

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC136973.1 - phase: 0 /pseudo
(381 letters)
Database: MTGI
36,976 sequences; 27,044,181 total letters
Searching..................................................done
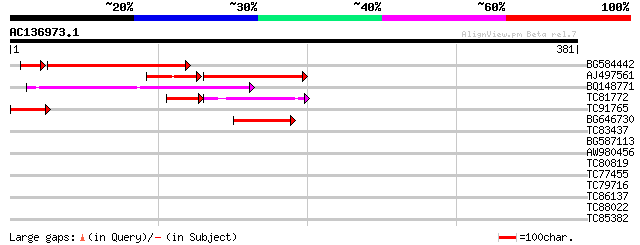
Score E
Sequences producing significant alignments: (bits) Value
BG584442 115 2e-27
AJ497561 weakly similar to GP|10140689|gb putative non-LTR retro... 70 2e-17
BQ148771 62 4e-10
TC81772 51 5e-10
TC91765 weakly similar to GP|19881779|gb|AAM01180.1 Putative ret... 44 1e-04
BG646730 41 7e-04
TC83437 weakly similar to PIR|D86384|D86384 unknown protein [imp... 38 0.007
BG587113 weakly similar to PIR|A84888|A8 hypothetical protein At... 34 0.082
AW980456 33 0.24
TC80819 weakly similar to GP|10140689|gb|AAG13524.1 putative non... 32 0.53
TC77455 similar to GP|22335695|dbj|BAC10549. nine-cis-epoxycarot... 30 1.2
TC79716 weakly similar to SP|O24495|GL2M_ARATH Hydroxyacylglutat... 29 3.4
TC86137 weakly similar to PIR|A96673|A96673 probable cytochrome ... 29 3.4
TC88022 weakly similar to PIR|T50009|T50009 hypothetical protein... 28 5.9
TC85382 similar to GP|21740531|emb|CAD41510. OSJNBa0029H02.26 {O... 28 7.7
>BG584442
Length = 775
Score = 115 bits (287), Expect(2) = 2e-27
Identities = 59/97 (60%), Positives = 73/97 (74%), Gaps = 1/97 (1%)
Frame = +1
Query: 26 QKISSWSSKCLSKAGREVMIKYVLQAIPSYVMSIFRLSNTLLDEIE-MMNSFWWGHGGSS 84
+KI+ +KCLSK EVMIKY LQ+I SYVMSIF L N+ +DEIE +MN+F W H G +
Sbjct: 379 KKINF*RNKCLSKVM*EVMIKYALQSISSYVMSIFLLLNSQVDEIEKIMNTFSWVHVGEN 558
Query: 85 NRGLNWLSWEKLSVHKNDGGMGFKDFVAFNVAMLGKQ 121
+G++W+S EKL VHKN GGMGF DF FN+ MLGKQ
Sbjct: 559 RKGMHWMS*EKLFVHKNYGGMGFTDFTTFNIPMLGKQ 669
Score = 25.4 bits (54), Expect(2) = 2e-27
Identities = 10/17 (58%), Positives = 13/17 (75%)
Frame = +3
Query: 8 MVGRSKKATFSFIKDRV 24
M+ S KATFS+IKD +
Sbjct: 324 MLSTSNKATFSYIKDEI 374
>AJ497561 weakly similar to GP|10140689|gb putative non-LTR retroelement
reverse transcriptase {Oryza sativa (japonica
cultivar-group)}, partial (2%)
Length = 621
Score = 70.1 bits (170), Expect(2) = 2e-17
Identities = 33/70 (47%), Positives = 45/70 (64%)
Frame = +2
Query: 131 SLVSKIFKARYYPNSNFLDAKLGHNPSFVWRSILSAKVVVRQGARWKIGAGFDIPIISEP 190
SL+SKI K +Y+P +F A LGHNPSF WRS+LS + ++ G RW IG G I + S
Sbjct: 410 SLLSKILKFKYFPQWDFSYANLGHNPSFTWRSLLSTQSLLTLGHRWMIGDGSQINVSSMS 589
Query: 191 WIGSGSSIPP 200
WI + ++ P
Sbjct: 590 WIRNRPTLRP 619
Score = 36.6 bits (83), Expect(2) = 2e-17
Identities = 16/37 (43%), Positives = 27/37 (72%)
Frame = +3
Query: 93 WEKLSVHKNDGGMGFKDFVAFNVAMLGKQGWQLQTKP 129
+ L+ HK+ GG+G ++ FN++MLGKQ W+L ++P
Sbjct: 303 FNNLARHKSVGGLGLQN-QEFNISMLGKQRWKLLSQP 410
>BQ148771
Length = 680
Score = 62.0 bits (149), Expect = 4e-10
Identities = 40/155 (25%), Positives = 79/155 (50%), Gaps = 2/155 (1%)
Frame = -3
Query: 12 SKKATFSFIKDRVWQKISSWSSKCLSKAGREVMIKYVLQAIPSYVMSIFRLSNTLLDEIE 71
+KK+ FS + +V +++W + LS A R + K V++A+P Y M + ++EI+
Sbjct: 618 TKKSRFS-VYYQVHVMLANWKANHLSLARRVTLAKSVIEAVPLYPMMTTIIPKACIEEIQ 442
Query: 72 -MMNSFWWGHGGSSNRGLNWLSWEKLSVHKNDGGMGFKDFVAFNVAMLGKQGWQLQTKPD 130
