

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC136973.10 + phase: 0
(330 letters)
Database: MTGI
36,976 sequences; 27,044,181 total letters
Searching..................................................done
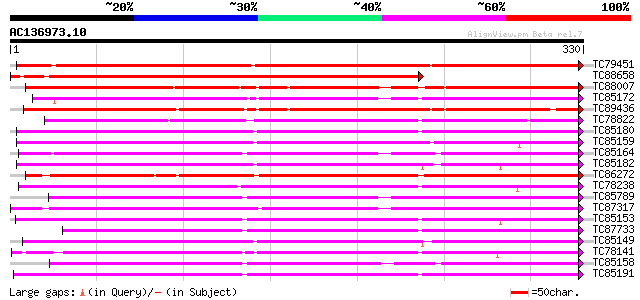
Score E
Sequences producing significant alignments: (bits) Value
TC79451 weakly similar to GP|4760700|dbj|BAA77387.1 peroxidase 1... 337 4e-93
TC88658 similar to GP|7453849|gb|AAF63024.1| peroxidase prx12 pr... 302 1e-82
TC88007 similar to PIR|T05993|T05993 probable peroxidase (EC 1.1... 253 7e-68
TC85172 weakly similar to GP|4760704|dbj|BAA77389.1 peroxidase 3... 246 1e-65
TC89436 similar to PIR|T09240|T09240 peroxidase (EC 1.11.1.7) pr... 240 6e-64
TC78822 similar to PIR|T02962|T02962 peroxidase (EC 1.11.1.7) is... 239 8e-64
TC85180 homologue to PIR|JC4780|JC4780 peroxidase (EC 1.11.1.7) ... 239 1e-63
TC85159 homologue to PIR|T09665|T09665 peroxidase (EC 1.11.1.7) ... 238 2e-63
TC85164 similar to SP|P22195|PER1_ARAHY Cationic peroxidase 1 pr... 236 7e-63
TC85182 homologue to PIR|JC4780|JC4780 peroxidase (EC 1.11.1.7) ... 235 2e-62
TC86272 similar to GP|4204761|gb|AAD11482.1| peroxidase precurso... 234 3e-62
TC78238 similar to PIR|T05723|T05723 peroxidase (EC 1.11.1.7) pr... 234 3e-62
TC85789 similar to GP|5002344|gb|AAD37428.1| peroxidase 3 precur... 234 5e-62
TC87317 similar to GP|5381253|dbj|BAA82306.1 peroxidase {Nicotia... 231 3e-61
TC85153 similar to GP|2245683|gb|AAC98519.1| peroxidase precurso... 229 1e-60
TC87733 similar to GP|10177520|dbj|BAB10915. peroxidase {Arabido... 227 4e-60
TC85149 similar to PIR|JC4780|JC4780 peroxidase (EC 1.11.1.7) 1B... 227 6e-60
TC78141 similar to GP|5002348|gb|AAD37430.1| peroxidase 5 precur... 226 9e-60
TC85158 similar to SP|P22195|PER1_ARAHY Cationic peroxidase 1 pr... 226 1e-59
TC85191 similar to PIR|JC4780|JC4780 peroxidase (EC 1.11.1.7) 1B... 225 2e-59
>TC79451 weakly similar to GP|4760700|dbj|BAA77387.1 peroxidase 1
{Scutellaria baicalensis}, partial (93%)
Length = 1229
Score = 337 bits (864), Expect = 4e-93
Identities = 178/326 (54%), Positives = 224/326 (68%)
Frame = +2
Query: 5 LSIATLVIVILSVSTTLASSTSLKYGFYKTTCSSVEAIVRRAVNKAVSLNPGIAAGLIRM 64
L+ A +V+VI ++ S L+ GFY +C E IV+ V K+ + NPGIAAGL+RM
Sbjct: 59 LNYAIIVLVIYFLNGNAHSQ--LEVGFYTYSCGMAEFIVKDEVRKSFNKNPGIAAGLVRM 232
Query: 65 HFHDCFVRGCDGSVLLDSIPGIQSERDHPANNPSLRGFEVINEAKAQIEAACPKTVSCAD 124
HFHDCF+RGCD SVLLDS +E+D PAN PSLRGFEVI+ AKA++E C VSCAD
Sbjct: 233 HFHDCFIRGCDASVLLDSTLSNIAEKDSPANKPSLRGFEVIDNAKAKLEEECKGIVSCAD 412
Query: 125 ILAFAARDSARKVSGGRIDYSVPSGRRDGRVSIFDEVTQNLPPPTFSAEQLIDNFDRKGL 184
I+AFAARDS G + Y VP+GRRDG++S+ + LPPPTF+ QL F +KGL
Sbjct: 413 IVAFAARDSVELAGG--LGYDVPAGRRDGKISLASDTRTELPPPTFNVNQLTQLFAKKGL 586
Query: 185 SVDEMVTLSGAHSIGVSHCSSFSKRLYSFNLTFPQDPSMDPNFARLLKSKCPPPQSQSIN 244
+ DEMVTLSGAH+IG SHCS+FSKRLY+F+ T QDPS+DP++A LLK +CP + N
Sbjct: 587 TQDEMVTLSGAHTIGRSHCSAFSKRLYNFSSTSIQDPSLDPSYAALLKRQCPQGNTNQ-N 763
Query: 245 PTVVLDGSTPNDLDNMYYKRLKNNRGLLTSDQTLLNSGLTRRMVLKNARHAAIWNVKFAK 304
V +D S+P D YY + NRGL TSDQTLL + T R V +NAR+ +W+ KFA
Sbjct: 764 LVVPMDPSSPGTADVGYYNDILANRGLFTSDQTLLTNTGTARKVHQNARNPYLWSNKFAD 943
Query: 305 AMVHMGSLDVLTGSEGEIRERCSVVN 330
AMV MG + VLTG+ GEIR C VVN
Sbjct: 944 AMVKMGQVGVLTGNAGEIRTNCRVVN 1021
>TC88658 similar to GP|7453849|gb|AAF63024.1| peroxidase prx12 precursor
{Spinacia oleracea}, partial (44%)
Length = 823
Score = 302 bits (773), Expect = 1e-82
Identities = 156/239 (65%), Positives = 187/239 (77%), Gaps = 1/239 (0%)
Frame = +1
Query: 1 MHSILSIATLVIVILSVSTTLASSTSLKYGFYKTTCSSVEAIVRRAVNKAVSLNPGIAAG 60
MH ++S +LV++ILS+S+ +S SL+ GFY +C S EAIVR A++KAVSLNPGI AG
Sbjct: 118 MHQMVS--SLVLIILSISSL--ASASLQVGFYSYSCPSAEAIVRSAIDKAVSLNPGIGAG 285
