27,044,181
Number of sequences in database: 36,976
Lambda K H
0.324 0.137 0.437
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 7,123,126
Number of Sequences: 36976
Number of extensions: 97115
Number of successful extensions: 860
Number of sequences better than 10.0: 28
Number of HSP's better than 10.0 without gapping: 853
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 859
length of query: 192
length of database: 9,014,727
effective HSP length: 91
effective length of query: 101
effective length of database: 5,649,911
effective search space: 570641011
effective search space used: 570641011
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 15 ( 7.0 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 40 (21.6 bits)
S2: 55 (25.8 bits)
Medicago: description of AC136973.11