+ F WG S R + + WE +S K G+G + N A + K GW + + +
Sbjct: 441 KLQRKFVWGDTEVSRR-YHAVGWETMSKPKTIYGLGLRRLDVMNKACIMKLGWSIYSGSN 265
Query: 131 SLVSKIFKARYYPNSNFLDAKLGH-NPSFVWRSIL 164
SL +++ + +Y + + + L S +W++++
Sbjct: 264 SLCTEVMRGKYQRSESLEEIFLEKPTDSSLWKALV 160
>TC81772
Length = 982
Score = 50.8 bits (120), Expect(2) = 5e-10
Identities = 29/71 (40%), Positives = 41/71 (56%)
Frame = -2
Query: 131 SLVSKIFKARYYPNSNFLDAKLGHNPSFVWRSILSAKVVVRQGARWKIGAGFDIPIISEP 190
SL SKIF L +GHNPS++W S+ + ++V+ +G WKIG G I I ++
Sbjct: 765 SLRSKIFFTL-----ELLGVVIGHNPSYIWCSV*TLRMVLEEGHHWKIGDGSSINIWTDH 601
Query: 191 WIGSGSSIPPV 201
W+ SG IP V
Sbjct: 600 WL-SGMYIPCV 571
Score = 30.4 bits (67), Expect(2) = 5e-10
Identities = 12/25 (48%), Positives = 16/25 (64%)
Frame = -1
Query: 106 GFKDFVAFNVAMLGKQGWQLQTKPD 130
GF D ++ MLGKQ W+L T P+
Sbjct: 853 GFHDIYGLDLVMLGKQVWKLLTNPN 779
>TC91765 weakly similar to GP|19881779|gb|AAM01180.1 Putative retroelement
{Oryza sativa (japonica cultivar-group)}, partial (1%)
Length = 625
Score = 43.5 bits (101), Expect = 1e-04
Identities = 19/27 (70%), Positives = 21/27 (77%)
Frame = +3
Query: 1 KYLGLPSMVGRSKKATFSFIKDRVWQK 27
KYLGLPSM+GR + TFS IK VWQK
Sbjct: 489 KYLGLPSMIGRDRTTTFSSIKGGVWQK 569
Score = 33.5 bits (75), Expect = 0.14
Identities = 16/29 (55%), Positives = 20/29 (68%)
Frame = +1
Query: 17 FSFIKDRVWQKISSWSSKCLSKAGREVMI 45
F ++ +KI+SWS CLSKA REVMI
Sbjct: 538 FLLLRVECGKKINSWSDICLSKACREVMI 624
>BG646730
Length = 799
Score = 41.2 bits (95), Expect = 7e-04
Identities = 15/42 (35%), Positives = 27/42 (63%)
Frame = +1
Query: 151 KLGHNPSFVWRSILSAKVVVRQGARWKIGAGFDIPIISEPWI 192
++G+ PS+ WRS+ + K V+ G+RW I G ++ I + W+
Sbjct: 382 EIGNQPSYAWRSMFNIKDVIDLGSRWSISNGQNVRIWKDDWL 507
>TC83437 weakly similar to PIR|D86384|D86384 unknown protein [imported] -
Arabidopsis thaliana, partial (6%)
Length = 951
Score = 37.7 bits (86), Expect = 0.007
Identities = 21/62 (33%), Positives = 39/62 (62%), Gaps = 1/62 (1%)
Frame = +2
Query: 2 YLGLPSMVGRSKKATF-SFIKDRVWQKISSWSSKCLSKAGREVMIKYVLQAIPSYVMSIF 60
YLG+P + G S++ +F I +R+ +++ W+S+ LS GR V++K VL ++ Y +
Sbjct: 761 YLGMP-IEGNSRRLSFWEPIVNRIKARLTGWNSRFLSFGGRLVLLKSVLTSLSVYALPSS 937
Query: 61 RL 62
+L
Sbjct: 938 KL 943
>BG587113 weakly similar to PIR|A84888|A8 hypothetical protein At2g45230
[imported] - Arabidopsis thaliana, partial (10%)
Length = 767
Score = 34.3 bits (77), Expect = 0.082
Identities = 17/34 (50%), Positives = 24/34 (70%)
Frame = -3
Query: 148 LDAKLGHNPSFVWRSILSAKVVVRQGARWKIGAG 181
L+A LG S+ WRSI SA+ +++QGA+ IG G
Sbjct: 762 LNAPLGSWASYAWRSIHSAQHLIKQGAKVIIGNG 661
>AW980456
Length = 779
Score = 32.7 bits (73), Expect = 0.24
Identities = 19/66 (28%), Positives = 38/66 (56%)
Frame = -1
Query: 255 DLEGGEKWPLFCEKCISYMHRRSHQQ*PFAQTRVLKWHLETKSPPQSQEFGMENL*RLFS 314
DLEG + W + C++C+S++ R + + + +R L +LE S + +E++* LF+
Sbjct: 200 DLEG*KAWEILCKECLSFLCRGTLRFFLPSPSR*LVRNLEA*STS*GSKLSLEDV*GLFT 21
Query: 315 NSCETT 320
+T+
Sbjct: 20 YENQTS 3
>TC80819 weakly similar to GP|10140689|gb|AAG13524.1 putative non-LTR
retroelement reverse transcriptase {Oryza sativa
(japonica cultivar-group)}, partial (2%)