Query: 61 LIRMHFHDCFVRGCDGSVLLDSIPGIQ-SERDHPANNPSLRGFEVINEAKAQIEAACPKT 119
LIRMHFHDCFVRGCD SVLL S PG +E+D+ NNPSL GFEVI+EAKAQ+E CP+T
Sbjct: 286 LIRMHFHDCFVRGCDASVLLASTPGNPIAEKDNFINNPSLHGFEVIDEAKAQLEVVCPQT 465
Query: 120 VSCADILAFAARDSARKVSGGRIDYSVPSGRRDGRVSIFDEVTQNLPPPTFSAEQLIDNF 179
VSCADIL FA RDS K+SGG I+Y VPSGRRDGRVSI DEV +N+P P +A+QLI N
Sbjct: 466 VSCADILTFATRDSILKLSGGTINYDVPSGRRDGRVSISDEVPKNIPSPFLNADQLIANL 645
Query: 180 DRKGLSVDEMVTLSGAHSIGVSHCSSFSKRLYSFNLTFPQDPSMDPNFARLLKSKCPPP 238
+KG S+DEMVTLSGAHS+GVSH SFS R Y F+ T QDPSMDP+ A K++CPPP
Sbjct: 646 AQKGWSMDEMVTLSGAHSLGVSHGPSFSNR*YPFSDTISQDPSMDPSLAESWKTRCPPP 822
>TC88007 similar to PIR|T05993|T05993 probable peroxidase (EC 1.11.1.7)
F17M5.180 - Arabidopsis thaliana, partial (92%)
Length = 1254
Score = 253 bits (646), Expect = 7e-68
Identities = 142/321 (44%), Positives = 198/321 (61%)
Frame = +2
Query: 10 LVIVILSVSTTLASSTSLKYGFYKTTCSSVEAIVRRAVNKAVSLNPGIAAGLIRMHFHDC 69
++I +++ L +Y +C VE +V+ VN+A+ +P +AA LIRMHFHDC
Sbjct: 101 MLIEVITCQFGFGFGGGLNMNYYLMSCPFVEPVVKNIVNRALDNDPTLAAALIRMHFHDC 280
Query: 70 FVRGCDGSVLLDSIPGIQSERDHPANNPSLRGFEVINEAKAQIEAACPKTVSCADILAFA 129
F++GCDGS+LLDS +E+D PA N SLRG+EVI++ K ++E CP VSCADILA A
Sbjct: 281 FIQGCDGSILLDSTKDNTAEKDSPA-NLSLRGYEVIDDIKDELENRCPGVVSCADILAMA 457
Query: 130 ARDSARKVSGGRIDYSVPSGRRDGRVSIFDEVTQNLPPPTFSAEQLIDNFDRKGLSVDEM 189
A + A +GG + Y++P GR+DGR S ++ T+NLP P+F+A +LI F + G S EM
Sbjct: 458 ATE-AVFYAGGPV-YNIPKGRKDGRRSKIED-TRNLPSPSFNASELITQFGQHGFSAQEM 628
Query: 190 VTLSGAHSIGVSHCSSFSKRLYSFNLTFPQDPSMDPNFARLLKSKCPPPQSQSINPTVVL 249
V LSGAH++GV+ CSSF RL DP++D FAR L C + N
Sbjct: 629 VALSGAHTLGVARCSSFKNRLSQV------DPALDTEFARTLSRTC----TSGDNAEQPF 778
Query: 250 DGSTPNDLDNMYYKRLKNNRGLLTSDQTLLNSGLTRRMVLKNARHAAIWNVKFAKAMVHM 309
D +T ND DN+Y+ L G+L SDQTL +S TR +V A + A++ + F +AMV M
Sbjct: 779 D-ATRNDFDNVYFNALLRKNGVLFSDQTLYSSPRTRNIVNAYAMNQAMFFLDFQQAMVKM 955
Query: 310 GSLDVLTGSEGEIRERCSVVN 330
G LD+ GS GE+R C +N
Sbjct: 956 GLLDIKQGSNGEVRSNCRKIN 1018
>TC85172 weakly similar to GP|4760704|dbj|BAA77389.1 peroxidase 3
{Scutellaria baicalensis}, partial (88%)
Length = 1282
Score = 246 bits (627), Expect = 1e-65
Identities = 144/324 (44%), Positives = 192/324 (58%), Gaps = 7/324 (2%)
Frame = +1
Query: 14 ILSVSTTLASS-------TSLKYGFYKTTCSSVEAIVRRAVNKAVSLNPGIAAGLIRMHF 66
++ +TTL SS + L FY T C I+RR + AVS + + A L+R+HF
Sbjct: 106 VIPSATTLVSSLIPTPNYSELSSTFYGTRCPRALYIIRREIIAAVSRDRRLGASLLRLHF 285
Query: 67 HDCFVRGCDGSVLLDSIPGIQSERDHPANNPSLRGFEVINEAKAQIEAACPKTVSCADIL 126
HDCFV+GCD SVLL P Q E++ N SLRGFE I+ KA+IEA CP VSCADIL
Sbjct: 286 HDCFVQGCDASVLLKDTPTFQGEQNARPNANSLRGFEFIDSLKAKIEAVCPNVVSCADIL 465
Query: 127 AFAARDSARKVSGGRIDYSVPSGRRDGRVSIFDEVTQNLPPPTFSAEQLIDNFDRKGLSV 186
A AARDS + GG I + V GRRD + F+ +LP P + LI F +KG S
Sbjct: 466 AVAARDSVATL-GGPI-WGVRLGRRDSTTANFNAANSDLPSPFLNLSGLIAAFKKKGFSA 639
Query: 187 DEMVTLSGAHSIGVSHCSSFSKRLYSFNLTFPQDPSMDPNFARLLKSKCPPPQSQSINPT 246
DEMV LSGAH+IG + C+ F R+Y+ + +++P + R L++ C P++ N
Sbjct: 640 DEMVALSGAHTIGKAKCAVFKNRIYN-------ESNINPYYRRSLQNTC--PRNGGDNNL 792
Query: 247 VVLDGSTPNDLDNMYYKRLKNNRGLLTSDQTLLNSGLTRRMVLKNARHAAIWNVKFAKAM 306
LD +TP D+ YY+ L RGLL SDQ L N G T VL AR+ ++ FAKAM
Sbjct: 793 ANLDSTTPAFFDSAYYRNLLFKRGLLHSDQELYNGGSTDYKVLAYARNPYLFRFDFAKAM 972
Query: 307 VHMGSLDVLTGSEGEIRERCSVVN 330
+ MG+L LTG++G+IR+ CS VN
Sbjct: 973 IKMGNLSPLTGNQGQIRKYCSRVN 1044
>TC89436 similar to PIR|T09240|T09240 peroxidase (EC 1.11.1.7) prx11
precursor - spinach, partial (91%)
Length = 1127
Score = 240 bits (612), Expect = 6e-64
Identities = 145/323 (44%), Positives = 198/323 (60%), Gaps = 1/323 (0%)
Frame = +1
Query: 9 TLVIVILSVSTTLASSTSLKYGFYKTTCSSVEAIVRRAVNKAVSLNPGIAAGLIRMHFHD 68
T I+ L + S L +Y TC ++ I+ V A +P + A ++RM FHD
Sbjct: 52 TFPILFLLFTIFALSKAELHAHYYDQTCPQLDKIISETVLTASIHDPKVPARILRMFFHD 231
Query: 69 CFVRGCDGSVLLDSIPGIQSERDHPANNPSLRGFEVINEAKAQIEAACPKTVSCADILAF 128
CF+RGCD SVLLDS Q+E+D P N S+R F VI+EAKA++E ACP VSCADILA
Sbjct: 232 CFIRGCDASVLLDSTATNQAEKDGPPNI-SVRSFYVIDEAKAKLELACPGVVSCADILAL 408
Query: 129 AARDSARKVSGGRIDYSVPSGRRDGRVSIFDEVTQNLPPPTFSAEQLIDNFDRKGLSVDE 188
ARD +SGG + V GR+DGRVS + T NLP PT + QLI +F ++GL V +
Sbjct: 409 LARDVVA-MSGGPY-WKVLKGRKDGRVSKASD-TANLPAPTLNVGQLIQSFAKRGLGVKD 579
Query: 189 MVTLSGAHSIGVSHCSSFSKRLYSFNLTFPQDPSMDPNFARLLKSKCPPPQSQSINPTVV 248
MVTLSG H++G SHCSSF RL++F+ DP ++ FA LK+KCP P + N
Sbjct: 580 MVTLSGGHTLGFSHCSSFEARLHNFSSVHDTDPRLNTEFALDLKNKCPKPNNNQ-NAGQF 756
Query: 249 LDGSTPNDLDNMYYKRLKNNRGLLTSDQTLLNSGLTRRMVLKNARHAAIWNVKFAKAMVH 308
LD ST + DN YYK+L +G+ +SDQ+L+ TR +V AR +++ +FA +M+
Sbjct: 757 LD-STASVFDNDYYKQLLAGKGVFSSDQSLVGDYRTRWIVEAFARDQSLFFKEFAASMLK 933
Query: 309 MGSLDVLTGSE-GEIRERCSVVN 330
+G+ L GS+ GE+R C VVN
Sbjct: 934 LGN---LRGSDNGEVRLNCRVVN 993
>TC78822 similar to PIR|T02962|T02962 peroxidase (EC 1.11.1.7) isozyme 40K
precursor cationic - common tobacco, partial (70%)
Length = 1181
Score = 239 bits (611), Expect = 8e-64
Identities = 135/310 (43%), Positives = 186/310 (59%)
Frame = +2
Query: 21 LASSTSLKYGFYKTTCSSVEAIVRRAVNKAVSLNPGIAAGLIRMHFHDCFVRGCDGSVLL 80
++ SL+ FYK +C E IV+ + VS P + A LIR+HFHDCFVRGCD SVLL
Sbjct: 95 ISEGGSLRKNFYKKSCPQAEEIVKNITLQHVSSRPELPAKLIRLHFHDCFVRGCDASVLL 274
Query: 81 DSIPGIQSERDHPANNPSLRGFEVINEAKAQIEAACPKTVSCADILAFAARDSARKVSGG 140
+S G +E+D N SL GF+VI + K +E CP VSCADIL A RD+ +
Sbjct: 275 ESTAGNTAEKD-AIPNLSLAGFDVIEDIKEALEEKCPGIVSCADILTLATRDAFKN---- 439
Query: 141 RIDYSVPSGRRDGRVSIFDEVTQNLPPPTFSAEQLIDNFDRKGLSVDEMVTLSGAHSIGV 200
+ ++ V +GRRDG VS E N+P P + QL F K L++ ++V LSGAH+IGV
Sbjct: 440 KPNWEVLTGRRDGTVSRSIEALINIPAPFHNITQLRQIFANKKLTLHDLVVLSGAHTIGV 619
Query: 201 SHCSSFSKRLYSFNLTFPQDPSMDPNFARLLKSKCPPPQSQSINPTVVLDGSTPNDLDNM 260
HC+ FS RL++F QDPS++P +A LK+KC TV +D ++ DN
Sbjct: 620 GHCNLFSNRLFNFTGKGDQDPSLNPTYANFLKTKC--QGLSDTTTTVEMDPNSSTTFDND 793
Query: 261 YYKRLKNNRGLLTSDQTLLNSGLTRRMVLKNARHAAIWNVKFAKAMVHMGSLDVLTGSEG 320
YY L N+GL TSD LL + +R +V + + +F+++M MG+++VLTGS G
Sbjct: 794 YYPVLLQNKGLFTSDAALLTTKQSRNIVNELVSQNKFF-TEFSQSMKRMGAIEVLTGSNG 970
Query: 321 EIRERCSVVN 330
EIR +CSVVN
Sbjct: 971 EIRRKCSVVN 1000
>TC85180 homologue to PIR|JC4780|JC4780 peroxidase (EC 1.11.1.7) 1B
precursor - alfalfa, complete
Length = 1342
Score = 239 bits (609), Expect = 1e-63
Identities = 137/328 (41%), Positives = 189/328 (56%), Gaps = 2/328 (0%)
Frame = +1
Query: 5 LSIATLVIVILSVSTTLASSTSLKYGFYKTTCSSVEAIVRRAVNKAVSLNPGIAAGLIRM 64
++IA IV L S+ L FY TC +V +IVR + + + A L+R+
Sbjct: 31 VAIALCFIVALFGVLPFPSNAQLNPSFYSKTCPNVSSIVREVIRNVSKTDTRMLASLVRL 210
Query: 65 HFHDCFVRGCDGSVLLDSIPGIQSERDHPANNPSLRGFEVINEAKAQIEAACPKTVSCAD 124
HFHDCFV+GCD SVLL++ I SE+D N SLRG +V+N+ K +E ACP TVSCAD
Sbjct: 211 HFHDCFVQGCDASVLLNNTATIVSEQDAFPNRNSLRGLDVVNQIKTAVEKACPNTVSCAD 390
Query: 125 ILAFAARDSARKVSGGRIDYSVPSGRRDGRVSIFDEVTQNLPPPTFSAEQLIDNFDRKGL 184
ILA AA S+ G D+ VP GRRDG + QNLP P S +QL F +GL
Sbjct: 391 ILALAAELSSTLSQGP--DWKVPLGRRDGLTANQSLANQNLPAPFNSLDQLKAAFASQGL 564
Query: 185 SVDEMVTLSGAHSIGVSHCSSFSKRLYSFNLTFPQDPSMDPNFARLLKSKCPPPQSQSIN 244
S ++V LSGAH+ G +HCS F RLY+F+ T DP+++ + + L++ C P
Sbjct: 565 STTDLVALSGAHTFGRAHCSLFVSRLYNFSNTGSPDPTLNATYLQQLRNIC--PNGGPGT 738
Query: 245 PTVVLDGSTPNDLDNMYYKRLKNNRGLLTSDQTLLN-SGLTRRMVLKN-ARHAAIWNVKF 302
P D +TP+ D YY L+ +GLL SDQ L + SG ++ N A + F
Sbjct: 739 PLASFDPTTPDKFDKNYYSNLQVKKGLLQSDQELFSTSGADTISIVNNFATDQKAFFESF 918
Query: 303 AKAMVHMGSLDVLTGSEGEIRERCSVVN 330
AM+ MG++ VLTG++GEIR++C+ VN