Length = 1262
Score = 31.6 bits (70), Expect = 0.53
Identities = 13/30 (43%), Positives = 20/30 (66%)
Frame = +1
Query: 112 AFNVAMLGKQGWQLQTKPDSLVSKIFKARY 141
AFN+++LGK W+L + L ++ KARY
Sbjct: 46 AFNLSLLGKWCWRLLVDKEGLWHRVLKARY 135
>TC77455 similar to GP|22335695|dbj|BAC10549. nine-cis-epoxycarotenoid
dioxygenase1 {Pisum sativum}, partial (43%)
Length = 1865
Score = 30.4 bits (67), Expect = 1.2
Identities = 14/41 (34%), Positives = 22/41 (53%)
Frame = -3
Query: 103 GGMGFKDFVAFNVAMLGKQGWQLQTKPDSLVSKIFKARYYP 143
GG+G +D NV++L K W+L SL ++ + Y P
Sbjct: 951 GGLGVRDIRLVNVSLLAKWWWRLLQDQSSLWKEVLEDIYGP 829
>TC79716 weakly similar to SP|O24495|GL2M_ARATH Hydroxyacylglutathione
hydrolase mitochondrial precursor (EC 3.1.2.6)
(Glyoxalase II) (Glx II)., partial (94%)
Length = 1241
Score = 28.9 bits (63), Expect = 3.4
Identities = 19/36 (52%), Positives = 21/36 (57%), Gaps = 1/36 (2%)
Frame = +2
Query: 10 GRSKKAT-FSFIKDRVWQKISSWSSKCLSKAGREVM 44
GR KKAT FS + WQ ISS S + LSKA M
Sbjct: 14 GRQKKATSFSTKQLN*WQPISSLSFQMLSKASTTAM 121
>TC86137 weakly similar to PIR|A96673|A96673 probable cytochrome P450
F13O11.25 [imported] - Arabidopsis thaliana, partial
(69%)
Length = 1695
Score = 28.9 bits (63), Expect = 3.4
Identities = 16/59 (27%), Positives = 26/59 (43%)
Frame = -2
Query: 158 FVWRSILSAKVVVRQGARWKIGAGFDIPIISEPWIGSGSSIPPVGDDMLALQPYSVGSN 216
F W L +KV + W I +G D+ +++ GS S + +D L VG +
Sbjct: 494 FGWMKHLGSKVTTKGTPCWAIISGVDVVLVAGQHFGSRKSCRAIREDG*VLNENMVGQS 318
>TC88022 weakly similar to PIR|T50009|T50009 hypothetical protein T31P16.40
- Arabidopsis thaliana, partial (87%)
Length = 1093
Score = 28.1 bits (61), Expect = 5.9
Identities = 12/30 (40%), Positives = 17/30 (56%)
Frame = -1
Query: 108 KDFVAFNVAMLGKQGWQLQTKPDSLVSKIF 137
+ F F VAM G W + +P +LVS +F
Sbjct: 427 QSFPGFLVAMWGTTAWTILMEPKTLVSNVF 338
>TC85382 similar to GP|21740531|emb|CAD41510. OSJNBa0029H02.26 {Oryza
sativa}, partial (26%)
Length = 849
Score = 27.7 bits (60), Expect = 7.7
Identities = 18/49 (36%), Positives = 26/49 (52%)
Frame = -1
Query: 26 QKISSWSSKCLSKAGREVMIKYVLQAIPSYVMSIFRLSNTLLDEIEMMN 74
QKIS W + L K + ++K Q I SY M + +L + EI+M N
Sbjct: 441 QKISGWFAGALHKNKAKRVLKKQSQIIQSYTM-MLKLIKGIKIEIKMRN 298
Database: MTGI
Posted date: Oct 22, 2004 3:39 PM
Number of letters in database: 27,044,181
Number of sequences in database: 36,976
Lambda K H
0.339 0.148 0.518
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 13,965,537
Number of Sequences: 36976
Number of extensions: 211549
Number of successful extensions: 1483
Number of sequences better than 10.0: 30
Number of HSP's better than 10.0 without gapping: 1473
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 1482
length of query: 381
length of database: 9,014,727
effective HSP length: 98
effective length of query: 283
effective length of database: 5,391,079
effective search space: 1525675357
effective search space used: 1525675357
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 15 ( 7.3 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 39 (21.9 bits)
S2: 59 (27.3 bits)
Medicago: description of AC136973.1