Sbjct: 919 KAAMIKMGNIGVLTGNQGEIRKQCNFVN 1002
>TC85159 homologue to PIR|T09665|T09665 peroxidase (EC 1.11.1.7) pxdC
precursor - alfalfa, complete
Length = 1340
Score = 238 bits (608), Expect = 2e-63
Identities = 132/329 (40%), Positives = 192/329 (58%), Gaps = 3/329 (0%)
Frame = +2
Query: 5 LSIATLVIVILSVS-TTLASSTSLKYGFYKTTCSSVEAIVRRAVNKAVSLNPGIAAGLIR 63
LS+A L V++ + +S+ L FY+ TC +V +IVR + +P I A LIR
Sbjct: 77 LSLAALCCVVVVLGGLPFSSNAQLDNSFYRDTCPNVHSIVREVLRNVSKTDPRILASLIR 256
Query: 64 MHFHDCFVRGCDGSVLLDSIPGIQSERDHPANNPSLRGFEVINEAKAQIEAACPKTVSCA 123
+HFHDCFV+GCD S+LL++ I SE+ NN S+RG +V+N+ K +E ACP TVSCA
Sbjct: 257 LHFHDCFVQGCDASILLNTTSTITSEQTAFGNNNSIRGLDVVNQIKTAVENACPNTVSCA 436
Query: 124 DILAFAARDSARKVSGGRIDYSVPSGRRDGRVSIFDEVTQNLPPPTFSAEQLIDNFDRKG 183
DILA AA S+ +G D+ VP GRRD + NLP P F+ QL NFD +G
Sbjct: 437 DILALAAEISSVLANGP--DWKVPLGRRDSLTANLTLANINLPSPAFNLTQLKSNFDNQG 610
Query: 184 LSVDEMVTLSGAHSIGVSHCSSFSKRLYSFNLTFPQDPSMDPNFARLLKSKCPPPQSQSI 243
L ++V LSGAH+IG C F RLY+F+ T DP+++ + + L++ CP S
Sbjct: 611 LDATDLVALSGAHTIGRGQCRFFVDRLYNFSNTGNPDPTLNTTYLQTLRTICPNGGPGS- 787
Query: 244 NPTVVLDGSTPNDLDNMYYKRLKNNRGLLTSDQTLLNSGLTRRMVLKNA--RHAAIWNVK 301
LD +TP+ D+ YY L+ +GL SDQ L ++ + + N+ + ++
Sbjct: 788 -TLTDLDPATPDTFDSAYYSNLRIQKGLFQSDQVLSSTSGADTIAIVNSFNNNQTLFFEA 964
Query: 302 FAKAMVHMGSLDVLTGSEGEIRERCSVVN 330
F +M+ M + VLTGS+GEIR++C+ VN
Sbjct: 965 FKASMIKMSRIKVLTGSQGEIRKQCNFVN 1051
>TC85164 similar to SP|P22195|PER1_ARAHY Cationic peroxidase 1 precursor (EC
1.11.1.7). [Peanut] {Arachis hypogaea}, partial (92%)
Length = 1392
Score = 236 bits (603), Expect = 7e-63
Identities = 132/326 (40%), Positives = 193/326 (58%), Gaps = 1/326 (0%)
Frame = +3
Query: 6 SIATLVIVILSVSTTLASSTSLKYGFYKTTCSSVEAIVRRAVNKAVSLNPGIAAGLIRMH 65
++A ++I I+ + S+ L FY TTCS V + ++R ++ AV + A ++R+H
Sbjct: 42 TMAKIIIPIILCFVGIVSA-QLSTDFYSTTCSDVLSTIKREIDSAVGNEARMGASILRLH 218
Query: 66 FHDCFVRGCDGSVLLDSIPGIQSERDHPANNPSLRGFEVINEAKAQIEAACPKTVSCADI 125
FHDCFV+GCD SVLLD E+ AN SLRGF+VI+ K ++E+ CP TVSCADI
Sbjct: 219 FHDCFVQGCDASVLLDDTSSFTGEKTAGANANSLRGFDVIDTIKTELESLCPNTVSCADI 398
Query: 126 LAFAARDSARKVSGGRIDYSVPSGRRDGRVSIFDEVTQNLPPPTFSAEQLIDNFDRKGLS 185
L+ AARDS V+ G ++V GRRD + +LP P LI +FD KG +
Sbjct: 399 LSVAARDSV--VALGGPSWTVQLGRRDSITASLSLANSDLPGPGSDLSGLITSFDNKGFT 572
Query: 186 VDEMVTLSGAHSIGVSHCSSFSKRLYSFNLTFPQDPSMDPNFARLLKSKCPPP-QSQSIN 244
EMV LSG+H+IG + C F R+Y+ D ++D +FA L++ CP +++
Sbjct: 573 PKEMVALSGSHTIGQASCRFFRTRIYN-------DDNIDSSFATSLQANCPTTGGDDNLS 731
Query: 245 PTVVLDGSTPNDLDNMYYKRLKNNRGLLTSDQTLLNSGLTRRMVLKNARHAAIWNVKFAK 304
P LD +TPN DN Y++ L++ +GL +SDQ L N G T V + + ++ + FA
Sbjct: 732 P---LDTTTPNTFDNSYFQNLQSQKGLFSSDQALFNGGSTDSDVDEYSSDSSSFATDFAN 902
Query: 305 AMVHMGSLDVLTGSEGEIRERCSVVN 330
AMV MG+L+ +TGS G+IR C V+N
Sbjct: 903 AMVKMGNLNPITGSNGQIRTNCRVIN 980
>TC85182 homologue to PIR|JC4780|JC4780 peroxidase (EC 1.11.1.7) 1B
precursor - alfalfa, complete
Length = 1373
Score = 235 bits (600), Expect = 2e-62
Identities = 133/330 (40%), Positives = 191/330 (57%), Gaps = 4/330 (1%)
Frame = +3
Query: 5 LSIATLVIVILSVSTTLASSTSLKYGFYKTTCSSVEAIVRRAVNKAVSLNPGIAAGLIRM 64
++IA IV++ +S+ L FY+ TC +V +IVR + +P + L+R+
Sbjct: 162 VAIALCCIVVVLGGLPFSSNAQLDPSFYRNTCPNVSSIVREVIRSVSKKDPRMLGSLVRL 341
Query: 65 HFHDCFVRGCDGSVLLDSIPGIQSERDHPANNPSLRGFEVINEAKAQIEAACPKTVSCAD 124
HFHDCFV+GCD SVLL+ + SE+D N SLRG +V+N+ K +E ACP TVSCAD
Sbjct: 342 HFHDCFVQGCDASVLLNKTDTVVSEQDAFPNRNSLRGLDVVNQIKTAVEKACPNTVSCAD 521
Query: 125 ILAFAARDSARKVSGGRIDYSVPSGRRDGRVSIFDEVTQNLPPPTFSAEQLIDNFDRKGL 184
ILA +A S+ G D+ VP GRRDG + +NLP P + +QL F +GL
Sbjct: 522 ILALSAELSSTLADGP--DWKVPLGRRDGLTANQLLANKNLPAPFNTTDQLKAAFAAQGL 695
Query: 185 SVDEMVTLSGAHSIGVSHCSSFSKRLYSFNLTFPQDPSMDPNFARLLKSKCP--PPQSQS 242
++V LSGAH+ G +HCS F RLY+FN T DP+++ + + L++ CP P +
Sbjct: 696 DTTDLVALSGAHTFGRAHCSLFVSRLYNFNGTGSPDPTLNTTYLQQLRTICPNGGPGTNL 875
Query: 243 INPTVVLDGSTPNDLDNMYYKRLKNNRGLLTSDQTLLNS--GLTRRMVLKNARHAAIWNV 300
N D +TP+ D YY L+ +GLL SDQ L ++ T +V K A +
Sbjct: 876 TN----FDPTTPDKFDKNYYSNLQVKKGLLQSDQELFSTSGSDTISIVNKFATDQKAFFE 1043
Query: 301 KFAKAMVHMGSLDVLTGSEGEIRERCSVVN 330
F AM+ MG++ VLTG +GEIR++C+ VN
Sbjct: 1044SFKAAMIKMGNIGVLTGKQGEIRKQCNFVN 1133
>TC86272 similar to GP|4204761|gb|AAD11482.1| peroxidase precursor {Glycine
max}, partial (86%)
Length = 1350
Score = 234 bits (597), Expect = 3e-62
Identities = 140/324 (43%), Positives = 200/324 (61%), Gaps = 3/324 (0%)
Frame = +1
Query: 10 LVIVILSVSTTLASSTSLKYGFYKTTCSSVEAIVRRAVNKAVSLNPGIAAGLIRMHFHDC 69
L++ IL+ ST L+ GFY +C E IV V++ + P +AA LIRMHFHDC
Sbjct: 202 LILCILAAST----HAQLELGFYTKSCPKAEQIVANFVHEHIRNAPSLAAALIRMHFHDC 369
Query: 70 FVRGCDGSVLLDSIPGIQSERDHPANNPSLRGFEVINEAKAQIEAACPKTVSCADILAFA 129
FVRGCD SVLL+S Q+E++ P N ++RGF+ I+ K+ +EA CP VSCADI+A +
Sbjct: 370 FVRGCDASVLLNST-NQQAEKNAPPNL-TVRGFDFIDRIKSLVEAECPGVVSCADIIALS 543
Query: 130 ARDSARKVSGGRIDYSVPSGRRDGRVSIFDEVTQNLPPPTFSAEQLIDNFDRKGLSVDEM 189
ARDS G + VP+GRRDG VS E QN+P P + L F +GL + ++
Sbjct: 544 ARDSIAATGGPY--WKVPTGRRDGVVSNLLEANQNIPAPFSNFTTLQTLFANQGLDMKDL 717
Query: 190 VTLSGAHSIGVSHCSSFSKRLYSFNLTFPQDPSMDPNFARLLKS-KCPPPQSQSINPTVV 248
V LSGAH+IG+S C+SFS RLY+F QDPS+D +A+ LK+ KC ++ + N T+V
Sbjct: 718 VLLSGAHTIGISLCTSFSNRLYNFTGKGDQDPSLDSEYAKNLKTFKC---KNINDNTTIV 888
Query: 249 -LDGSTPNDLDNMYYKRLKNNRGLLTSDQTLLNSGLTRRMVLKNARHA-AIWNVKFAKAM 306
LD + N D YY ++ RGL SD LL + +T+ +V + + + + +FAK++
Sbjct: 889 ELDPGSRNTFDLGYYSQVVKRRGLFESDSALLTNSVTKALVTQFLQGSLENFYAEFAKSI 1068
Query: 307 VHMGSLDVLTGSEGEIRERCSVVN 330
MG + V TGS+G IR+ C++VN
Sbjct: 1069EKMGQIKVKTGSQGVIRKHCALVN 1140
>TC78238 similar to PIR|T05723|T05723 peroxidase (EC 1.11.1.7) precursor
seed coat - soybean, partial (90%)
Length = 1243
Score = 234 bits (597), Expect = 3e-62
Identities = 134/327 (40%), Positives = 192/327 (57%), Gaps = 2/327 (0%)
Frame = +2
Query: 6 SIATLVIVILSVSTTLASSTSLKYGFYKTTCSSVEAIVRRAVNKAVSLNPGIAAGLIRMH 65
SI L +V A L FY TC + IV R + +A +P I A LIR+H
Sbjct: 32 SINVLGVVFWCAVLMHAGYAQLSPSFYSQTCPFLYPIVFRVIFEASLTDPRIGASLIRLH 211
Query: 66 FHDCFVRGCDGSVLLDSIPGIQSERDHPANNPSLRGFEVINEAKAQIEAACPKTVSCADI 125
FHDCFV+GCDGSVLL++ I SE+D N SLRG +V+N+ K +E CP TVSCADI
Sbjct: 212 FHDCFVQGCDGSVLLNNTNTIVSEQDALPNINSLRGLDVVNQIKTAVENECPATVSCADI 391
Query: 126 LAFAARDSARKVSGGRIDYSVPSGRRDGRVSIFDEVTQNLPPPTFSAEQLIDNFDRKGLS 185
L AA+ V GG + +P GRRD + QNLP P F+ +QL F +GL+
Sbjct: 392 LTIAAQ--VASVLGGGPSWQIPLGRRDSLTANQALANQNLPAPFFTLDQLKAAFLVQGLN 565
Query: 186 VDEMVTLSGAHSIGVSHCSSFSKRLYSFNLTFPQDPSMDPNFARLLKSKCPPPQSQSINP 245
++VTLSGAH+ G + CS+F RLY+FN T D +++ + + L+ C PQ+ + N
Sbjct: 566 TTDLVTLSGAHTFGRAKCSTFINRLYNFNSTGNPDQTLNTTYLQTLREIC--PQNGTGNN 739
Query: 246 TVVLDGSTPNDLDNMYYKRLKNNRGLLTSDQTLLNSGLTRRMVLKN--ARHAAIWNVKFA 303
LD +TPN DN +Y L++++GLL SDQ L ++ + + N + + A++ F
Sbjct: 740 LTNLDLTTPNQFDNKFYSNLQSHKGLLQSDQELFSTPNADTIAIVNSFSSNQALFFENFR 919
Query: 304 KAMVHMGSLDVLTGSEGEIRERCSVVN 330
+M+ M ++ VLTG+EGEIR +C+ +N
Sbjct: 920 VSMIKMANISVLTGNEGEIRLQCNFIN 1000
>TC85789 similar to GP|5002344|gb|AAD37428.1| peroxidase 3 precursor
{Phaseolus vulgaris}, partial (92%)
Length = 1467
Score = 234 bits (596), Expect = 5e-62
Identities = 132/310 (42%), Positives = 177/310 (56%), Gaps = 2/310 (0%)
Frame = +2
Query: 23 SSTSLKYGFYKTTCSSVEAIVRRAVNKAVSLNPGIAAGLIRMHFHDCFVRGCDGSVLLDS 82
S + L FY C +V V V+ AV+ P + L+R+HFHDCFV GCDGSVLLD
Sbjct: 212 SFSQLSENFYAKKCPNVFKAVNSVVHSAVAREPRMGGSLLRLHFHDCFVNGCDGSVLLDD 391
Query: 83 IPGIQSERDHPANNPSLRGFEVINEAKAQIEAACPKTVSCADILAFAARDSARKVSGGRI 142
P + E+ N SLRGFEVI+ K+++E+ CP VSCADI+A AARDS V+ G
Sbjct: 392 TPSNKGEKTALPNKDSLRGFEVIDAIKSKVESVCPGVVSCADIVAIAARDSV--VNLGGP 565
Query: 143 DYSVPSGRRDGRVSIFDEVTQNLPPPTFSA-EQLIDNFDRKGLSVDEMVTLSGAHSIGVS 201
+ V GRRD + + ++ + PP FS LI+ F +GLS +MV LSGAH+IG +
Sbjct: 566 FWKVKLGRRDSKTASLNDANSGVIPPPFSTLNNLINRFKAQGLSTKDMVALSGAHTIGKA 745
Query: 202 HCSSFSKRLYSFNLTFPQDPSMDPNFARLLKSKCPPPQSQ-SINPTVVLDGSTPNDLDNM 260
C+ + R+Y+ D ++D FA+ + CP N VLD TPN DN+
Sbjct: 746 RCTVYRDRIYN-------DTNIDSLFAKSRQRNCPRKSGTIKDNNVAVLDFKTPNHFDNL 904
Query: 261 YYKRLKNNRGLLTSDQTLLNSGLTRRMVLKNARHAAIWNVKFAKAMVHMGSLDVLTGSEG 320
YYK L N +GLL SDQ L N G T +V + + + FA AM+ MG+ LTGS G
Sbjct: 905 YYKNLINKKGLLHSDQELFNGGSTDSLVKSYSNNQNAFESDFAIAMIKMGNNKPLTGSNG 1084
Query: 321 EIRERCSVVN 330
EIR++C N
Sbjct: 1085EIRKQCRRAN 1114
>TC87317 similar to GP|5381253|dbj|BAA82306.1 peroxidase {Nicotiana
tabacum}, partial (94%)
Length = 1237
Score = 231 bits (589), Expect = 3e-61
Identities = 128/330 (38%), Positives = 185/330 (55%)
Frame = +3
Query: 1 MHSILSIATLVIVILSVSTTLASSTSLKYGFYKTTCSSVEAIVRRAVNKAVSLNPGIAAG 60
M S L I ++V++S+ + ++ +L +Y ++C + V+ V A+S + A
Sbjct: 84 MTSNLMICFSLLVLVSIGS---ANANLSKDYYYSSCPKLFETVKCEVQSAISKETRMGAS 254
Query: 61 LIRMHFHDCFVRGCDGSVLLDSIPGIQSERDHPANNPSLRGFEVINEAKAQIEAACPKTV 120
L+R+ FHDCFV GCDGS+LLD E+ N S RGFEVI++ K+ +E CP V
Sbjct: 255 LLRLFFHDCFVNGCDGSILLDDTSSFTGEKTANPNKNSARGFEVIDKIKSAVEKVCPGAV 434
Query: 121 SCADILAFAARDSARKVSGGRIDYSVPSGRRDGRVSIFDEVTQNLPPPTFSAEQLIDNFD 180
SCADIL ARDS + G D V GRRD R + ++P PT S QLI F+
Sbjct: 435 SCADILTITARDSVEILGGPTWD--VKLGRRDARTASKSAANNDIPAPTSSLNQLISRFN 608
Query: 181 RKGLSVDEMVTLSGAHSIGVSHCSSFSKRLYSFNLTFPQDPSMDPNFARLLKSKCPPPQS 240
GLS ++V LSG H+IG + C++F +Y+ D ++D +FAR +S CP
Sbjct: 609 ALGLSTKDLVALSGGHTIGQARCTTFRAHIYN-------DSNIDTSFARTRQSGCPKTSG 767
Query: 241 QSINPTVVLDGSTPNDLDNMYYKRLKNNRGLLTSDQTLLNSGLTRRMVLKNARHAAIWNV 300
N LD +TP DN Y+K L +++GLL SDQ L N G T +V + + + + ++
Sbjct: 768 SGDNNLAPLDLATPTSFDNHYFKNLVDSKGLLHSDQQLFNGGSTDSIVHEYSLYPSSFSS 947
Query: 301 KFAKAMVHMGSLDVLTGSEGEIRERCSVVN 330
F AM+ MG + LTGS GEIR++C VN
Sbjct: 948 DFVTAMIKMGDISPLTGSNGEIRKQCRSVN 1037
>TC85153 similar to GP|2245683|gb|AAC98519.1| peroxidase precursor {Glycine
max}, complete
Length = 1413
Score = 229 bits (584), Expect = 1e-60
Identities = 128/329 (38%), Positives = 192/329 (57%), Gaps = 2/329 (0%)
Frame = +3
Query: 4 ILSIATLVIVILSVSTTLASSTSLKYGFYKTTCSSVEAIVRRAVNKAVSLNPGIAAGLIR 63
++ A +V++ +S+ L FY TC + +I + + K +P + A +IR
Sbjct: 60 VIVTALCCVVVVFGGLPFSSNAQLDPYFYGKTCPKLHSIAFKVLRKVAKTDPRMPASIIR 239
Query: 64 MHFHDCFVRGCDGSVLLDSIPGIQSERDHPANNPSLRGFEVINEAKAQIEAACPKTVSCA 123
+HFHDCFV+GCD SVLL++ I SE+D N SLRG +VIN+ K ++E ACP VSCA
Sbjct: 240 LHFHDCFVQGCDASVLLNNTATIVSEQDAFPNINSLRGLDVINQIKTKVEKACPNRVSCA 419
Query: 124 DILAFAARDSARKVSGGRIDYSVPSGRRDGRVSIFDEVTQNLPPPTFSAEQLIDNFDRKG 183
DIL A+ S+ V G + VP GRRD + QNLP P FS ++L F +G
Sbjct: 420 DILTLASGISS--VLTGGPGWEVPLGRRDSLTANQSLANQNLPGPNFSLDRLKSAFAAQG 593
Query: 184 LSVDEMVTLSGAHSIGVSHCSSFSKRLYSFNLTFPQDPSMDPNFARLLKSKCPPPQSQSI 243
L+ ++V LSGAH+ G + C RLY+FN T DP++D + + L+++C PQ+ +
Sbjct: 594 LNTVDLVALSGAHTFGRARCLFILDRLYNFNNTGKPDPTLDTTYLQQLRNQC--PQNGTG 767
Query: 244 NPTVVLDGSTPNDLDNMYYKRLKNNRGLLTSDQTLLNS--GLTRRMVLKNARHAAIWNVK 301
N V D +TP+ LD +Y L+ +GLL SDQ L ++ T +V A ++
Sbjct: 768 NNRVNFDPTTPDTLDKNFYNNLQGKKGLLQSDQELFSTPGADTISIVNSFANSQNVFFQN 947
Query: 302 FAKAMVHMGSLDVLTGSEGEIRERCSVVN 330
F +M+ MG++DVLTG +GEIR++C+ +N
Sbjct: 948 FINSMIKMGNIDVLTGKKGEIRKQCNFIN 1034
>TC87733 similar to GP|10177520|dbj|BAB10915. peroxidase {Arabidopsis
thaliana}, partial (89%)
Length = 1270
Score = 227 bits (579), Expect = 4e-60
Identities = 127/301 (42%), Positives = 178/301 (58%), Gaps = 1/301 (0%)
Frame = +1
Query: 31 FYKTTCSSVEAIVRRAVNKAVSLNPGIAAGLIRMHFHDCFVRGCDGSVLLDSIPGIQSER 90
FY +C + IV+ + AV+ P IAA L+R+HFHDCFV+GCD S+LLD+ I SE+
Sbjct: 148 FYDYSCPQAQNIVKSILANAVAKEPRIAASLLRLHFHDCFVKGCDASILLDNSGSIISEK 327
Query: 91 DHPANNPSLRGFEVINEAKAQIEAACPKTVSCADILAFAARDSARKVSGGRIDYSVPSGR 150
N S RGFEVI+E K +E CP TVSCADILA AARDS V G ++ VP GR
Sbjct: 328 GSNPNRNSARGFEVIDEIKYALEKECPHTVSCADILAIAARDST--VLAGGPNWEVPLGR 501
Query: 151 RDGRVSIFDEVTQNLPPPTFSAEQLIDNFDRKGLSVDEMVTLSGAHSIGVSHCSSFSKRL 210
RD + N+P P + + ++ F +GL + ++V LSG+H+IG S C+SF +RL
Sbjct: 502 RDSLGASLSGSNNNIPAPNNTFQTILTKFKLQGLDIVDLVALSGSHTIGKSRCTSFRQRL 681
Query: 211 YSFNLTFPQDPSMDPNFARLLKSKCPPPQSQSINPTVVLDGSTPNDLDNMYYKRLKNNRG 270
Y+ QD ++D +A L+++C P+S LD TP DN Y+K L +G
Sbjct: 682 YNQTGNGKQDFTLDQYYAAELRTQC--PRSGGDQNLFFLDYVTPTKFDNNYFKNLLAYKG 855
Query: 271 LLTSDQTLLNSGL-TRRMVLKNARHAAIWNVKFAKAMVHMGSLDVLTGSEGEIRERCSVV 329
LL+SD+ LL + +V A ++ +FAK+M+ MG++ LTGS G IR C V+
Sbjct: 856 LLSSDEILLTKNQESAELVKLYAERNDLFFEQFAKSMIKMGNISPLTGSRGNIRTNCRVI 1035
Query: 330 N 330
N
Sbjct: 1036N 1038
>TC85149 similar to PIR|JC4780|JC4780 peroxidase (EC 1.11.1.7) 1B precursor
- alfalfa, partial (95%)
Length = 1274
Score = 227 bits (578), Expect = 6e-60
Identities = 131/328 (39%), Positives = 186/328 (55%), Gaps = 5/328 (1%)
Frame = +2
Query: 8 ATLVIVILSVSTTLASSTSLKYGFYKTTCSSVEAIVRRAVNKAVSLNPGIAAGLIRMHFH 67
A +V++ +S L FY+ TC V +I+R + +P + A L+R+HFH
Sbjct: 47 ALCCVVVVLGGLPFSSDAQLDPSFYRDTCPKVHSIIREVIRNVSKTDPRMLASLVRLHFH 226
Query: 68 DCFVRGCDGSVLLDSIPGIQSERDHPANNPSLRGFEVINEAKAQIEAACPKTVSCADILA 127
DCFV GCD SVLL+ I SE++ N SLRG +V+N+ K +E ACP TVSCADILA
Sbjct: 227 DCFVLGCDASVLLNKTDTIVSEQEAFPNINSLRGLDVVNQIKTAVEKACPNTVSCADILA 406
Query: 128 FAARDSARKVSGGRIDYSVPSGRRDGRVSIFDEVTQNLPPPTFSAEQLIDNFDRKGLSVD 187
+A+ S+ G ++ VP GRRDG + QNLP P S +QL F +GLS
Sbjct: 407 LSAQISSILADGP--NWKVPLGRRDGLTANQSLANQNLPAPFNSLDQLKSAFAAQGLSTT 580
Query: 188 EMVTLSGAHSIGVSHCSSFSKRLYSFNLTFPQDPSMDPNFARLLKSKCP---PPQSQSIN 244
++V LSGAH+ G + C+ + RLY+F+ T DP+++ + + L+ CP PP N
Sbjct: 581 DLVALSGAHTFGRARCTFITDRLYNFSSTGKPDPTLNTTYLQELRKICPNGGPP-----N 745
Query: 245 PTVVLDGSTPNDLDNMYYKRLKNNRGLLTSDQTLLN-SGL-TRRMVLKNARHAAIWNVKF 302
D +TP+ D YY L+ +GLL SDQ L + SG T +V K + + F
Sbjct: 746 NLANFDPTTPDKFDKNYYSNLQGKKGLLQSDQELFSTSGADTISIVNKFSADKNAFFDSF 925
Query: 303 AKAMVHMGSLDVLTGSEGEIRERCSVVN 330
AM+ MG++ VLTG +GEIR+ C+ VN
Sbjct: 926 EAAMIKMGNIGVLTGKKGEIRKHCNFVN 1009
>TC78141 similar to GP|5002348|gb|AAD37430.1| peroxidase 5 precursor
{Phaseolus vulgaris}, partial (92%)
Length = 1254
Score = 226 bits (576), Expect = 9e-60
Identities = 134/332 (40%), Positives = 194/332 (58%), Gaps = 3/332 (0%)
Frame = +2
Query: 2 HSILSIATLVIVILSVSTTLASSTSLKYGFYKTTCSSVEAIVRRAVNKAVSLNPGIAAGL 61
+SI ++ L+ ++L+ S +ST FY TC SV +IVR V +A+ +P I A L
Sbjct: 53 YSIFTV--LIFLLLNPSHAQLTST-----FYSNTCPSVSSIVRNVVQQALQNDPRITASL 211
Query: 62 IRMHFHDCFVRGCDGSVLLDSIPGIQ-SERDHPANNPSLRGFEVINEAKAQIEAACPKTV 120
R+HFHDCFV GCD S+LLD I SE++ NN S RGF+V+++ K +E +CP V
Sbjct: 212 TRLHFHDCFVNGCDASLLLDQGGNITLSEKNAVPNNNSARGFDVVDKIKTSVENSCPSVV 391
Query: 121 SCADILAFAARDSARKVSGGRIDYSVPSGRRDGRVSIFDEVTQNLPPPTFSAEQLIDNFD 180
SCADILA AA S +SGG ++V GRRDG ++ ++P PT S + F
Sbjct: 392 SCADILALAAEASV-SLSGGP-SWNVLLGRRDGLIANQSGANTSIPNPTESLANVTAKFA 565
Query: 181 RKGLSVDEMVTLSGAHSIGVSHCSSFSKRLYSFNLTFPQDPSMDPNFARLLKSKCPPPQS 240
GL+ ++V LSGAH+ G C F++RL++F+ T DP+++ + L+ C PQ+
Sbjct: 566 AVGLNTSDLVALSGAHTFGRGQCRFFNQRLFNFSGTGKPDPTLNSTYLATLQQNC--PQN 739
Query: 241 QSINPTVVLDGSTPNDLDNMYYKRLKNNRGLLTSDQTLL--NSGLTRRMVLKNARHAAIW 298
S N LD S+PN+ DN Y+K L N+GLL +DQ L N T +V A + +
Sbjct: 740 GSGNTLNNLDPSSPNNFDNNYFKNLLKNQGLLQTDQELFSTNGAATISIVNNFASNQTAF 919
Query: 299 NVKFAKAMVHMGSLDVLTGSEGEIRERCSVVN 330
F ++M++MG++ L GS+GEIR C VN
Sbjct: 920 FEAFVQSMINMGNISPLIGSQGEIRSDCKKVN 1015
>TC85158 similar to SP|P22195|PER1_ARAHY Cationic peroxidase 1 precursor (EC
1.11.1.7). [Peanut] {Arachis hypogaea}, partial (86%)
Length = 1161
Score = 226 bits (575), Expect = 1e-59
Identities = 127/308 (41%), Positives = 178/308 (57%), Gaps = 1/308 (0%)
Frame = +2
Query: 24 STSLKYGFYKTTCSSVEAIVRRAVNKAVSLNPGIAAGLIRMHFHDCFVRGCDGSVLLDSI 83
S L FY TC V + +++ V A+ + A L+R+HFHDCFV+GCD SVLLD
Sbjct: 134 SAQLSSNFYFRTCPLVLSTIKKEVISALINERRMGASLLRLHFHDCFVQGCDASVLLDDT 313
Query: 84 PGIQSERDHPANNPSLRGFEVINEAKAQIEAACPKTVSCADILAFAARDSARKVSGGRID 143
+ E+ N SLRGF+VI++ K+++E CP TVSCADILA AARDS V+ G +
Sbjct: 314 SSFRGEKTAGPNANSLRGFDVIDKIKSEVEKLCPNTVSCADILAVAARDSV--VALGGLS 487
Query: 144 YSVPSGRRDGRVSIFDEVTQNLPPPTFSAEQLIDNFDRKGLSVDEMVTLSGAHSIGVSHC 203
++V GRRD + F +LP P LI+ F+ KG + EMV LSG+H+IG + C
Sbjct: 488 WTVQLGRRDSTTASFGLANSDLPGPGSDLSGLINAFNNKGFTPKEMVALSGSHTIGEASC 667
Query: 204 SSFSKRLYSFNLTFPQDPSMDPNFARLLKSKCPPPQSQ-SINPTVVLDGSTPNDLDNMYY 262
F R+Y+ N ++D +FA L+S CP +++P LD ++PN DN Y+
Sbjct: 668 RFFRTRIYNEN-------NIDSSFANSLQSSCPRTGGDLNLSP---LDTTSPNTFDNAYF 817
Query: 263 KRLKNNRGLLTSDQTLLNSGLTRRMVLKNARHAAIWNVKFAKAMVHMGSLDVLTGSEGEI 322
K L+N +GL SDQ L + T+ V R+ + V FA AM M +L LTGS G++
Sbjct: 818 KNLQNQKGLFHSDQVLFDEVTTKSQVNSYVRNPLSFKVDFANAMFKMANLGPLTGSSGQV 997
Query: 323 RERCSVVN 330
R+ C VN
Sbjct: 998 RKNCRSVN 1021
>TC85191 similar to PIR|JC4780|JC4780 peroxidase (EC 1.11.1.7) 1B precursor
- alfalfa, partial (96%)
Length = 1307
Score = 225 bits (574), Expect = 2e-59
Identities = 126/330 (38%), Positives = 191/330 (57%), Gaps = 2/330 (0%)
Frame = +2
Query: 3 SILSIATLVIVILSVSTTLASSTSLKYGFYKTTCSSVEAIVRRAVNKAVSLNPGIAAGLI 62
S+++ A +V++ +S+ L FY TC +++IV + + K + + A +I
Sbjct: 95 SLIATALCCVVVVFGGLPFSSNAQLSPDFYAKTCPQLQSIVFQILEKVSKTDSRMPASII 274
Query: 63 RMHFHDCFVRGCDGSVLLDSIPGIQSERDHPANNPSLRGFEVINEAKAQIEAACPKTVSC 122
R+HFHDCFV+GCD SVLL+ I SE+D N SLR +VIN+ K ++E CP VSC
Sbjct: 275 RLHFHDCFVQGCDASVLLNKTSTIASEQDAGPNINSLRRLDVINQIKTEVEKVCPNKVSC 454
Query: 123 ADILAFAARDSARKVSGGRIDYSVPSGRRDGRVSIFDEVTQNLPPPTFSAEQLIDNFDRK 182
ADIL AA S+ V G + VP GRRD + +NLP P+ S +QL +F +
Sbjct: 455 ADILTLAAGVSS--VLSGGPGWIVPLGRRDSLTANQSLANRNLPGPSSSLDQLKSSFAAQ 628
Query: 183 GLSVDEMVTLSGAHSIGVSHCSSFSKRLYSFNLTFPQDPSMDPNFARLLKSKCPPPQSQS 242
GL+ ++V LSGAH++G + C RLY F+ T DP++DP + + L+ +C PQ+
Sbjct: 629 GLNTVDLVALSGAHTLGRARCLFILDRLYDFDNTGKPDPTLDPTYLKQLQKQC--PQNGP 802
Query: 243 INPTVVLDGSTPNDLDNMYYKRLKNNRGLLTSDQTLLNS-GLTRRMVLKN-ARHAAIWNV 300
N V D +TP+ D YY L+ +GLL SDQ L ++ G ++ N + ++
Sbjct: 803 GNNVVNFDPTTPDKFDKNYYNNLQGKKGLLQSDQELFSTPGADTISIVNNFGNNQNVFFQ 982
Query: 301 KFAKAMVHMGSLDVLTGSEGEIRERCSVVN 330
F +M+ MG++ VLTG +GEIR++C+ VN
Sbjct: 983 NFINSMIKMGNIGVLTGKKGEIRKQCNFVN 1072
Database: MTGI
Posted date: Oct 22, 2004 3:39 PM
Number of letters in database: 27,044,181
Number of sequences in database: 36,976
Lambda K H
0.319 0.133 0.385
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 9,979,910
Number of Sequences: 36976
Number of extensions: 135045
Number of successful extensions: 1059
Number of sequences better than 10.0: 154
Number of HSP's better than 10.0 without gapping: 831
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 846
length of query: 330
length of database: 9,014,727
effective HSP length: 96
effective length of query: 234
effective length of database: 5,465,031
effective search space: 1278817254
effective search space used: 1278817254
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.8 bits)
S2: 58 (26.9 bits)
Medicago: description of AC136973.10


